Table 1.
Data collection, phasing and refinement statistics
| RRV SeM-RdRpSER3 | RRV RdRpSER3 | SINV RdRp | SINV RdRpCS | |
|---|---|---|---|---|
| Data collection | Se-SAD phasing | Native | Native | Native |
| PDB ID | - | 7F0S | 7VB4 | 7VW5 |
| Space group | P 21 21 21 | P 21 21 21 | P 1 | P 1 |
| Unit cell dimensions | ||||
| a, b, c (Å) | 64.15, 68.58, 100.98 | 64.84, 68.35, 99.65 | 67, 68.29, 71.07 | 67.45, 68.94, 71.42 |
| α, β, γ (°) | 90, 90, 90 | 90, 90, 90 | 115.97, 106.71, 95.90 | 116.311, 107.05, 95.10 |
| Wavelength | 0.979 (peak) | 0.999 (native) | 1.000 | 0.946 |
| Resolution range (Å) | 46.85–2.83 (2.93–2.83) | 23.41–2.6 (2.69–2.6) | 40.52–1.9 (1.968–1.9) | 34.48–2.3 (2.382–2.3) |
| Total reflections | 296 954 (29 212) | 148 510 (15 410) | 524 533 (36 101) | 161 646 (17 056) |
| Unique reflections | 11 129 (1107) | 14 118 (1399) | 78 572 (6646) | 45 591 (4603) |
| Mean I/sigma(I) | 17.19 (0.30) | 23.81 (1.78) | 18.84 (1.90) | 13.25 (3.46) |
| Completeness (%) | 99.93 (99.82) | 99.66 (100.00) | 92.26 (71.47) | 94.41 (93.72) |
| Multiplicity | 26.7 (26.3) | 10.5 (11.0) | 6.7 (5.4) | 3.5 (3.7) |
| R merge b | - | 0.066 (1.469) | 0.06941 (0.9519) | 0.05755 (0.3683) |
| R meas c | - | 0.070 (1.544) | 0.07531 (1.054) | 0.06833 (0.4318) |
| R pim d | - | 0.023 (0.470) | 0.02882 (0.4429) | 0.03631 (0.2233) |
| CC1/2 e | - | 0.984 (0.632) | 0.999 (0.708) | 0.998 (0.901) |
| FOM | 0.70 (0.66) | - | - | - |
| Phase error (°) | 35.60 (40.21) | - | - | - |
| Refinement Statistics | ||||
| Reflections used in refinement | - | 14 112 (1399) | 78 574 (5923) | 45 652 (4415) |
| Reflections used for R-free | - | 696 (67) | 1858 (146) | 1169 (112) |
| Rwork f | - | 0.265 (0.3877) | 0.1875 (0.3210) | 0.1950 (0.2564) |
| Rfree g | - | 0.288 (0.4466) | 0.2239 (0.3549) | 0.2446 (0.3374) |
| Number of nonhydrogen atoms | - | 3109 | 8064 | 7648 |
| macromolecules | - | 3108 | 7160 | 7267 |
| ligands | - | - | 28 | 4 |
| solvent (water) | - | 1 | 876 | 377 |
| Protein residues | - | 419 | 917 | 930 |
| RMSD (bonds) | - | 0.006 | 0.003 | 0.002 |
| RMSD (angles) | - | 0.99 | 0.58 | 0.53 |
| Ramachandran statistics | ||||
| Ramachandran favored (%) | - | 91.81 | 97.45 | 97.6 |
| Ramachandran allowed (%) | - | 8.19 | 2.44 | 2.4 |
| Ramachandran outliers (%) | - | 0.00 | 0.11 | 0 |
| Rotamer outliers (%) | - | 0.00 | 0.13 | 0.25 |
| Clashscore | - | 12.18 | 3.21 | 4.69 |
| Average B-factor | - | 108.36 | 28.32 | 44.61 |
| macromolecules | - | 108.37 | 27.71 | 44.54 |
| ligands | - | - | 34.43 | 52.67 |
| solvent | - | 69.40 | 33.15 | 45.86 |
| Number of TLS groups | - | 6 | 7 | 7 |
The values presented in parentheses are of the highest resolution shell.
 .
.
 , where Rmeas is the precision indicator of individual observation for unmerged data.
, where Rmeas is the precision indicator of individual observation for unmerged data.
 , where Rpim is the precision indicator for the merged data.
, where Rpim is the precision indicator for the merged data.
 , where CC1/2 is of two half datasets.
, where CC1/2 is of two half datasets.
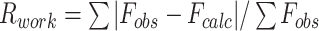 .
.
 , where Rfree was randomly sampled from 5% reflections.
, where Rfree was randomly sampled from 5% reflections.
