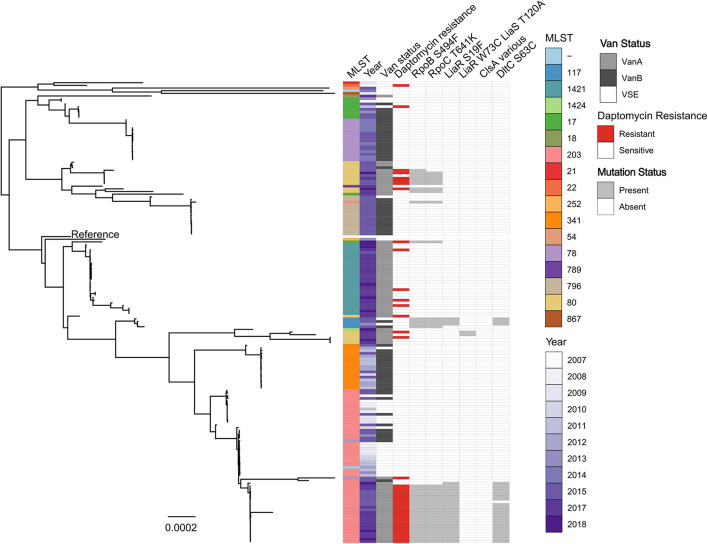
FIGURE 3.
Maximum likelihood core-SNP phylogeny of 173 Australian E. faecium study isolates. Coloured blocks show the multi-locus sequence type (MLST), country of origin (country), year of isolation (year), van status (i.e., vanA, vanB, and VSE) and daptomycin status (resistant or susceptible) for each isolate within the phylogeny. Grey blocks show the presence of the RNAP β-subunit S494F mutation (RpoB S494F), the RNAP β′-subunit T641K mutation (RpoC T641K), the LiaR S19F mutation, the DltC S63C mutation, the LiaR W73C and LiaS T120A co-occurring mutations or mutations within ClsA previously associated with daptomycin resistance in VREfm (ClsA various) for each isolate within the phylogeny. The vanA-VREfm strain D0 (TX16) (Diaz et al., 2014) was used as the reference.