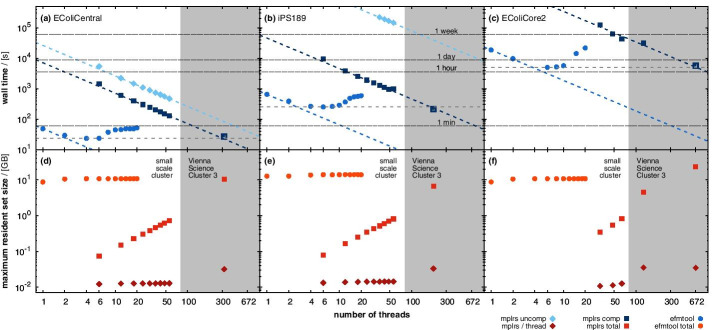
Fig. 4.
Performance comparison of efmtool [12] (circles) and mplrs [1] (full squares) enumerating EFMs in the flux cone (9) of the three compressed metabolic models: EColiCentral [28] (a, d), EColiCore2 [29] (b, e) and iPS189 [30] (c, f). Top panels compare run time as function of thread number. Additionally, we checked mplrs’s run time behavior for the uncompressed models (triangles). Dotted lines indicate efmtool’s minimum wall time, and a power-law fit to mplrs’s wall time, respectively. The intersections of the dotted lines mark (predicted) points where mplrs is as fast as the fastest efmtool run. We validated these points by running mplrs with the predicted number of threads (open squares). Note, open squares were not used in the power-law fit. The 4 thin gray lines indicate 1 min, 1 h, 1 day and 1 week—as stated in the upper middle plot. Bottom panels compare maximum resident set size during EFM enumeration as function of thread number. For mplrs, we also plotted the maximum resident set size per thread (open squares). Two different clusters were used for this analysis as indicated by the different background shadings