Figure 4.
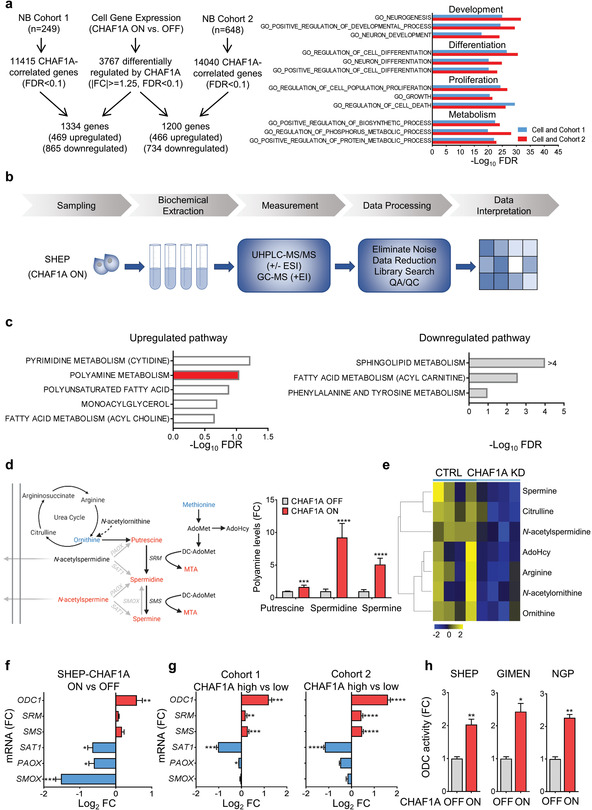
CHAF1A gene expression and pathway analyses of NB cells and patients. a) Left: overlap of differentially expressed genes (DEGs, |(fc)| > = 1.25, FDR < 0.1) between control (CHAF1A OFF) and CHAF1A‐overexpressing SHEP cells (CHAF1A ON, 96 h) and CHAF1A‐correlated genes (FDR < 0.1) in patient cohort 1 (n = 249) and 2 (n = 648). Right: GO pathway enrichment analysis of the overlapped genes (ranked by −Log10FDR, FDR<0.05). b) Work flow of the metabolomics analysis: global metabolomics analysis was performed by GC‐MS and LC‐MS (DiscoveryHD4 platform, Metabolon Inc.) in CHAF1A‐overexpressing SHEP cells (DOX 1 µg mL−1 for 0, 24, and 72 h, n = 5). c) Metabolite enrichment analysis depicts the pathways significantly up‐ and down‐regulated by CHAF1A (DOX 24 h, FDR < 0.25); Benjamini–Hochberg corrected two‐sided homoscedastic t‐test. d) Left: schematic presentation (redrawn from Gamble et al.[ 52 ]) of the polyamine pathway with metabolite changes in SHEP cells with or without CHAF1A overexpression for 24 h (red = upregulated metabolites, p ≤ 0.05; blue = downregulated metabolites, p ≤ 0.05). Right: polyamine levels in SHEP cells with or without CHAF1A overexpression for 24 h. Data are mean ± SD (n = 5). e) Targeted polyamine analysis in IMR32 cells with conditional KD of CHAF1A (DOX 1 µg mL−1 for 5 days). Differential metabolites (FDR < 0.25) are presented in the heatmap (yellow = upregulated; blue = downregulated) (n = 4). f) Polyamine synthetic and catabolic gene expression in SHEP cells with or without CHAF1A overexpression (24 h). Data are mean ± SD (n = 2); *p < 0.05, **p < 0.01, ***p < 0.001; two‐sided unpaired t‐test. g) Polyamine gene expression in patients with high and low CHAF1A expression (average CHAF1A mRNA expression ± 1SD, Figure 1) in patient cohorts 1 and 2. Data are mean ± SEM (n = 44 in cohort 1 and n = 107 in cohort 2); *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; two‐sided unpaired t‐test. h) ODC1 activity in SHEP, GIMEN, and NGP cells with or without CHAF1A overexpression (8 h). One unit is defined as the fluorescence change per minute. Data are normalized by the protein amount and presented as the fold change compared to control (mean ± SD, n = 2); *p < 0.05, **p < 0.01; two‐sided unpaired t‐test. MTA = 5'‐methylthioadenosine; AdoMet = S‐(5'‐Adenosyl)‐L‐methionine; AdoHyc = S‐(5′‐Adenosyl)‐L‐homocysteine; FC = fold change.
