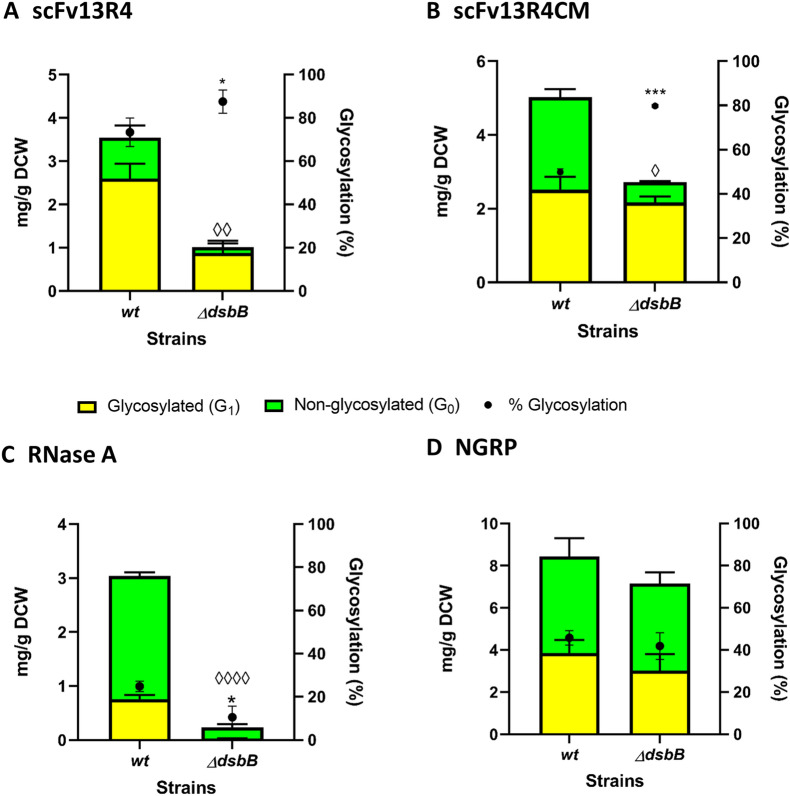
Fig. 6.
Glycosylation of disulphide bond-containing proteins in oxidoreductase mutant (ΔdsbB) of E. coli. Quantitative Western blot analysis (densitometry) of A scFv13R4, B scFv13R4CM, C RNase A and D control non-disulphide bond-containing protein NGRP expressed in periplasmic of glyco-competent E. coli wild-type (wt) or ΔdsbB strain. Total proteins (A–D) were quantified using pre-determined purified scFv13R4CM, RNase A, or NGRP standard curve (5 ng to 100 ng). The data were converted into yield mg/g of dry cell weight (DCW) based on normalisation and calculation with measured OD600 of the samples Glycosylated (yellow bar) and non-glycosylated (green bar) protein as shown (left y-axis). % Glycosylation (% G1/G0 + G1) is indicated (black circle, right y-axis). Additional file 1: Figure S16, A and B showed representative Western blot images for scFv13R4CM and RNase A produced in wt and ΔdsbB strains. Representative Western blot image for NGRP and scFv13R4 have been shown before in Figs. 3B, C, and 4B, C. Statistical analysis was conducted by unpaired t-test with Welch’s correction to control sample expressed in wt strain (P < 0.05*, < 0.01**, for % glycosylation; P < 0.05◊, < 0.01◊◊, < 0.0001◊◊◊◊, for total protein mg/g DCW). All data were processed from three biological replicates. Error bars indicate standard deviation from mean values