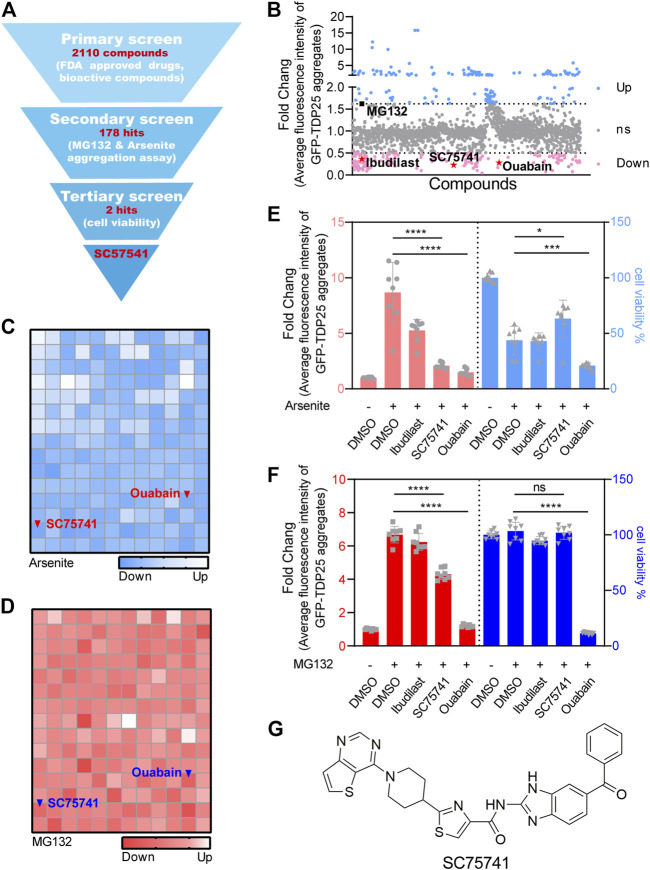
FIGURE 1.
Identification of TDP25 degrader by high-throughput screening. (A). Flowchart of the drug screening and its results. (B). H4GT25 cells were pretreated with 1 μg/mL DOX (doxycycline) for 24 h, incubated with 2110 FDA approved drugs and bioactive compounds for 24 h, and the GFP-TDP25 fluorescence was analyzed by fluorescence microscopy and compared with cells treated with DMSO. The results were presented in the forms of scatter diagram. Skyblue and pink colors represent the degree of increase and decrease of GFP-TDP25 expression, respectively, following drug treatments. Shown in black scatterplot is MG132, and shown in red are ibudilast, ouabain, and SC75741. (C). H4GT25 cells were pretreated with 1 μg/mL DOX for 24 h, treated with arsenite (0.5 mM) for 1 h, incubated with 178 hits for 24 h, and the GFP-TDP25 fluorescence was analyzed by fluorescence microscopy and compared with cells treated with DMSO. The results were presented in the forms of blue heatmap. (D). H4GT25 cells were pretreated with 1 μg/mL DOX for 24 h, treated with MG132 (1 μM) for 12 h, incubated with 178 hits for 24 h, and the GFP-TDP25 fluorescence was analyzed by fluorescence microscopy and compared with cells treated with DMSO. The results were presented in the forms of red heatmap. (E). H4GT25 cells were pretreated with 1 μg/mL DOX for 24 h, treated with arsenite (0.5 mM) for 1 h, and treated with ibudilast (10 μM), SC75741 (10 μM) or ouabain (10 μM) for another 24 h, fluorescence of GFP-TDP25 was analyzed by fluorescence microscopy and Cell viability was determined using CellTiterGlo® assay (data represents Mean ± SD.; n = 8, **** p < 0.0001, *** p < 0.001, * p < 0.05, two-tailed t test). (F). H4GT25 cells were pretreated with 1 μg/mL DOX for 24 h, treated with MG132 (1 μM) for 12 h, and treated with ibudilast (10 μM), SC75741 (10 μM) or ouabain (10 μM) for another 24 h, fluorescence of GFP-TDP25 was analyzed by fluorescence microscopy and Cell viability was determined using CellTiterGlo® assay (data represents Mean ± SD.; n = 8, **** p < 0.0001, two-tailed t test). (G). Chemical structure of SC75741.