Fig. 1.
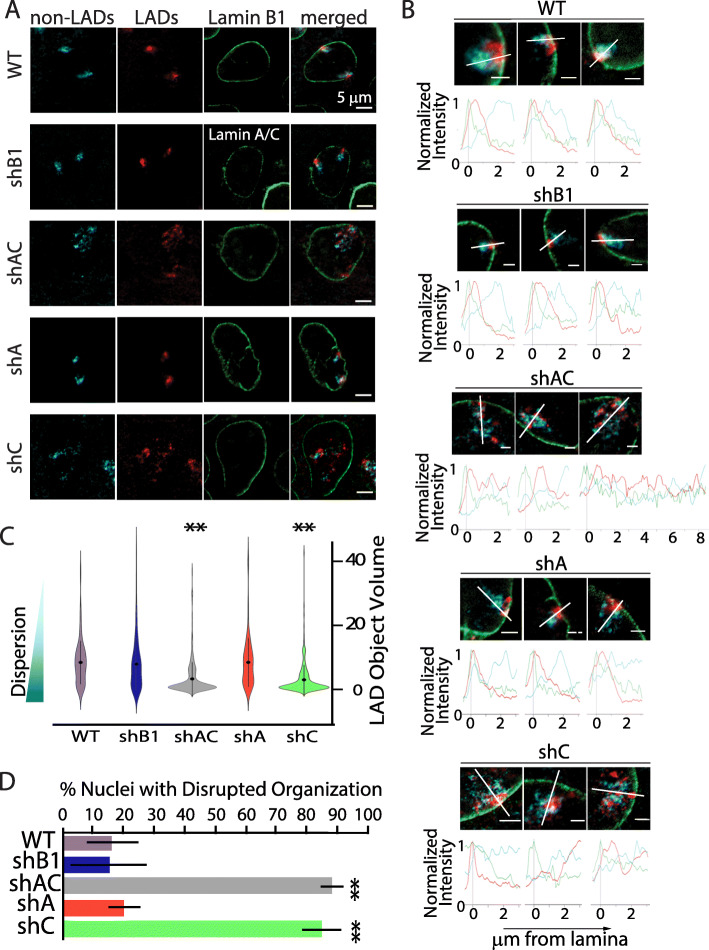
Specific knockdown of lamin C disrupts LAD aggregation and localization at the single cell level. A Representative images showing the organization of chromosome 11 with nonLADs in cyan, LADs in red, and lamin B1 (or lamin A/C for shB1) in green. ShA representative image shows one territory that is attached in the medial plane and one territory that is attached at the top of the nucleus where a portion of the LAD aggregate hangs down into the medial plane. B Representative images of chromosome conformation paints to chromosome nonLADS (cyan), LADs (red), and lamin B1 (green) in primary MEFs. Images were chosen to represent the spectrum of phenotypes for each knockdown. Normalized fluorescence intensity histogram plots, from nuclear lamina (0μm) to 3μm into the nucleus, were plotted for all chromosome 11 territories (except territory 3 of shAC which required a longer measurement) to display the extent of LAD signals. The line each plot travels through is represented by a white line. Scale bar = 2μm. C Violin plots showing the distribution of the volumes of segmented objects based on LAD signals, with smaller objects indicating greater dispersion of LADs (n>50 territories per condition). ShAC (gray) and shC (green) show significant LAD dispersion. Statistically significant changes from wild-type dispersion were noted in shAC and shC conditions (marked with **, p<0.001). D Quantification of nuclei with disrupted organization for each knockdown condition. Error bars represent 1 standard deviation. ** indicates statistically significant differences in percent disrupted nuclei when compared to WT control (t-test p value <0.001; n>200 nuclei per condition)