Figure 1.
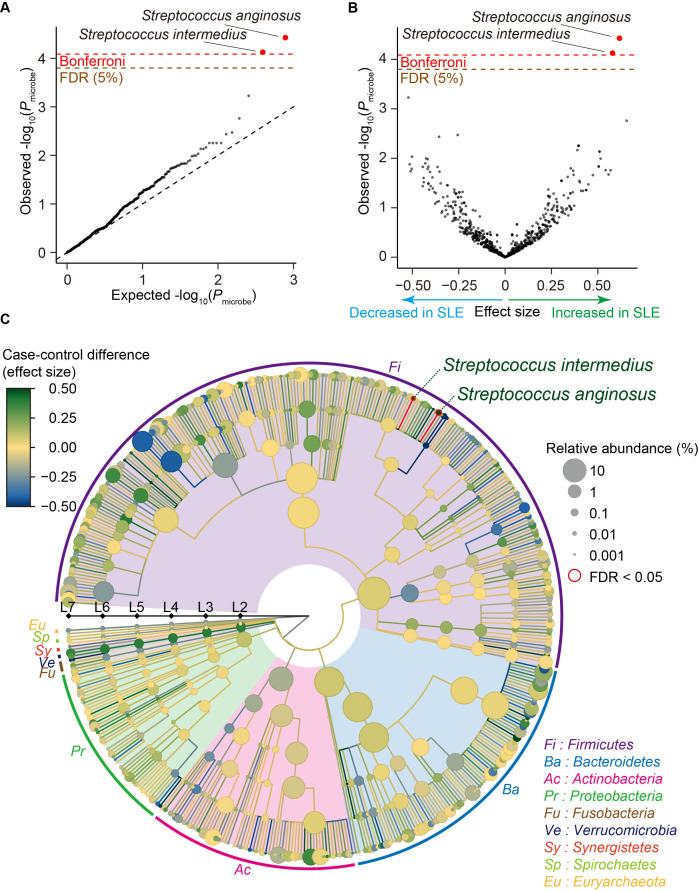
Result of the SLE MWAS based on the phylogenetic abundance data. (A) A quantile-quantile plot of the phylogenetic MWAS p values (P microbe) of the clades. The x-axis indicates log-transformed empirically estimated median P microbe. The y-axis indicates observed −log10(P microbe). The diagonal dashed line represents y=x, which corresponds to the null hypothesis. The horizontal red line indicates the empirical Bonferroni-corrected threshold (α=0.05), and the brown line indicates the empirically estimated FDR threshold (FDR=0.05). Clades with P microbe less than the Bonferroni thresholds are plotted as red dots, and other clades are plotted as black dots. (B) A volcano plot. The x-axis indicates effect sizes in linear regression. The y-axis, horizontal lines and dot colours are the same as in (A). (C) A phylogenetic tree. Levels L2–L7 are from the inside layer to the outside layer. The size and the colour of dots represent relative abundances and effect sizes, respectively. The two clades with significant case-control associations (FDR<0.05) are outlined in red. FDR, false discovery ratio; MWAS, metagenome-wide association study; SLE, systemic lupus erythematosus.