Figure 6.
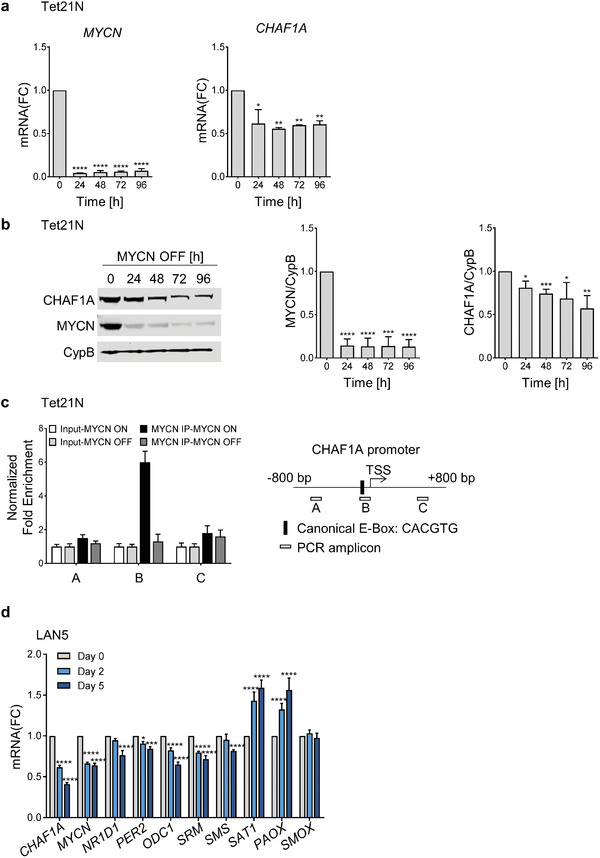
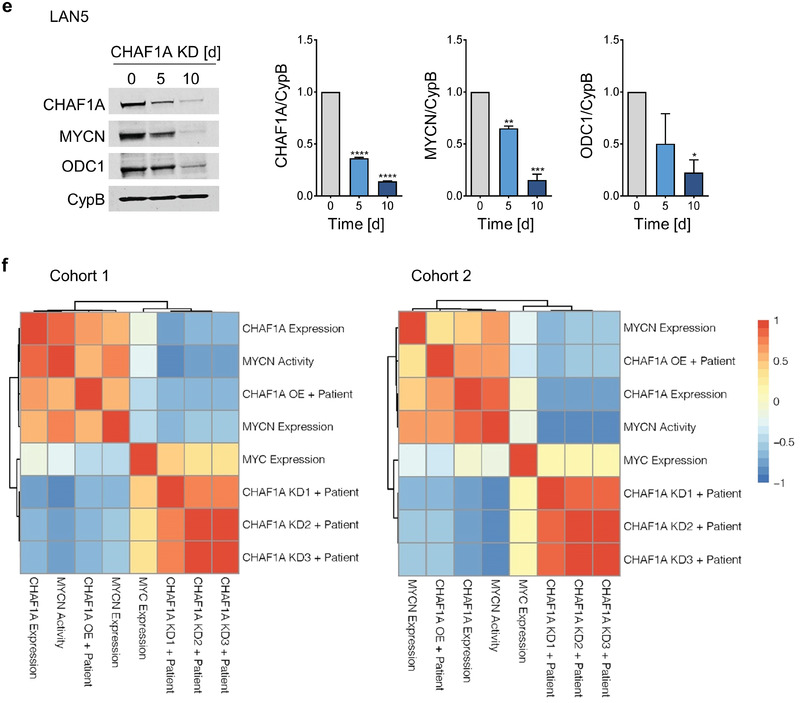
CHAF1A is a direct target of MYCN. a,b) mRNA and protein expression of CHAF1A in TET‐21/N cells when MYCN is turned off upon DOX treatment (2 µg mL−1, 24–96 h). GAPDH is used as housekeeping gene, CypB as protein loading control. Data are mean ± SD (n = 2–3); * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001; one‐way ANOVA with Dunnett's multiple comparisons test. c) MYCN ChIP‐qPCR assays in TET‐21/N cells. Input (white bars) and MYCN‐ChIP (black bars) samples were analyzed by qPCR using specific primers for CHAF1A (Table S5, Supporting Information). Data from two independent experiments are shown (mean ± SEM, n = 2). d) mRNA expression of CHAF1A, MYCN, MYCN targets and polyamine genes in LAN5 cells upon CHAF1A KD (DOX 1 µg mL−1 for 2–5 days). GAPDH served as control. Mean ± SD (n = 3); * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001; two‐way ANOVA with Dunnett's multiple comparisons test. e) Protein expression of CHAF1A, MYCN, and ODC1 in LAN5 cells upon CHAF1A KD (DOX 1 µg mL−1 for 0–10 days). CypB served as protein loading control. Mean ± SD (n = 2); * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001; one‐way ANOVA with Dunnett's multiple comparisons test. f) Correlation of CHAF1A, MYCN, and MYCN signature scores in patient cohort 1 (n = 249) and cohort 2 (n = 648). Signatures are defined in the methods section. FC = fold change.