Fig 1. PEDAR mediates large target deletion and simultaneous insertion at an endogenous genomic locus.
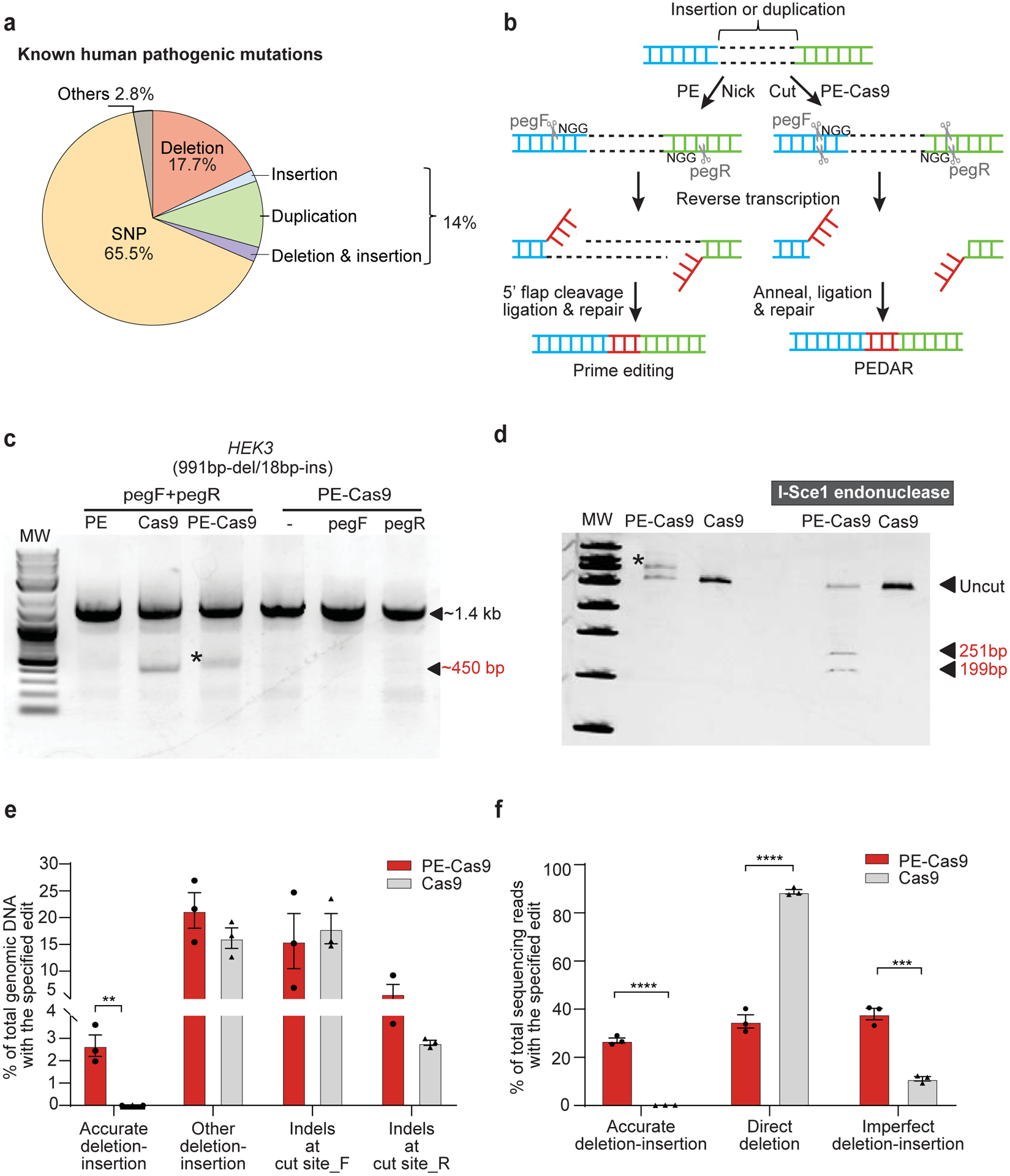
a, Classification of the 60,008 known human pathogenic genetic variants reported in the ClinVar database1. b, Overview of using prime editing (left) and PEDAR (right) to generate accurate deletion-insertion. c, Deleting a 991-bp DNA fragment and simultaneous insertion of I-Sce1 recognition sequence (18bp) at the HEK3 locus (Chr9:107422166–107423588). Target genomic region was amplified using primers that span the cut sites. The paired pegRNAs targeting complementary DNA strand are denoted as pegF and pegR. HEK293T cells were transfected with PE, Cas9, or PE-Cas9 with or without single or paired pegRNAs. The ~450-bp band is the expected deletion amplicon (denoted with *), and the ~1.4-kb band is the amplicon without deletion. d, Deletion amplicons from Cas9- or PE-Cas9-treated groups shown in (c) were incubated with or without I-SceI endonuclease and analyzed in 4–20% TBE gel. Digested products are marked by arrows with expected sizes. Original amplicon is marked as “uncut”. The band with insertion of i-Sce1 recognition sequence is denoted with *. e, PE-Cas9 or Cas9-mediated absolute rates of all the editing events in total genomic DNA at HEK3 site. Data represent mean ± SEM (n = 3 biologically independent samples). P= 0.0053 (**), two-tailed t-test. f, Deep sequencing of deletion amplicons shown in (c). Bar chart shows distribution of all deletion events, including accurate deletion-insertion, direct deletion (deletion without any insertions), or imperfect deletion-insertion. Editing rate = the reads with indicated editing / total deletion events. Data represent mean ± SEM (n = 3 biologically independent samples). P=0.0000140 (****), 0.002 (***), two-tailed t-test.
