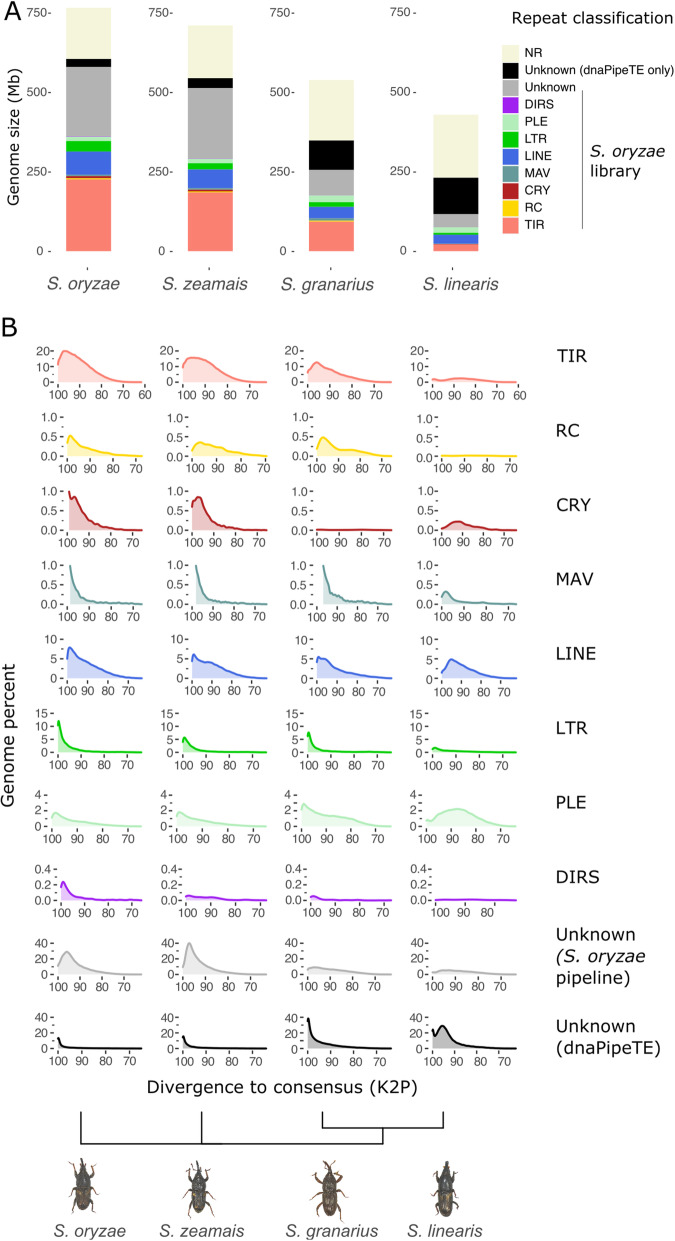
Fig. 6.
TE landscape across Sitophilus species. A Proportion of TE per species estimated from short reads with dnaPipeTE and a custom TE library including Repbase (release 2017) and annotated TE consensus discovered in S. oryzae. S. oryzae, S. zeamais, and S. granarius harbor similar TE content, while S. granarius presents a smaller TE load, and S. linearis harbors the smallest TE content and the higher proportion of unknown repeats. The proportion of unknown repeats only found by dnaPipeTE (black) increases from S. oryzae to S. linearis with the phylogenetic distance. B Distribution of divergence values between raw reads and repeats contig assembled with dnaPipeTE (blastn) across four Sitophilus species. S. oryzae appears to share its TE landscape with S. zeamais and S. granarius, but the three species display a distinct repeatome than S. linearis, in spite of their phylogenetic proximity. SO2: S. oryzae’s TE library produced in this analysis, DPTE: DNApipeTE TE annotation (repeats only found by dnaPipeTE)