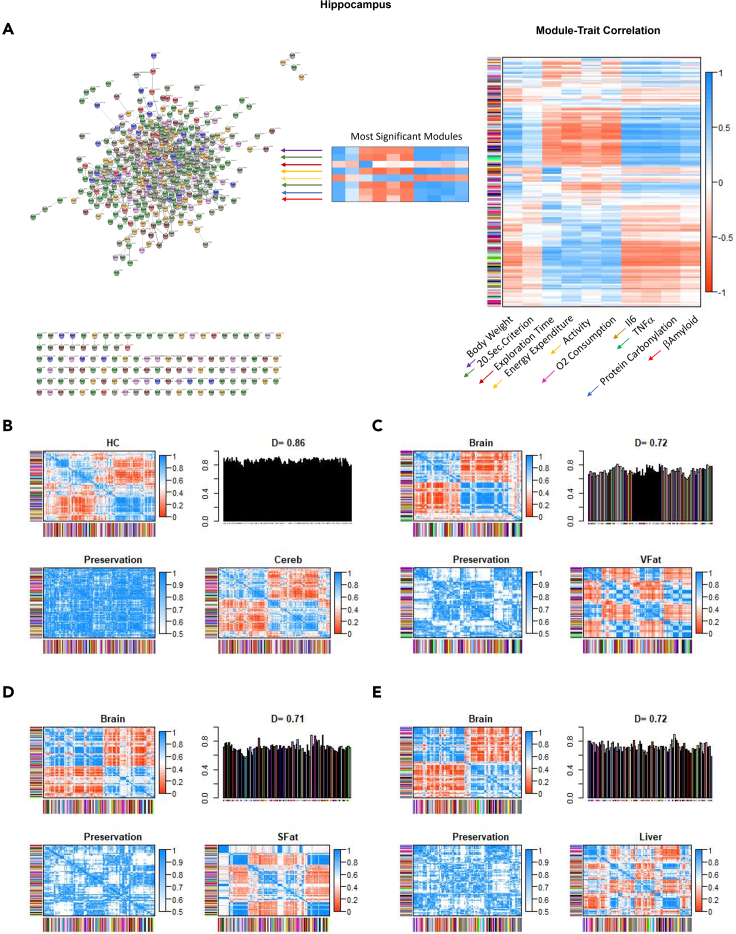
Figure 6.
Differential eigengene network analysis and module-trait correlation of datasets from tissues of mice
(A–E)(A) Module-trait correlations. Each row in the table (right panel) corresponds to different gene groupings, and each column to selected phenotypical features. Based on the highest correlations with these phenotypic features (body weight, time to 20 s criterion, exploration time, locomotion, energy expenditure, oxygen consumption, plasma IL-6, plasma TNFα, protein carbonylation, and Aβ40). Modules which correlated best with the 10 phenotypic traits (there were 7 such modules as some correlated best with more than one trait) were placed into a cytoscape network plot using the STRING protein-protein interaction dataset (left panel). We note that although most of these module genes are assigned to such interactions in this dataset, a significant minority were not. Summary plot of consensus eigengene networks and their differential analysis from (B) hippocampus and cerebellum datasets, (C) brain and visceral fat datasets, (D) brain and subcutaneous fat datasets, (E) brain and liver datasets. Heat maps show high (red) and low (or negative, green) adjacency. Preservation heatmap is the 1-absolute difference of the eigengene networks in the two sets. Bar plot shows mean preservation of adjacency for each eigengene to other eigengenes with a D value calculated as the arithmetic mean of these measurements. N = 6/group.