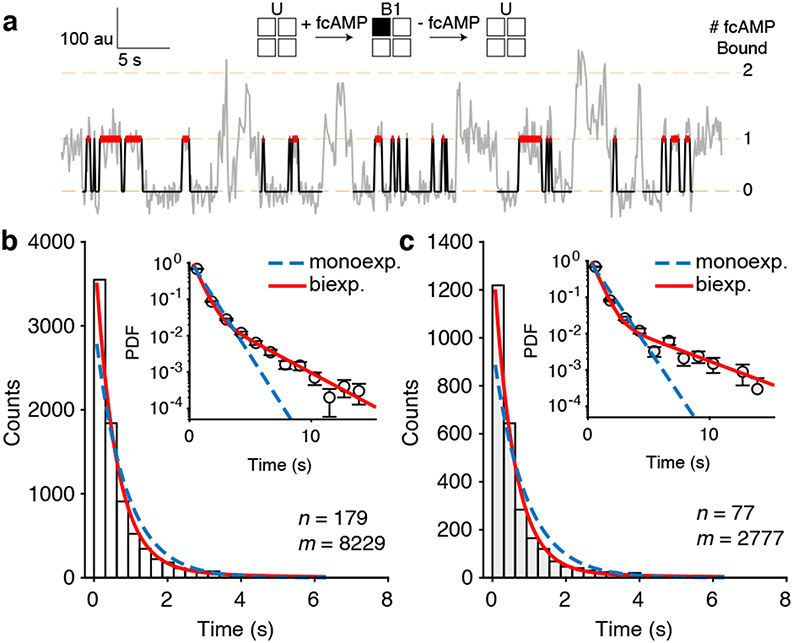
Fig. 3: Ligand binding induces a conformational change at each HCN subunit.
(a) Representative fcAMP binding trajectory with identified isolated-B1 dwell times (red). (b, c) Dwell time distributions of isolated-B1 events for HCN1SM (b) and HCN2SM (c) at 250 nM fcAMPoverlaid with monoexponential (blue dashed) and biexponential (red) fits (Methods). In each panel, n is the total number of molecules and m is the total number of dwell times. Abscissa values correspond to the center of the histogram bars. Inset of (b) and (c) shows probability density function (PDF) of each distribution with the ordinate on a log scale to highlight less frequent but long-lived bound durations. Error bars for each bin are calculated as the error of binomial distribution. See Extended Data Fig. 6-7 for isolated-B1 dwell time distributions and Supplementary Table 3 for all fit parameters.