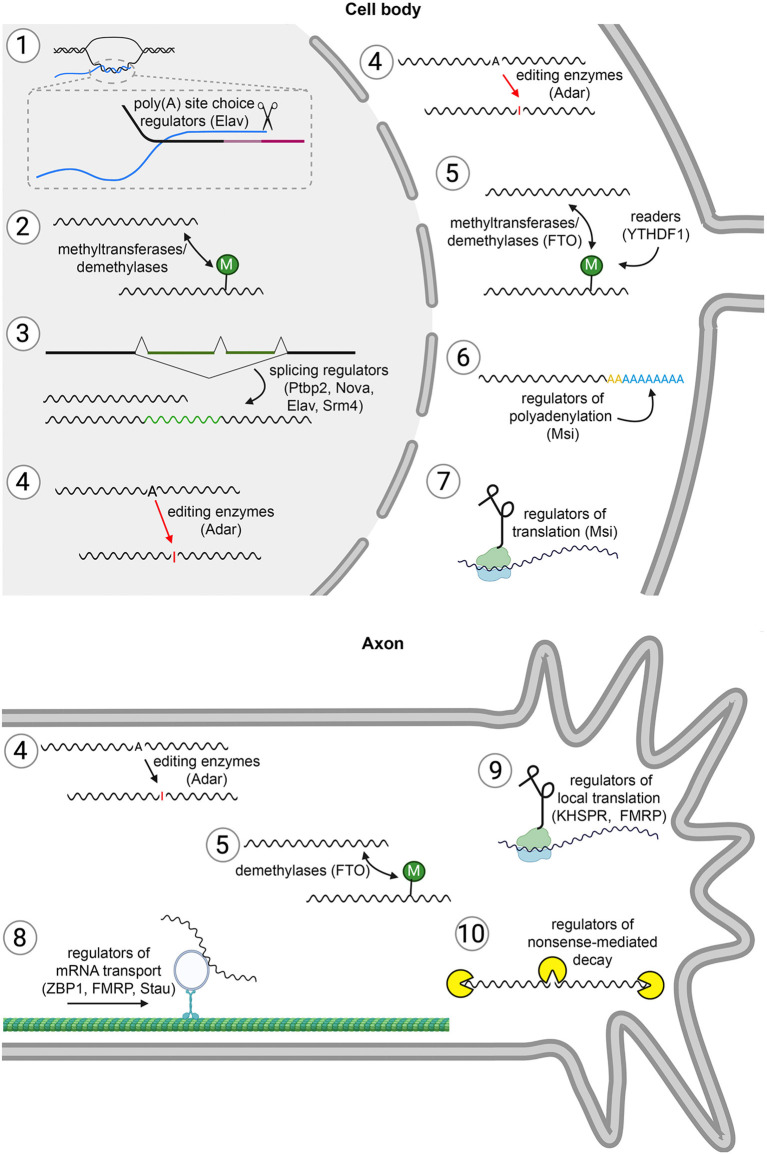
Figure 1.
Different steps of mRNA metabolism. (1) Alternative cleavage and polyadenylation. The choice between alternative polyadenylation signals diversifies 3′UTRs and thereby determines, which regulatory motifs are included in an mRNA molecule. Thus, this process expands the potential for post-transcriptional regulation of gene expression. As in the rest of the figure, an example of an RNA-binding protein that controls the process, and which we discuss in this article, is indicated in brackets. Blue: mRNA. Shades of pink: alternative 3′UTRs encoded in the DNA. (2) RNA modification. Nucleotides are modified for example by methyltransferases and demethylases. RNA modification can occur both in the nucleus and in the cytoplasm. This impacts on splicing, translation, and stability of an mRNA. (3) Alternative splicing (AS). Regulation of gene expression by AS is a means to increase proteome diversity, and also to include or exclude regulatory elements that for example can provide temporal or spatial control of expression. (4) RNA editing. The coding and regulatory regions in an mRNA molecule can be edited in the nucleus or in the cytoplasm, for example by conversion of adenosines (A) into inosines (I). (5) Dynamic RNA modifications. In the cell body and axonal cytoplasm, modifications can be added or removed from mRNA molecules. RBPs that recognize the modifications (“readers”) can then for example modulate translation. (6) Polyadenylation. This dynamic process can occur in the nucleus [yellow poly(A) tails] and in the cytoplasm [blue poly(A) tails]. Polyadenylation is a means to regulate translation and mRNA stability. (7) Translation. In neurons, protein synthesis is heavily regulated to provide temporal and spatial control of proteome composition during development. (8) Localization. mRNA molecules can be transported to axon terminals, and (9) locally translated. (10) Degradation. Different pathways, such as nonsense-mediated decay, degrade mRNAs after translation.