Significance
Isolated and infrequently colonized, islands harbor many of nature’s most renowned evolutionary radiations. Despite this evolutionary exuberance, island occupation has long been considered irreversible: The much tougher competitive and predatory milieu on mainlands prevents colonization, much less evolutionary diversification, from islands to continents. To test these postulates, we examined neotropical Anolis lizards, asking what happens when mainland and island evolutionary radiations collide. Far from being a dead end, we show that island-to-mainland colonization seeded an extensive radiation that achieved its ecomorphological disparity in ways distinct from their island ancestors. Moreover, when the incumbent and island-derived radiations collided, the ensuing interactions favored the latter, together highlighting a persistent role of both historical contingency and determinism in adaptive radiation.
Keywords: Anolis, macroevolution, adaptive radiation, convergence, diversification
Abstract
Oceanic islands are known as test tubes of evolution. Isolated and colonized by relatively few species, islands are home to many of nature’s most renowned radiations from the finches of the Galápagos to the silverswords of the Hawaiian Islands. Despite the evolutionary exuberance of insular life, island occupation has long been thought to be irreversible. In particular, the presumed much tougher competitive and predatory milieu in continental settings prevents colonization, much less evolutionary diversification, from islands back to mainlands. To test these predictions, we examined the ecological and morphological diversity of neotropical Anolis lizards, which originated in South America, colonized and radiated on various islands in the Caribbean, and then returned and diversified on the mainland. We focus in particular on what happens when mainland and island evolutionary radiations collide. We show that extensive continental radiations can result from island ancestors and that the incumbent and invading mainland clades achieve their ecological and morphological disparity in very different ways. Moreover, we show that when a mainland radiation derived from island ancestors comes into contact with an incumbent mainland radiation the ensuing interactions favor the island-derived clade.
Historically, conceptions of island evolution have been contradictory. Because oceanic islands initially have few species, island resources often can be underutilized, presenting an “ecological opportunity” for evolutionary diversification. Indeed, essentially all textbook cases of adaptive radiation come from clades that evolved on islands or in island-like settings (e.g., lakes) (1–3). Yet, the reduced species richness on islands has led to the presumption that interspecific interactions are less intense there and that island species rarely reinvade, much less diversify, in continental settings because they are not adapted to strong competitive and predatory interactions (4–7). Many examples now show this premise to be incorrect: Mainland-to-island colonization is not a one-way street (8–13). Nonetheless, a disparity in evolutionary outcomes is still evident: Continental species can give rise to spectacular adaptive radiations on islands, yet the converse has scarcely been reported.
Island anoles (Fig. 1 A–C) are a textbook example of adaptive radiation (1, 14). Anoles have radiated independently on each of the main islands of the Greater Antilles, resulting in highly similar suites of habitat specialist species—termed ecomorphs—on each island (15). Correlations between habitat use and morphology suggest species have evolved to capitalize on different microhabitats, and detailed studies of behavior, biomechanics, and natural selection have bolstered our understanding of the adaptive basis of these radiations (14).
Fig. 1.
Morphological and ecological diversity in Caribbean (A–C) and Mainland (D–F) Anolis lizards. (A) Anolis guamuhaya, a twig anole, (B) Anolis occultus, a twig anole, (C) Anolis bartschi, (D) Anolis proboscis, (E) Anolis uniformis, (F) Anolis alvarezdeltoroi.
Anoles, however, are distributed much more widely than the Greater Antilles, their range encompassing all of the West Indies, Central America (except parts of Mexico), the northern half of South America, and the southeastern United States. For reasons both biological (anoles have lower abundance and are more cryptic on the mainland) and historical [the pioneer in anole studies, Ernest Williams, focused his work on island species (15)], the diversity of mainland anoles (Fig. 1 D–F) has received much less attention. This geographical discrepancy in research effort has occurred even though ecological and morphological diversity of mainland anoles rivals that of the islands (1, 16–19), local mainland communities support as many as 11 to 15 sympatric anole species (14), and more described species occur on the mainland (204 species) than on islands (166 species—counts excluding 9 species that secondarily colonized islands; SI Appendix, Fig. S1) (20).
A curious quirk of mainland anole diversity is that it is the result of two, partially overlapping evolutionary radiations. Anolis originated in South America ∼51 Ma (Bayesian credible interval of crown age = 42.4 to 61.7 Ma; dates from Poe et al. (see Fig. 2 legend), diversifying there while that continent was isolated from Central America (20, 21) (we henceforth refer to this clade of mainland anoles as M1; Fig. 2 A and B). A colonization event ∼43 to 51 Ma then gave rise to the anole faunas of the Greater Antilles and northern Lesser Antilles (referred to herein as GA; Fig. 2B) (20). In contrast to the diverse radiation of the GA anoles, a second clade colonized the islands of the southern Lesser Antilles (herein SLA) but did not undergo extensive diversification, no doubt a result of the small size of those islands (22). Approximately 35 Ma, an anole from the Greater Antilles colonized previously anole-free Central America, seeding an expansive species radiation (M2) in which anoles diversified and dispersed throughout the region. This second mainland radiation eventually invaded South America, where species came into contact with members of the older, incumbent mainland radiation (Fig. 2B and ref. 20).
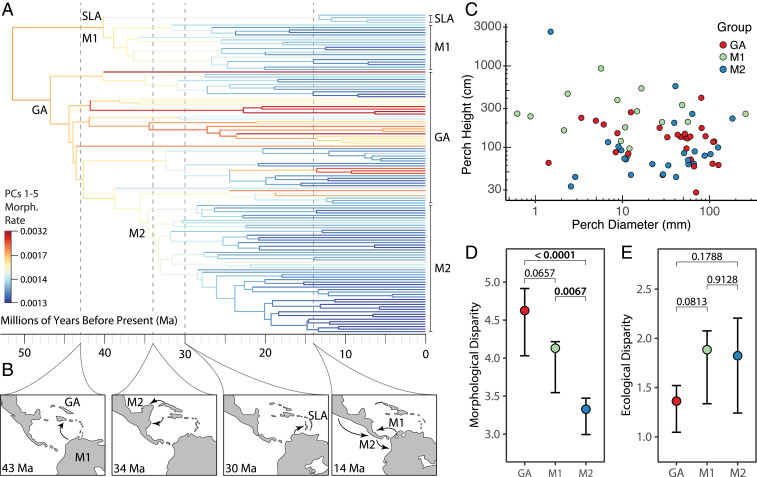
Fig. 2.
Phylogeny, paleogeographic colonization history, and morphological and ecological disparity across Anolis. (A) Morphological rates of evolution across Anolis using PCs 1 to 5 as inferred by BAMM and a time-calibrated phylogeny (20). Warmer colors indicate faster rates (see inset legend and note the nonlinear scale). (B) Colonization history of Anolis, with paleogeographic reconstructions drawn from Scotese et al. (81) for reference time points. From left to right: Colonization of northern Lesser and Greater Antilles producing the radiation we refer to as GA (42.4 to 61.7 Ma); the clade ancestrally occupying South America we refer to as Mainland 1 (M1); colonization of Central America either from Cuba or Jamaica giving rise to the clade we refer to as Mainland 2 (M2: 29.9 to 41 Ma); colonization of the southern Lesser Antilles by a clade referred to herein as SLA (23.9 to 40.1 Ma); repeated southward dispersal by M2 beginning ∼15 Ma, and the earliest instance of limited northward dispersal by M1 ∼14 Ma. (C) Perch height plotted against perch diameter, each in log-scale. (D and E) Contemporary disparity calculated as the average Euclidean distance among all pairs of points, using (D) morphological or (E) ecological traits among GA, M1, and M2. Observed values are large, filled circles; 95% CIs calculated from 1,000 bootstrap replicates are plotted as error bars. P values correspond to the probability that the difference in disparity among groups equals zero, calculated from the bootstrap replicates.
Herein, we characterize the evolutionary outcomes of this mainland recolonization and subsequent radiation by ancestrally Caribbean Anolis lizards. Specifically, we use a nearly complete time-calibrated Anolis phylogeny (20) and measurements for 10 adaptively relevant morphological and two ecological traits to address the following questions. First, how do the dynamics of adaptive radiation differ between island and mainland groups? For instance, the pace of lineage and morphological diversification is commonly thought to be greatest early in adaptive radiation (1, 23); these patterns have been observed in anoles of the Greater Antilles (24), but does the tempo and mode of diversification differ in mainland radiations? Second, does a clade recolonizing the mainland adapt in ways more similar to the earlier diverging mainland clade or to their more immediate island ancestors? For example, perhaps environmental conditions are so distinct between island and mainland settings that recolonization of the mainland by island anoles leads to the reversion to ecomorphologies more typical of the earlier diverging mainland clade. The approaches used to address the previous two questions ultimately enable us to investigate our third and primary question: What happens when closely related but independently evolving adaptive radiations collide? Do the clades exclude each other from their ancestral ranges? Or is success in radiation and dispersal asymmetrical, the outcome favoring one clade over the other?
Results and Discussion
Alternative Paths to Adaptive Radiation.
Greater Antillean anoles are an iconic example of adaptive radiation, renowned for the extent of their adaptive disparity (Fig. 1 A–C). The suggestion (25) that adaptive radiation is primarily an island phenomenon leads to the expectation that island radiations should exhibit much greater ecological and morphological disparity than their mainland counterparts. Our data for anoles reveals support for this hypothesis to be equivocal. Contrary to the ecological opportunity hypothesis, mainland anoles exhibit substantially greater variation in habitat use. They do so by occupying microhabitats little utilized by island anoles high in the trees and low on the ground (SI Appendix, Fig. S2), as well as by occurring in macrohabitats, such as cloud forests and beaches, rarely frequented on islands (14).
All three radiations exhibit substantial ecomorphological diversity, including species large and small, stout and thin, with long legs and short legs, and big toepads or none at all. Nonetheless, despite their great ecological amplitude (Fig. 2C), mainland anoles exhibit less morphological disparity than their island counterparts (Figs. 1 and 2 D and E). These results are consistent with the findings of two recent papers (26, 27).
Although all three radiations exhibit great ecomorphological variety, we find that their morphological diversity has been the result of two different evolutionary paths. This dichotomy, however, does not fall along island–mainland lines; rather, it is the two mainland anoles that differ in evolutionary trajectories. Specifically, a pattern often seen in adaptive radiations, in which morphological evolution occurs rapidly early on before slowing as niche space fills (1, 28), is seen in GA and, to a lesser extent, M1 (Fig. 1A). M2, however, does not show such an early burst. Rather, the clade has consistently low rates of morphological evolution throughout its history (these low rates are also reported by ref. 26). Nonetheless, M2 still exhibits nearly three-quarters as much ecomorphological disparity as GA and 88% of M1 (Figs. 1 D–F and 2 D and E) despite being substantially younger (indeed, when scaled by divergence time, the ecomorphological disparity of the radiations are statistically indistinguishable; SI Appendix, Fig. S3).
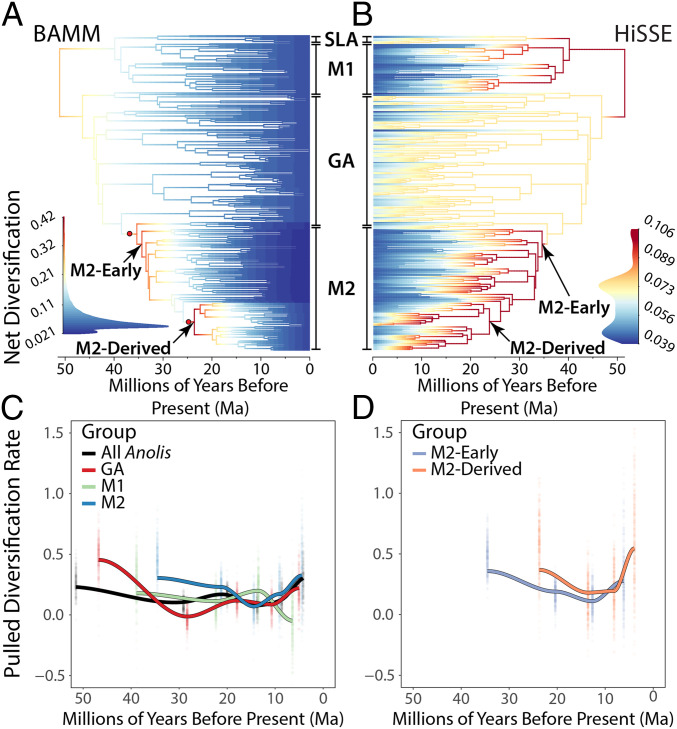
The answer to the paradox of M2’s great disparity despite low rates of morphological evolution lies in patterns of species proliferation. Although adaptive radiations often experience high rates of species diversification (i.e., net diversification: speciation − extinction) early in their histories (29), we found strong evidence of such elevated early rates only for M2. Specifically, using three different approaches (BAMM [Bayesian Analysis of Macroevolutionary Mixtures], HiSSE [Hidden State Speciation and Extinction], and pulled diversification rate [PDR]; details in SI Appendix), we failed to find consistent support for high rates of species diversification early in the radiations of GA and M1 (in each case, only one method of the three supported this pattern). In contrast, all three analyses revealed elevated species diversification rates early on for M2 followed by subsequent slowdowns [Fig. 3 A–C; see SI Appendix for discussion of comparison to other recent comparisons of anole diversification rate (27, 30)].
Fig. 3.
Species diversification dynamics of Anolis through time. Branch colors correspond to net diversification rates (see inset density plots) and red circles (in A) denote the position of rate shifts. (A) Best shift configuration according to BAMM. M2-Early denotes the early arising group of M2 (exclusive of M2-Derived; see below) inferred to experience the first round of accelerated diversification; M2-Derived is nested within M2 and was identified by BAMM as having a diversification rate increase at its base. (B) Model averaged net-diversification rate estimates obtained via HiSSE analysis. (C and D) PDR is a summary statistic of a “congruence class” of distinct, but statistically indistinguishable, diversification rate histories compatible with a single extant time tree that is equal to the net diversification rate assuming time-constant rates of speciation. Solid lines are the maximum likelihood estimate of the PDR for each discrete point in the time grid; estimates from parametric bootstrap replicates are plotted as individual points. (C) PDR for all Anolis (black) as well as the three anole radiations. PDR for SLA is not estimated due to limitations of sample size, nor for recent (i.e., < 5 Ma) timescales so as to avoid the “pull of the recent” (see Materials and Methods). (D) PDR trajectories for the two subgroups within M2.
These results point to two fundamentally different pathways to adaptive radiation: one through high rates of phenotypic evolution early in the radiation, as revealed by GA and, to a lesser extent, M1, and the other through high rates of species diversification early on without markedly high rates of concurrent morphological evolution, as evident in M2. Quantitative analysis of ongoing evolutionary rates supports this dichotomy, demonstrating that an inverse relationship exists among extant species in contemporary rates of diversification and morphological evolution (SI Appendix, Fig. S6). Despite these markedly different routes to radiation, all three clades exhibited extensive adaptive radiation (Fig. 2 C–E). Why M2 diversified in a fundamentally different manner than its island ancestors or the other mainland clade is a question ripe for future research endeavors.
Island versus Mainland Differences in Ecomorphological Adaptation.
The environment of Caribbean islands differs from that of the mainland Neotropics in many ways, including the overall greater biodiversity of the mainland, the greater prevalence of predators and insectivorous competitors [including many more squamate species (31, 32)], differences in forest structure, and even the incidence of hurricanes (16, 33–35). Given these differences, we might expect the species evolved in the mainland anole radiations to be more similar to each other than to those in an island radiation. Alternatively, because M2 is derived from GA, we might instead expect ecomorphologies to be more similar for these two groups than either is to M1.
In support of the former hypothesis, patterns of ecomorphological evolution in the two mainland radiations are similar in many respects. Not only is the magnitude of their ecological disparity statistically indistinguishable (Fig. 2E), but the two radiations overlap in morphological space to a much greater extent than either does with GA anoles when looking across all phylogenetic principal components (PCs) (Fig. 4A and SI Appendix, Fig. S7 and Table S1). Given that M2 is descended from GA, this similarity between M1 and M2 suggests a radiation-level convergence in which M2 has evolved away from its island ancestors, diversifying instead in morphospace proximate to the basal mainland clade.
Fig. 4.
Patterns and rates of morphological evolution across Anolis. (A) Two-dimensional hypervolumes of PCs 1 and 2 (24 and 22.4% of variance explained, respectively) for GA, M1, and M2 show that overlap is greatest between the two mainland clades. Hypervolumes determined using single-value vector machine (SVM) learning. Heavy, white outlined circles are centroids of each group’s hypervolume. Dark, opaque colored points are observed, and light, semitransparent points are uniformly distributed points that fall within the hypervolume inferred by the SVM. Morphospace overlap is greatest between M1 and M2 (Jaccard = 0.29, Sørenson = 0.22), followed by M2 and GA (Jaccard = 0.12, Sørenson = 0.21), and M1 and GA (Jaccard = 0.10, Sørenson = 0.19). (B–E) Ecomorphological relationships between (B) forefoot lamella count and perch height, (C) hindfoot lamella count and perch height, (D) forelimb length and perch diameter, and (E) hindlimb length and perch diameter. In B–E, variables are size-corrected, and regressions account for phylogenetic nonindependence (78). Slopes accounting for phylogenetic nonindependence are solid, and uncorrected are dashed. For each trait–habitat use comparison we conducted an ANCOVA in which we first tested for heterogeneity of slopes among GA, M1, and M2 and, when that was not significant, then tested for heterogeneity of intercepts. If we did not recover evidence for heterogeneity of intercepts (i.e., 4E) we report the P value for a single, global slope, but plot slopes per group for consistency. (F) Phylogenetic pattern of extreme convergence among species. Each convergent pair is indicated with a single line, drawn from left to right.
To further test the similarity of the mainland radiations, we examined the relationship between morphology (limb dimensions and toepad scales) and habitat use (perch height and diameter) for traits considered key to the anole adaptive radiation (14). The actual ecomorphological relationships do not neatly favor either hypothesis. For all four traits, the slope of the relationship is most different for the two mainland groups (Fig. 4 B–E and SI Appendix, Tables S2–S4); indeed, for the relationship between lamella number and perch height, the relationships are in opposite directions for the two mainland clades! For the same pairwise comparison to be the most different of the three possibilities in all four comparisons is highly improbable (P = 0.037). Clearly, ecomorphological adaptation is occurring in different ways in the two groups. Nonetheless, M2 has diverged from ancestral GA to be more similar to M1 in ecomorphological space (SI Appendix, Fig. S8). That is, M2 species are closer in trait space to M1 than they are to GA, despite the markedly different slopes of their phenotype–environment relationships.
Despite their similarities, the two mainland clades are not identical in ecological morphology. Rather, they occupy statistically distinct, albeit closely situated, regions of morphological space. Detailed examination indicates that M1 anoles differ from M2 anoles in multiple dimensions, both morphologically and ecologically (Fig. 5 and SI Appendix, Figs. S2 and S9).
Fig. 5.
Incumbent and invading mainland anoles differ in morphology and ecology. Boxplots show medians and interquartile range. P values for pairwise t tests are reported above each comparison. Significant P values are bolded. M1 anoles are typically larger than M2 species (A, t = 5.2, degrees of freedom [df] = 63, Cohen’s D = 1.03) and have shorter fore- (B, t = −2.09, df = 53, D = 0.43) and hindlimbs (C, t = −3.3, df = 53, D = 0. 68) and narrower (D, t = −6.1, df = 100, D = 1.1) and deeper (E, t = −2.04, df = 67, D = 0.41) heads. M1 is also significantly more arboreal (F, t = 3.7, df = 39, D = 1.1).
The similarity between the two mainland clades in morphology suggests that convergent evolution should be common. To quantify degree of convergence, we calculated the phylogenetically standardized trait similarity (PSTS) for each pair of species and identified the top 1% (i.e., the most similar pairs of species compared to expected similarity for each pair based on phylogenetic relationship and multivariate rates of trait evolution (Eq. 1; see SI Appendix for details). Thus, we are focusing solely on the most extreme cases of convergence. Indeed, we find that there are significantly more convergences between M1 and M2 than would be expected given their diversity alone (Fig. 4F and SI Appendix, Figs. S10–S13 and Table S5).
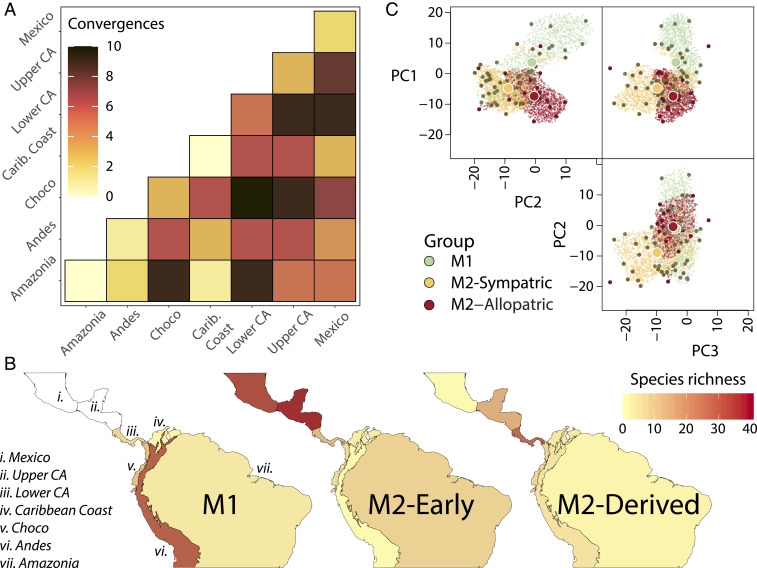
The ecomorphs of the Greater Antilles are famous examples of convergence, with a similar suite of habitat specialist ecomorphs having evolved repeatedly on each island, a pattern that has only recently been shown for mainland anoles (18, 36, 37). Surprisingly, GA anoles do not harbor more extreme convergences than expected in contrast to M2 anoles, which are significantly enriched for within-clade convergences: GA harbors 1.14 times more within-radiation convergences than expected under a neutral model of trait evolution (q = 0.923), whereas M2 harbors 2.00 times more convergences than expected (q = 0.003; Fig. 4F and SI Appendix, Figs. S10–S13 and Table S5). Most of the instances of convergence on the mainland are in species pairs occurring in different biogeographic regions in Central and South America (Fig. 6A), suggesting repeated exploration of similar adaptive space across regions. However, these mainland convergences are not organized into geographically discrete replicate adaptive radiations, as in the Greater Antilles.
Fig. 6.
Patterns of convergence, species richness, and morphological evolution across mainland Anolis. (A) Number of mainland convergences among or within each of the seven biogeographic regions. Darker colors (see inset legend) correspond to greater numbers of convergences. (B) Distribution of species richness across seven biogeographical areas in southern Mexico and Central and South America. (C) Three-dimensional hypervolumes of M1 and M2 when geographically overlapping with M1, and M2 when in allopatry with respect to M1 determined using single-value vector machine (SVM) learning and phylogenetic PC axes, plotted as in Fig. 3A. Morphospace overlap/similarity is greatest between M1 and M2–Sympatric (Jaccard = 0.24, Sørenson = 0.38), followed by M2–Allopatric & M2–Sympatric (Jaccard = 0.11, Sørenson = 0.20) and M1 and M2-Allopatric (Jaccard = 0.11, Sørenson = 0.19).
Other studies have detected convergence between mainland and island anoles (18, 37); although we detect such mainland–island convergences, we do not find that their number is exceptional. Moreover, we focused on the most extreme cases of convergence, which similarly may explain why we did not find more convergence than expected among GA anoles: Members of the same ecomorph class in the Greater Antilles are convergent but oftentimes still quite distinctive, as occurs, for example, in twig ecomorphs.
Though displaying extensive morphological disparity like GA (Fig. 2D), the adaptive radiation of the M1 clade in South America followed a different route, resulting in very little within-clade convergence (q = 0.11; Fig. 4F and SI Appendix, Figs. S10–S13 and Table S5). In this clade, ecomorphologically similar species tend to be closely related but occur in different biogeographic regions, suggesting an early diversification that quickly expanded geographically, filling ecological niches and precluding the opportunity for convergent adaptation (SI Appendix, Fig. S14).
In summary, an island–mainland difference is apparent in patterns of ecomorphological adaptation: The two mainland clades have evolved many similarities in comparison to island anoles. Nonetheless, differences exist between the mainland clades; patterns of convergent evolution reveal, just as with the rates of species and morphological diversification, that the two mainland clades have taken different routes to similar outcomes.
When Radiations Collide.
The two mainland anole clades were allopatric over much of their evolutionary history, separated by the Central American Seaway. However, as the Seaway began to close with the rise of the Isthmus of Panama, M1 and M2 came into contact (Figs. 2B and 6B). This contact between two large anole clades, part of the Great American Biotic Interchange, represents a rare example of closely related and adaptively diverse clades coming into contact (38).
Given the general similarity in the pattern of diversification of the two clades, one might expect a priority effect in which neither clade was able to make inroads into the area occupied by the other (39, 40). Alternatively, the differences that existed between clades in ecological morphology may have allowed the clades to partition resources and thus coexist in sympatry, possibly leading to character displacement to enhance these differences.
We find support for neither of these possibilities. Contrary to the possibility that priority effects prevented range expansions, both clades were able to expand into the ancestral area of the other (Fig. 2B), producing communities comprised of members of both M1 and M2 (18). However, as with the famous mammal story (41, 42), the Great Anole Biotic Interchange was asymmetric in favor of the northerners; anoles from Central America swept into South America, occupying the entire ancestral range of M1 (Fig. 6B). In contrast, South American M1 anoles had considerably less success moving northward [much less so than South American mammals (41, 42) and other taxa (43, 44)], advancing only to southern Costa Rica, thus failing to occupy much of the geographic area inhabited by M2.
Also similar to the mammal invasion, in situ diversification played an important role. Diversification rate analyses revealed that a subclade of M2 (herein M2-Derived) experienced elevated rates of diversification beginning ∼24 Ma, concurrent with the initial stages of the formation of the Isthmus of Panama [estimated ∼25 Ma (45); Fig. 3 A and D]. Members of this clade subsequently expanded southward (as did some members of M2-Early), invading South America as early as 15 Ma (20), coincident with extensive uplift 15 to 10 Ma (46) that subsequently led to the closure of the Central American Seaway (47). A result of this heightened diversification is that M2 constitutes one-third of the 103 anole species in South America; of these 17 fall within M2-Early and 18 within M2-Derived. By contrast, the M1 clade experienced no such increase in diversification rate at the time of the interchange; rather, it declined after 14 Ma. Moreover, relatively few M1 species occur in Central America (9 species), substantially fewer than the 41 M2 species in the same area (lower Central America) and only 9% of the anole species in all of Central America.
The asymmetry in invasion success is also evident in ecology and morphology. Morphologically, M2 anoles that cooccur with M1 species are very similar to allopatric M2 anoles (Fig. 6C). That is, 90.4% of the morphospace occupied by South American M2 species is shared by M2 species in Central America (SI Appendix, Fig. S15 and Table S6). Although some of this morphological variation came with the immigrants from Central America, in situ evolutionary diversification played a role as well, as evidenced by the extensive convergence between South and Central American M2 (Fig. 6A). In contrast, M1 anoles in Central America are less varied morphologically than they are in their ancestral range in South America, primarily being restricted to large species (SI Appendix, Fig. S16A). A null model that samples with replacement from the ancestral range found that the size distribution of Central American species was strongly nonrandom relative to the ancestral species pool in South America (P = 0.007; SI Appendix, Fig. S16B). These results indicate that random colonization of species could not produce such a large discrepancy in size distribution. More sophisticated analyses incorporating phylogenetic information and in situ diversification dynamics are needed to more fully investigate whether this difference could have arisen by chance. Ecologically, an interesting difference is apparent; not only do M1 anoles perch higher than M2 anoles (P = 0.00071; Fig. 5F), but this difference is exaggerated in sympatry; M2 anoles in sympatry with M1 anoles perch lower than M2 anoles in allopatry (P = 0.0025; SI Appendix, Fig. S17A).
This ecological niche partitioning could set the stage for the evolution of character displacement in sympatry, but we found little evidence for its occurrence (SI Appendix, Fig. S18). That is, sympatric M2 anoles are closer in morphospace to M1 anoles than are allopatric M2 species, contrary to the character displacement hypothesis (Fig. 6C and SI Appendix, Table S7). Indeed, rather than diverging, sympatric M2 anoles were often more similar to M1 than were allopatric M2 anoles when looking at individual traits (SI Appendix, Fig. S18).
In sum, sympatry with M1 had little effect on the ecological or morphological diversification of M2. However, the fact that M1’s expansion into areas occupied by M2 was only accomplished by large species—those most dissimilar in size to M2 anoles—suggests a role for interspecific interactions in constraining the expansion of M1 (SI Appendix, Fig. S16).
At present, we have no explanation for this discrepancy in outcome. When biotas meet, the direction of asymmetric interchange often is predicted by the extent of available habitat in each region; species from the larger region tend to overrun those from the smaller region (38). However, our results contradict this expectation given the much greater size of M1’s range in South America compared to M2’s range in Central America. Alternatively, successful invaders often come from more species-rich areas and M2’s diversity in their ancestral area (excluding M2 in South America: 101 species) is substantially greater than M1 in theirs (excluding M1 in Central America: 67 species). If the faunal exchange was simply a function of clade size, then we would expect proportionally similar numbers of species exchanged between the two regions. In contrast, we find that 12% of M1’s diversity is found outside of its ancestral range in South America as compared to M2, for which 27% of its diversity is found outside of its ancestral distribution in Central America.
Unfortunately, the population biology and ecology of mainland anoles is surprisingly poorly known, and new species are being described at a steady pace (18, 36, 48–53). Perhaps future work will uncover key life history, physiological or behavioral differences between the clades that can explain their differential success. If so, perhaps we can learn whether those differences evolved as a result of historical environmental differences between Central and South America, as a legacy of M2’s island ancestry (island anoles occur at much higher densities and experience strong intra- and interspecific competition), or for other reasons.
Regardless, the anole adaptive radiations deliver two important messages. First, islands, far from being evolutionary dead ends, are a cauldron of evolutionary innovation and diversification. Not only are island species capable of colonizing the mainland, but they can give rise to highly successful evolutionary radiations, so evolutionarily capable that they outperform mainland radiations when they come into contact. Our results emphasizes that mainland diversity is ripe for future study, both to gain a more complete understanding of the process of adaptive radiation on mainland settings and to better characterize the understudied extant mainland diversity (53).
Second, multiple paths exist to adaptive radiation. Evolutionary biologists have debated the relative importance of rapid ecomorphological differentiation and rapid species diversification as keys to adaptive radiation (1, 2, 29). The anole story illustrates that no single answer exists; rather, similar adaptive radiations can result from very different evolutionary histories.
More generally, although mainland and island radiations have diversified to produce very different ecomorphological configurations, a parallel exists in patterns of diversification across clades within each region. On the four islands of the Greater Antilles, essentially the same set of ecomorphs has evolved on each island, yet the sequence by which they have evolved has not been the same across islands (54). In a similar way, our results here show that the two mainland radiations have taken different routes to very similar outcomes. This parallel duality illustrates a higher-level convergence in the interplay of determinism and contingency, occurring not only within clades but across regions and radiations.
Materials and Methods
Phylogenetic and Geographic Data Collection.
To test the hypothesis that geographic factors influence lineage diversification in Anolis, we used the time-calibrated molecular phylogeny of Poe et al. (20) including 315 of 379 extant Anolis lizards (83% of described species). Each species was classified as being either a mainland (0) or island (1) species according to the data matrix of Poe et al. (20). Two species with widespread distributions in Cuba have colonized the coast of Central America; they are considered island species. In addition to the four anole radiations, nine mainland species colonized very small islands: Anolis agassizi, Anolis bicaorum, Anolis concolor, Anolis lineatus, Anolis nelsoni, Anolis pinchoti, Anolis roatensis, Anolis townsendi, and Anolis utilensis. Not surprisingly, given the size of the islands, none of these species have diversified. Because the purpose of these analyses is to compare patterns of diversification between island and mainland radiations, we have excluded them from the study.
We extracted the M1, M2, and GA subtrees (see main text) for downstream analyses (Fig. 1C) using APE v5.0 (55) in R (56). Due to its small size and restricted geographic distribution, SLA was not included in most analyses. In several cases, subgroups of these trees were also examined. When testing models of trait-dependent diversification, we treat island/mainland as a binary trait that is hypothesized to impact the diversification process. To contextualize our findings in the paleogeographic history of southern Mexico and Central and South America, we visualized paleogeographic rasters in GPlates (57).
Morphological and Habitat Use Data Collection.
To investigate patterns and rates of morphological evolution, we measured 10 traits on adult males that have been shown to be functionally relevant to the diversification of Anolis (14). Morphological measurements for all but lamella counts were collected between 2009 and 2019 from 188 of the 379 extant species of Anolis, totaling 1,965 specimens. Traits measured were snout vent length (SVL), head depth, head width, head length, tail length, fore-limb length, hind-limb length, and counts of forefoot (toe III) and hindfoot (toe IV) lamellae. Additional details are available in SI Appendix. Lamella counts were collected from a total of 946 preserved specimens across 230 species with the goal of sampling five individuals where possible. Details on the collection of these data can be found in SI Appendix.
To test whether relationships between morphology and ecology in Anolis are consistent between the mainland and island species, we collected data on perch height and perch diameter for 1,555 males across 72 species. Data on habitat use were collected by walking through the habitat at daytime and by measuring, for each undisturbed individual, its height above the ground and the diameter of its perch using a tape measure or a laser distance meter.
To account for the confounding effect of correlations between body size and all other morphological and habitat measurements, we calculated the residuals of each Ln-transformed variable against Ln(SVL) using a standard linear regression. In comparisons of univariate traits, we included data from species not included in the phylogeny; these analyses are conducted without incorporating phylogenetic information. All other analyses utilize data only from species included in the phylogeny and incorporate phylogenetic information. Specifically, to improve statistical power, we applied univariate analyses (ANOVAs and Welch’s two-sample t tests of individual traits as in Fig. 5) to all species, including those not in the time-calibrated phylogeny. These tests included the comparisons among 1) M1/M2/GA, 2) M1/M2-Sympatric/M2-Allopatric, 3) M1/M2-South America/M2-elsewhere, and 4) among M1 and M2 of different biogeographic regions on the mainland (i.e., South America, lower Central America, upper Central America, and Mexico). All other morphological analyses are conducted in a phylogenetic context and so use only those species shared between both datasets.
Rates of Morphological Evolution.
To test whether rates of morphological diversification were driven by patterns of colonization from either the mainland to the Antilles or vice versa we used BAMM (58). Although use of the fossil record can inform estimates of historical rates and patterns of morphological evolution (59), the pre–late-Pleistocene fossil record of anoles is scant (60). Consequently, our analyses are limited to the use of contemporary samples. These analyses were repeated across all 10 traits and for measurements collected from males. As this method cannot handle missing data, species for which data were unavailable for any of the 10 traits were excluded. On average, only 51% of the species in our time-calibrated phylogeny were sampled for morphological traits. Consequently, we conservatively specified our prior on the number of rate shifts as two, or less than half that used in the diversification rate analysis (five). Assuming a prior of two rate shifts, we ran six MC3 chains for 100 million generations, sampling every 5,000 generations and discarding the first 20% of generations as burn-in. As each analysis was conducted on individual traits [Ln(SVL) and size-corrected residuals for other traits], a total of 20 analyses were conducted. We used the plotPrior function in BAMMtools (61) to confirm that our prior on number of rate shifts did not influence our posterior. We also investigated the rate of multivariate trait evolution by analyzing phylogenetic PCs 1 to 5 in the same manner as described above. We report these results individually, but additionally summarized these results by combining (postburn-in) the posterior distribution of shifts and associated rates by appending the resultant Markov chain Monte Carlos (MCMCs). We evenly sampled 25,000 total events (5,000 per PC) across this combined posterior and plotted the resulting rates along the phylogeny. Because samples were drawn evenly along the combined posterior, rates estimated for each PC is represented equally in the final sampled posterior.
Trait Disparity.
To obtain estimates of contemporary disparity we used the R package dispRity (62). Only species that were measured for all 10 morphological and two ecological traits were included for this analysis. To standardize unit measurements, we scaled all 10 traits to mean = 0, SD = 1. Disparity is calculated as the average Euclidean distance in phenotypic trait values between species and was obtained for each subtree (i.e., M1, M2, and GA). We tested two hypotheses: 1) that disparity differs among mainland (M1 and M2 combined) and GA anoles and 2) that disparity differs among the three major clades. To quantify uncertainty in our estimates of contemporary diversity we conducted 1,000 nonparametric bootstraps. From these bootstrapped distributions we calculated 95% confidence intervals. To test whether contemporary diversity significantly differed between groups we tested the hypothesis that differences in disparity among bootstrapped distributions equals zero. To test the hypothesis that each group differs in the rate at which disparity accumulates we repeated the above analyses, but scaling disparity by patristic distance separating each species pair.
Geography-Dependent Diversification.
To test the hypothesis that mainland and island Anolis experienced different diversification rates we competed seven models of trait-dependent and seven models of trait-independent diversification using HiSSE (63). HiSSE tests for associations between diversification rates and binary characters while allowing for heterogeneity in diversification rates within character-states. As data for this analysis, we used the full time-calibrated molecular phylogeny with species designated as either mainland (0) or island (1). Parameter optimization was improved through the use of simulated annealing to first traverse the likelihood surface to identify ideal starting parameter values for subsequent maximum likelihood optimization.
To correct for the influence of incomplete sampling on diversification rate parameter estimation (64) we specified the sampling fraction as the proportion of extant described mainland (82%) and island (85%) species sampled in our phylogeny (see SI Appendix). For both this analysis and the analysis described below, we focused primarily on net diversification, which we acknowledge will primarily be driven by the estimation of speciation rates. That is, although extinction may well play a significant role in the diversification process, our limited ability to accurately estimate extinction rates will hamper meaningful interpretation of this parameter (65).
Lineage Diversification Rate Shifts.
To complement the above analyses we tested whether diversification rates vary in a geographically concordant manner among anole lineages by analyzing the complete phylogeny using BAMM v2.5 (61, 66). BAMM tests for the presence of distinct diversification rate regimes within a phylogeny in a manner agnostic to geography. Reversible-jump MCMC is used to sample and pinpoint the locations of diversification rate shifts, and diversification rate parameters are fit to the subtending branches (66). Assuming a prior of five rate shifts, we ran four independent runs of six MC3 chains for a total of 50 million generations, sampling every 5,000 generations, discarding the first 20% of generations for burn-in. Clade specific sampling fractions were specified each for clades M1 (62%), M2 (91%), GA (84%), and SLA (89%) (Fig. 1C). We subsequently used the Gelman diagnostic (67) to confirm that the four runs had converged on the same posterior distribution and the Geweke diagnostic to confirm that each run had converged using the R package coda (68). Further visual confirmation of convergence was assessed in coda. Because all runs converged, we present results from only one of these runs. As with HiSSE, all species were included.
PDR Estimation.
Louca and Pennell (69) recently demonstrated that extant time trees alone cannot reliably estimate speciation and extinction rates separately using homogeneous birth–death models and instead are consistent with a large number of diversification histories (for a contrasting view see ref. 70). That is, each time tree is consistent with a set of different, but statistically indistinguishable, diversification dynamics called a “congruence class.” One summary statistic, the PDR, summarizes the congruence class; the PDR is equal to the net diversification rate when the speciation rate is constant through time. In effect, the PDR is analogous to the net diversification rate—positive values imply an increase in diversity through time, whereas negative values imply a loss of diversity through time. To characterize the diversification dynamics of anoles within the new framework, we estimated the PDR for each Anolis group (GA, M1, and M2 and also for mainland groups M2-Early and M2-Derived). For each subtree, we 1) used a custom script (provided by Matthew Pennell) to identify the ideal number of time-points needed to estimate the PDR, 2) fit the PDR using maximum likelihood, 3) simulated 1,000 phylogenies of equal size and age under the fitted parameters, and 4) fit the PDR to each simulated tree. In effect, steps 3 to 4 represent a parametric bootstrapping procedure. For each subtree, rho, or the proportion of extant species included in the phylogeny, was specified. In turn, we obtain time-series estimates of the PDR, which is equal to the net diversification rate assuming time-constant speciation rates and incomplete taxon sampling. We excluded estimates more recent than 5 Ma so as to avoid “pull of the recent” effects (i.e., upwardly biased estimate of diversification caused by failure to identify young, morphologically cryptic taxa as distinct species).
Morphospace Comparison.
We asked whether patterns of morphospace occupation differed among mainland and island clades. We first make this comparison among GA, M1, and M2. We further ask whether M2 exhibits ecological character displacement when in sympatry with M1. If so, we predict that M2 species allopatric from M1 will be more similar in morphospace to M1 than M2 species that are sympatric with M1. To test this prediction we 1) conducted a phylogenetic principal component analysis (71) accounting for shared phylogenetic history with phytools v0.6-99 (72) using the correlation matrix (because we had both ordinal and continuous traits) and 2) constructed n-dimensional hypervolumes using a one-class support vector machine (SVM) learning model using the first three PC axes (73). As with previous analyses, only species measured for all 10 traits were included in these analyses. Hypervolumes are visualized by randomly sampling 1,000 uniformly distributed points that are determined by the SVM to fall within the inferred hypervolume. These random (unobserved) points are in turn plotted as small, semitransparent points. These hypervolumes were subsequently used to calculate distance and similarity/overlap between GA, M1, and M2 in multivariate space. These morphospace and hypervolume analyses were repeated for M2 either in sympatry or allopatry with M1. Hypervolumes were subsequently summarized using four statistics: Jaccard’s similarity, Sørenson’s similarity, and fraction of hypervolume unique for either of the two groups in the comparison. To supplement our multivariate approach, we conducted simple pairwise comparisons (t test assuming unequal variance) of morphological and ecological data between 1) all three major geographic clades and 2) M1 and M2, either when in sympatry or allopatry.
Quantifying the Extent of Convergence.
To quantify the extent of extreme convergence within and among anole groups, we identified species pairs that are much more similar in morphology than would be expected under a fitted model of trait evolution, accounting for phylogeny. Specifically, we replicated the procedure of Mazel et al. (74) but inverted the values so as to provide a more intuitive metric of convergence. In brief, we quantified the extent to which observed trait similarities deviated from expectation under a model of neutral trait evolution by calculating the standard effect size. This metric is given by Eq. 1:
| [1] |
This procedure allows for the identification of both the most extreme observed convergences (positive PSTS) and divergences (negative PSTS); however, we focus solely on convergence. Note that this approach simply allows for the identification of the most extremely similar species pairs given a particular sample of species; in this sense, we are quantifying relative, not absolute, convergence. That is, this metric cannot be compared across distinct datasets, nor does it indicate that unidentified pairs aren’t convergent to a lesser extent.
As with previous multivariate analyses, only species measured for all 10 traits were included in this analysis. As our distance metric we used Euclidean distances. We tested two models of trait evolution: multivariate Brownian motion and multivariate Ornstein–Uhlenbeck, each fitted using mvMORPH v1.1.0 (75). Under the best-fit model we then simulated 20,000 comparable multivariate trait distributions in mvMORPH to obtain a null distribution of trait similarities. We subsequently used these 20,000 simulated trait distributions to calculate PSTS for each pairwise comparison. We define the extreme convergences as the top 1% of the pairwise PSTS values. To minimize potential instances of phylogenetic stasis being mistakenly identified as convergence, we excluded any convergence between sister species as determined using the complete phylogeny using the distTips function in adephylo (76). However, this procedure only led to the removal of two species pairs: Anolis dolichocephalus–Anolis hendersoni and Anolis lionotus–Anolis poecilopus.
To compare the extent of extreme convergence in different groups we depict these extreme convergences as a heat map in which for each cell the number of observed extreme convergences is scaled relative to the number expected given the number of pairwise comparisons. A binomial test is used to examine whether extent of extreme convergence differed significantly among comparisons, with P values corrected for multiple tests using the method of Benjamini and Hochberg (77). To further explore patterns of extreme convergence on the mainland we calculated the number of extreme convergences observed among species occurring within each of the seven biogeographic regions. Results for M2 alone are qualitatively very similar and thus not shown.
Ecological Morphology of Mainland and Island Anolis.
To test the hypothesis that the relationship between morphology and ecology differs among groups, we conducted a series of phylogenetic regressions of morphological traits against habitat use measurements. Specifically, we tested for relationships between 1) hind- and forefoot lamella count and perch height and 2) hind- and forelimb length against perch diameter. To account for phylogenetic nonindependence we conducted phylogenetic least squares (PGLS) in the R package caper v1.0.1 (78). For each trait–habitat use comparison, we conducted an analysis of covariance (ANCOVA) in which we first tested for heterogeneity of slopes among GA, M1, and M2 and, when that was not significant, then tested for differences in intercepts. If neither was significant, we report P values for the global slope. The procedure used follows that of Fuentes-G et al. (79). To test the hypothesis that M2 species are closer in ecomorphological space to M1 than to GA, we calculated the distance from each M2 species to the fitted slopes for M1 and GA for all four ecomorphological relationships. To quantify similarity of slope between M1, M2, and GA, we fit all possible models including two of the three groups in which trait value is predicted by ecological value plus an interaction term with clade. From these models, we recoded the slope per group, calculated the difference between these slopes (a metric of how similar the slopes are), and extracted the P value for the interaction term. We also tested whether perch height or perch diameter differed between clades (GA, M1, and M2) or mainland groups (M1, M2-Allopatric, and M2-Sympatric) using an ANOVA. When the ANOVAs were significant, we made pairwise comparisons using Welch’s two-sample t tests. For relationships in which a significant interaction term was recovered (i.e., slopes differed among clades) we sought to quantify how similar these relationships were between groups. Thus, we subsequently repeated the above procedure using all pairwise combinations of clades. Thus, for each pairwise comparison, we tested whether intercepts or slopes differed significantly (SI Appendix, Table S2).
Acknowledgments
For comments on earlier drafts we thank M. Alfaro, F. Burbrink, M. Muñoz, E. Burress, L. Mahler, D. Reznick, and R. Ricklefs. L. Mahler contributed to the collection of the lamella data. For assistance in the field, logistics, advice, or in other ways, we thank M. Aja, C. Anderson, N. Aristizabal, T. Aveling, F. Ayala, T. Barros, E. Bell, O. Bent, S. Bessudo, E. Boada, N. Bolaños, H. Botero, M. Calderón, R. Camacho, S. Campbell-Staton, S. Challenger, J. Christensen, C. Crandell, M. DaCosta Cottam, J. Daltry, E. Díaz Franjul, M. Díaz Franjul, T. Eldred, P. Endera, A.-C. Fabre, K. Fenstermacher, T. Fies, K. Godbeer, G. Gómez, R. González, D. Irschick, S. Janzan, C. Jiménez, V. Jimenez Arcos, D. Kizirian, N. Lawrence, K. Lindsay, S. Lobos, D. Losos, M. Losos, G. Londoño, E. López, L. Mahler, J. Mailles, C. Mariën, C. M. Gomez, A. Marmol, J. Meyers, M. Muñoz, S. Montuelle, R. Moreno, N. Murillo, J. O’Reilly, A. Otto, A. Pacheco, L.F. Payán, M. Pennell, S. Poe, S. Quintero, G. Rivas, A. Rodrıguez, J. Rosado, G. Saborío, E. Salamanca, D. Salazar, C. Santoro, M. Sasa, E. Schaad, D. Schluter, S. Steele, J. Stops, J. Tayton, B. Vanhooydonck, P. Vanmiddlesworth, J. Velasco, P. Velozo, L. Vonnahme, S. Walton, J. Weaver, and K. Wollenberg. We thank the following institutions for assistance: American Museum of Natural History; Departamento de Biología, Facultad Experimental de Ciencias, Universidad del Zulia; Discovery Bay Marine Laboratory; Estación de Biología Tropical Los Tuxtlas; Museum of Comparative Zoology, Harvard University; Organization for Tropical Studies; Smithsonian Tropical Research Institute; Pine Cay Homeowners Association; and Redonda Restoration Programme. Research was conducted under permits from Autoridad Nacional del Ambiente de Panama; Autoridad Nacional de Licencias Ambientales, Colombia; Ministerio del Ambiente y Energía–Sistema Nacional de Areas de Conservacion, Costa Rica; CARMABI Foundation and the government of Curaçao; Department of Agriculture, Bahamas; Department of Environment Government of Antigua and Barbuda; Direction de l’Environnement de l’Aménagement et du Logement de la Martinique; Dominican Ministry of Agriculture and Fisheries Forestry, Wildlife and Parks Division; and Turks and Caicos Department of Environment and Coastal Resources. This work was supported by the NSF (NSF IOS-1354620 to J.B.L. and NSF OAC-1835893 to L.J.H.), the David M. Fite Fund, Putnam Expedition Grants from the Museum of Comparative Zoology, and the David Rockefeller Center for Latin American Studies, Harvard University. A.H.P. was supported by the NSF Postdoctoral Research Fellowship in Biology under grant DBI-2109838.
Footnotes
The authors declare no competing interest.
See QnAs, e2116186118, in vol. 118, issue 42.
This article contains supporting information online at https://www.pnas.org/lookup/suppl/doi:10.1073/pnas.2024451118/-/DCSupplemental.
Data Availability
Scripts and ecological and morphological measurements have been deposited in GitHub (https://github.com/austinhpatton/AnolisRadiation) (80). Ecological, morphological, and all other study data are included in the article and/or supporting information.
References
- 1.Schluter D., The Ecology of Adaptive Radiation (Oxford University Press, Oxford, 2000). [Google Scholar]
- 2.Gillespie R. G., et al., Comparing adaptive radiations across space, time, and taxa. J. Hered. 111, 1–20 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Stroud J. T., Losos J. B., Ecological opportunity and adaptive radiation. Annu. Rev. Ecol. Evol. Syst. 47, 507–532 (2016). [Google Scholar]
- 4.Wilson E. O., The nature of the taxon cycle in the Melanesian ant fauna. Am. Nat. 95, 169–193 (1961). [Google Scholar]
- 5.MacArthur R. H., Wilson E. O., Island Biogeography (Princeton University Press, 1967). [Google Scholar]
- 6.Kalmar A., Currie D. J., A global model of island biogeography. Glob. Ecol. Biogeogr. 15, 72–81 (2006). [Google Scholar]
- 7.Elton C. S., The Ecology of Invasions by Animals and Plants (Springer, 1958). [Google Scholar]
- 8.Filardi C. E., Moyle R. G., Single origin of a pan-Pacific bird group and upstream colonization of Australasia. Nature 438, 216–219 (2005). [DOI] [PubMed] [Google Scholar]
- 9.Bellemain E., Ricklefs R. E., Are islands the end of the colonization road? Trends Ecol. Evol. 23, 461–468 (2008). [DOI] [PubMed] [Google Scholar]
- 10.Fabre P.-H., et al., Dynamic colonization exchanges between continents and islands drive diversification in paradise-flycatchers (Terpsiphone, Monarchidae). J. Biogeogr. 39, 1900–1918 (2012). [Google Scholar]
- 11.Nishiumi I., Kim C.-H., Assessing the potential for reverse colonization among Japanese birds by mining DNA barcode data. J. Ornithol. 156, 325–331 (2015). [Google Scholar]
- 12.Patiño J., et al., Approximate Bayesian Computation reveals the crucial role of oceanic islands for the assembly of continental biodiversity. Syst. Biol. 64, 579–589 (2015). [DOI] [PubMed] [Google Scholar]
- 13.Rowe K. C., et al., Oceanic islands of Wallacea as a source for dispersal and diversification of murine rodents. J. Biogeogr. 46, 2752–2768 (2019). [Google Scholar]
- 14.Losos J. B., Lizards in an Evolutionary Tree: Ecology and Adaptive Radiation of Anoles (University of California Press, 2009). [Google Scholar]
- 15.Williams E. E., “The origin of faunas. Evolution of lizard congeners in a complex island fauna: A trial analysis” in Evolutionary Biology, Dobzhansky T., Hecht M. K., Steere W. C., Eds. (Springer, 1972), pp. 47–89. [Google Scholar]
- 16.Irschick D. J., Vitt L. J., Zani P. A., Losos J. B., A comparison of evolutionary radiations in mainland and Caribbean Anolis lizards. Ecology 78, 2191–2203 (1997). [Google Scholar]
- 17.Pinto G., Mahler D. L., Harmon L. J., Losos J. B., Testing the island effect in adaptive radiation: Rates and patterns of morphological diversification in Caribbean and mainland Anolis lizards. Proc. Biol. Sci. 275, 2749–2757 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Poe S., Anderson C. G., The existence and evolution of morphotypes in Anolis lizards: Coexistence patterns, not adaptive radiations, distinguish mainland and island faunas. PeerJ 6, e6040 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Moreno-Arias R. A., Calderón-Espinosa M. L., Patterns of morphological diversification of mainland Anolis lizards from northwestern South America. Zool. J. Linn. Soc. 176, 632–647 (2016). [Google Scholar]
- 20.Poe S., et al., A phylogenetic, biogeographic, and taxonomic study of all extant species of Anolis (Squamata; Iguanidae). Syst. Biol. 66, 663–697 (2017). [DOI] [PubMed] [Google Scholar]
- 21.Nicholson K. E., et al., Mainland colonization by island lizards. J. Biogeogr. 32, 929–938 (2005). [Google Scholar]
- 22.Losos J. B., Schluter D., Analysis of an evolutionary species-area relationship. Nature 408, 847–850 (2000). [DOI] [PubMed] [Google Scholar]
- 23.Simpson G. G., The Major Features of Evolution (Columbia University Press, 1953). [Google Scholar]
- 24.Mahler D. L., Revell L. J., Glor R. E., Losos J. B., Ecological opportunity and the rate of morphological evolution in the diversification of Greater Antillean anoles. Evolution 64, 2731–2745 (2010). [DOI] [PubMed] [Google Scholar]
- 25.Webb C. O., Ackerly D. D., McPeek M. A., Donoghue M. J., Phylogenies and community ecology. Annu. Rev. Ecol. Syst. 33, 475–505 (2002). [Google Scholar]
- 26.Feiner N., Jackson I. S. C., Stanley E. L., Uller T., Evolution of the locomotor skeleton in Anolis lizards reflects the interplay between ecological opportunity and phylogenetic inertia. Nat. Commun. 12, 1–10 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Burress E. D., Muñoz M. M., Ecological opportunity from innovation, not islands, drove the anole lizard adaptive radiation. Syst. Biol., 10.1093/sysbio/syab031 (2021). [DOI] [PubMed] [Google Scholar]
- 28.Simpson G. G., Tempo and Mode in Evolution (Columbia University Press, 1944). [Google Scholar]
- 29.Givnish T. J., Adaptive radiation versus ‘radiation’ and ‘explosive diversification’: Why conceptual distinctions are fundamental to understanding evolution. New Phytol. 207, 297–303 (2015). [DOI] [PubMed] [Google Scholar]
- 30.Poe S., et al., Comparative evolution of an archetypal adaptive radiation: Innovation and opportunity in Anolis lizards. Am. Nat. 191, E185–E194 (2018). [DOI] [PubMed] [Google Scholar]
- 31.Vitt L. J., De la Torre S., Guía para la investigación de las lagartijas de Cuyabeno (Museo de Zoologia Centro de Biodiversidad, ed. 1, 1996). [Google Scholar]
- 32.Savage J., The Amphibians and Reptiles of Costa Rica: A Herpetofauna Between Two Continents, Between Two Seas (University of Chicago Press, 2002). [Google Scholar]
- 33.Donihue C. M., et al., Hurricane effects on Neotropical lizards span geographic and phylogenetic scales. Proc. Natl. Acad. Sci. U.S.A. 117, 10429–10434 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Green H. W., “Species richness in tropical predators” in Tropical Rainforests: Diversity and Conservation, Almeda F., Pringle C. M., Eds. (California Academy of Sciences and Pacific Division, American Association for the Advancement of Science, 1988), pp. 259–280. [Google Scholar]
- 35.Simard M., Pinto N., Fisher J. B., Baccini A., Mapping forest canopy height globally with spaceborne lidar. J. Geophys. Res. Biogeosci. 116, G04021 (2011). [Google Scholar]
- 36.Schaad E. W., Poe S., Patterns of ecomorphological convergence among mainland and island Anolis lizards. Biol. J. Linn. Soc. Lond. 101, 852–859 (2010). [Google Scholar]
- 37.Huie J. M., Prates I., Bell R. C., de Queiroz K., Convergent patterns of adaptive radiation between island and mainland Anolis lizards. Biol. J. Linn. Soc. Lond. 134, 85–110 (2021). [Google Scholar]
- 38.Vermeij G. J., When biotas meet: Understanding biotic interchange. Science 253, 1099–1104 (1991). [DOI] [PubMed] [Google Scholar]
- 39.Carlquist S. J., Island Biology (Columbia University Press, 1974). [Google Scholar]
- 40.Fukami T., Historical contingency in community assembly: integrating niches, species pools, and priority effects. Annu. Rev. Ecol. Evol. Syst. 46, 1–23 (2015). [Google Scholar]
- 41.Webb S. D., Mammalian faunal dynamics of the great American interchange. Paleobiology 2, 220–234 (1976). [Google Scholar]
- 42.Marshall L. G., Land mammals and the great American interchange. Am. Sci. 76, 380–388 (1988). [Google Scholar]
- 43.Leigh E. G., O’Dea A., Vermeij G. J., Historical biogeography of the Isthmus of Panama. Biol. Rev. Camb. Philos. Soc. 89, 148–172 (2014). [DOI] [PubMed] [Google Scholar]
- 44.Weir J. T., Bermingham E., Schluter D., The Great American Biotic Interchange in birds. Proc. Natl. Acad. Sci. U.S.A. 106, 21737–21742 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Farris D. W., et al., Fracturing of the Panamanian Isthmus during initial collision with: South America. Geology 39, 1007–1010 (2011). [Google Scholar]
- 46.Montes C., et al., Evidence for middle Eocene and younger land emergence in central Panama: Implications for Isthmus closure. Geol. Soc. Am. Bull. 124, 780–799 (2012). [Google Scholar]
- 47.Montes C., et al., Middle Miocene closure of the Central American seaway. Science 348, 226–229 (2015). [DOI] [PubMed] [Google Scholar]
- 48.Poe S., Ryan M. J., Description of two new species similar to Anolis insignis (Squamata: Iguanidae) and resurrection of Anolis (Diaphoranolis) brooksi. Amphib. Reptile Conserv. 11, 1–16 (2017). [Google Scholar]
- 49.Poe S., Scarpetta S., Schaad E. W., A new species of Anolis (Squamata: Iguanidae) from Panama. Amphib. Reptile Conserv. 9, 1–13 (2015). [Google Scholar]
- 50.Velasco J. A., Herrel A., Ecomorphology of Anolis lizards of the Choco′ region in Colombia and comparisons with Greater Antillean ecomorphs. Biol. J. Linn. Soc. Lond. 92, 29–39 (2007). [Google Scholar]
- 51.Velasco J. A., Poe S., González-Salazar C., Flores-Villela O., Solitary ecology as a phenomenon extending beyond insular systems: Exaptive evolution in Anolis lizards. Biol. Lett. 15, 20190056 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Velasco J. A., et al., Climatic and evolutionary factors shaping geographical gradients of species richness in Anolis lizards. Biol. J. Linn. Soc. Lond. 123, 615–627 (2018). [Google Scholar]
- 53.Moura M. R., Jetz W., Shortfalls and opportunities in terrestrial vertebrate species discovery. Nat. Ecol. Evol. 5, 631–639 (2021). [DOI] [PubMed] [Google Scholar]
- 54.Losos J. B., Jackman T. R., Larson A., Queiroz K., Rodríguez-Schettino L., Contingency and determinism in replicated adaptive radiations of island lizards. Science 279, 2115–2118 (1998). [DOI] [PubMed] [Google Scholar]
- 55.Paradis E., Claude J., Strimmer K., APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004). [DOI] [PubMed] [Google Scholar]
- 56.R Core Team , R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2019). [Google Scholar]
- 57.Müller R. D., et al., GPlates: Building a virtual earth through deep time. Geochem. Geophys. Geosyst. 19, 2243–2261 (2018). [Google Scholar]
- 58.Rabosky D. L., Donnellan S. C., Grundler M., Lovette I. J., Analysis and visualization of complex macroevolutionary dynamics: An example from Australian scincid lizards. Syst. Biol. 63, 610–627 (2014). [DOI] [PubMed] [Google Scholar]
- 59.Simões T. R., Vernygora O., Caldwell M. W., Pierce S. E., Megaevolutionary dynamics and the timing of evolutionary innovation in reptiles. Nat. Commun. 11, 1–14 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Sherratt E., et al., Amber fossils demonstrate deep-time stability of Caribbean lizard communities. Proc. Natl. Acad. Sci. U.S.A. 112, 9961–9966 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Rabosky D. L., et al., BAMM tools: An R package for the analysis of evolutionary dynamics on phylogenetic trees. Methods Ecol. Evol. 5, 701–707 (2014). [Google Scholar]
- 62.Guillerme T., dispRity: A modular R package for measuring disparity. Methods Ecol. Evol. 9, 1755–1763 (2018). [Google Scholar]
- 63.Beaulieu J. M., O’Meara B. C., Detecting hidden diversification shifts in models of trait dependent speciation and extinction. Syst. Biol. 65, 583–601 (2016). [DOI] [PubMed] [Google Scholar]
- 64.Stadler T., How can we improve accuracy of macroevolutionary rate estimates? Syst. Biol. 62, 321–329 (2013). [DOI] [PubMed] [Google Scholar]
- 65.Louca S., Pennell M. W., Why extinction estimates from extant phylogenies are so often zero. Curr. Biol. 31, 3168–3173.e4 (2021). [DOI] [PubMed] [Google Scholar]
- 66.Rabosky D. L., Automatic detection of key innovations, rate shifts, and diversity-dependence on phylogenetic trees. PLoS One 9, e89543 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Gelman A., Rubin D. B., Inference from iterative simulation using multiple sequences. Stat. Sci. 7, 457–472 (1992). [Google Scholar]
- 68.Plummer M., Best N., Cowles K., Vines K., CODA: Convergence diagnosis and output analysis for MCMC. R News 6, 7–11 (2006). [Google Scholar]
- 69.Louca S., Pennell M. W., Extant timetrees are consistent with a myriad of diversification histories. Nature 580, 502–505 (2020). [DOI] [PubMed] [Google Scholar]
- 70.Morlon H., Hartig F., Robin S., Prior hypotheses or regularization allow inference of diversification histories from extant timetrees. bioRxiv [Preprint] (2020). https://www.biorxiv.org/content/10.1101/2020.07.03.185074v1 (Accessed 7 July 2020).
- 71.Revell L. J., Size-correction and principal components for interspecific comparative studies. Evolution 63, 3258–3268 (2009). [DOI] [PubMed] [Google Scholar]
- 72.Revell L. J., phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2012). [Google Scholar]
- 73.Blonder B., et al., New approaches for delineating n‐dimensional hypervolumes. Methods Ecol. Evol. 9, 305–319 (2018). [Google Scholar]
- 74.Mazel F., et al., The geography of ecological niche evolution in mammals. Curr. Biol. 27, 1369–1374 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Clavel J., Escarguel G., Merceron G., mvMORPH: An R package for fitting multivariate evolutionary models to morphometric data. Methods Ecol. Evol. 6, 1311–1319 (2015). [Google Scholar]
- 76.Jombart T., Balloux F., Dray S., adephylo: New tools for investigating the phylogenetic signal in biological traits. Bioinformatics 26, 1907–1909 (2010). [DOI] [PubMed] [Google Scholar]
- 77.Benjamini Y., Hochberg Y., Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995). [Google Scholar]
- 78.Orme D., The caper package: Comparative analyses of phylogenetics and evolution in R (2012).
- 79.Fuentes-G J. A., Housworth E. A., Weber A., Martins E. P., Phylogenetic ANCOVA: Estimating changes in evolutionary rates as well as relationships between traits. Am. Nat. 188, 615–627 (2016). [DOI] [PubMed] [Google Scholar]
- 80.Patton A. H., AnolisRadiation. GitHub. https://github.com/austinhpatton/AnolisRadiation. Deposited 13 September 2021.
- 81.Scotese C. R., Wright N., PALEOMAP Paleodigital Elevation Models (PaleoDEMS) for the Phanerozoic PALEOMAP Project. https://www.earthbyte.org/paleodem-resource-scotese-and-wright-2018/. Accessed 26 May 2019.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Scripts and ecological and morphological measurements have been deposited in GitHub (https://github.com/austinhpatton/AnolisRadiation) (80). Ecological, morphological, and all other study data are included in the article and/or supporting information.