Abstract
Background
Naked mole-rats (Heterocephalus glaber, NMRs) and blind mole-rats (Spalax galili, BMRs) are representative subterranean rodents that have evolved many extraordinary traits, including hypoxia tolerance, longevity, and cancer resistance. Although multiple candidate loci responsible for these traits have been uncovered by genomic studies, many of them are limited to functional changes to amino acid sequence and little is known about the contributions of other genetic events. To address this issue, we focused on gene losses (unitary pseudogenes) and systematically analyzed gene losses in NMRs and BMRs, aiming to elucidate the potential roles of pseudogenes in their adaptation to subterranean lifestyle.
Results
We obtained the pseudogene repertoires in NMRs and BMRs, as well as their respective aboveground relatives, guinea pigs and rats, on a genome-wide scale. As a result, 167, 139, 341, and 112 pseudogenes were identified in NMRs, BMRs, guinea pigs, and rats, respectively. Functional enrichment analysis identified 4 shared and 2 species-specific enriched functional groups (EFGs) in subterranean lineages. Notably, the pseudogenes in these EFGs might be associated with either regressive (e.g., visual system) or adaptive (e.g., altered DNA damage response) traits. In addition, several pseudogenes including TNNI3K and PDE5A might be associated with specific cardiac features observed in subterranean lineages. Interestingly, we observed 20 convergent gene losses in NMRs and BMRs. Given that the functional investigations of these genes are generally scarce, we provided functional evidence that independent loss of TRIM17 in NMRs and BMRs might be beneficial for neuronal survival under hypoxia, supporting the positive role of eliminating TRIM17 function in hypoxia adaptation. Our results also suggested that pseudogenes, together with positively selected genes, reinforced subterranean adaptations cooperatively.
Conclusions
Our study provides new insights into the molecular underpinnings of subterranean adaptations and highlights the importance of gene losses in mammalian evolution.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12915-022-01243-0.
Keywords: Gene losses, Naked mole-rat, Blind mole-rat, Subterranean adaptations, Pseudogenes
Background
Subterranean environment provides mammalian dwellers with highly stressful living conditions that are characterized mainly by extreme hypoxia/hypercapnia, darkness, and food scarcity. Over the course of their evolution for millions of years, subterranean mammals have evolved exquisite physiological and morphological modifications to cope with these challenges [1]. Excellent examples of subterranean adaptation were found in naked mole-rats (Heterocephalus glaber, NMRs) [2–8] and the blind mole-rats (Spalax galili, BMRs) [9–15]. NMRs and BMRs are phylogenetically distantly related subterranean rodents that evolved similar morphological and extraordinary physiological traits. To live in the burrow system, NMRs and BMRs evolved cylinder-shaped bodies, shortened limbs, strong claws, elongated incisors, and degenerated visual system [1]. They are highly tolerant to tissue hypoxia, resistant to oxidative stress induced by oxygen deficiency and insensitive to acid induced by hypercapnia [4, 5, 7, 11, 15]. In addition, they are both extremely long-lived rodents that live more than 20 years and are able to suppress spontaneous and experimentally induced tumorigenesis [3, 6, 14], even though cases of cancer were reported recently in zoo-housed NMRs [16]. They also evolved several species-specific traits, for example, eusociality [17], poikilothermy [18], and lack of a circadian sleep rhythm in NMRs [19]; enlarged brain structure related to sensing and orientation [20, 21], and increased muscle capillary density [22] in BMRs. These traits aroused general interests among researchers working on various fields such as evolutionary biology, genetics, aging, and cancer research. Understanding the molecular basis of extraordinary traits in NMRs and BMRs may help us address some of the most challenging questions in biology and medicine, including adaptations to extreme environments, mechanisms of hypoxia tolerance, mechanisms of aging, and cancer resistance.
Genetic and genomic studies in NMRs and BMRs have uncovered the molecular basis of some extraordinary traits. For examples, genome sequencing revealed that the amino acid substitutions in NMR UCP1 protein are associated with its unique thermoregulation, and fast evolution of BMR embryonic hemoglobin γ gene may contribute to hypoxia adaptation [4, 15]. Comparative genomics has uncovered hundreds of positively selected genes (PSGs) and convergently evolved genes that may contribute to phenotypic evolution in NMRs and BMRs [23–25], including the convergence on the SCN9A gene (encoding Nav1.7), which leads to the repeatedly evolved acid insensitivity [15, 26]. However, it should be noted that the positive selection and convergence rarely occurred in the same gene across subterranean lineages and that evolutionary changes in different genes might produce similar phenotypes [25]. In addition, transcriptome analysis identified many differentially expressed genes that are associated with longevity, hypoxia, and hypercapnia tolerance [4, 15, 27–29]. For example, two p53 target genes, CYTC and CASP9, are downregulated in BMRs under hypoxia, which may contribute to cellular protection against hypoxia-induced apoptosis [15]. SMAD3 gene is also upregulated in an aging NMR brain, which may slow down the growth of cancer cells and contribute to cancer resistance [4]. Even though these studies identified many candidate loci that are potentially responsible for certain traits, functional assays are largely absent to validate their contributions. Moreover, most of these studies were focusing on the functional modifications of protein-coding genes and many traits cannot be explained adequately. In this regard, it is important to assess the contributions of other genetic events to subterranean adaptations.
In addition to functional modifications of protein-coding genes, other types of genetic changes can contribute to phenotypic evolution, such as pseudogenes (gene losses) [30], regulatory element mutations [31], and epigenetic modifications [32]. Gene losses, which are usually associated with regressive evolution, were recently shown to play roles in adaptive phenotypic evolution more than we expected. Examples of adaptive gene losses include: (i) in cetaceans, losses of DSG4, DSC1, TGM5, and GSDMA may contribute to the evolution of an epidermis morphology suitable for aquatic environment, and loss of POLM may contribute to the improved tolerance of oxidative DNA lesions [33–36]; (ii) in fruit bats, losses of FAM3B and FFAR3 may help them adapt to sugar-rich diet by increasing insulin secretion and insulin sensitivity [33]; (iii) repeated losses of PNLIPRP1 in herbivores may increase their capacity of fat storage, which is beneficial given that herbivores usually consume fat-poor diet [37]. These observations are in consistence with the “less is more” hypothesis [38]. In terms of the subterranean mammals, however, almost all current studies addressing the role of gene losses in phenotypic evolution focused on eye degeneration [4, 15, 29, 39–41], while the contribution of gene losses to other subterranean adaptations has yet to be investigated.
In this study, to assess the contribution of gene losses to subterranean adaptations, we systematically identified gene losses in NMRs and BMRs, as well as in their terrestrial relatives, guinea pigs, and rats, which were used as aboveground controls. Comparison of gene losses between subterranean and superterranean lineages identified multiple pseudogenes that might be associated with specific subterranean physiological traits including an altered DNA damage response and a specialized cardiovascular system. In addition, we provide functional evidence for the contribution of eliminating TRIM17, a gene independently lost from NMRs and BMRs, to hypoxia adaptation. Thus, our work provides new insights into the molecular mechanisms of adaptations to subterranean lifestyle in both NMRs and BMRs.
Results
Compositions of the pseudogene repertoires
We first identified species-specific gene losses (unitary pseudogenes) in NMRs, BMRs, guinea pigs, and rats by mapping a complete mouse protein set to their genomic sequences, followed by disablement (stop codons and frameshifts) detection and filtering by rigorous criteria (Additional file 1: Fig. S1; see "Methods"). The mouse genome was used as reference because mouse is the phylogenetically closest to NMRs and BMRs with high-quality genome assembly and annotations available at present, which would mostly minimize the effects caused by phylogenetic distance. Several factors affect the validity of detected gene losses including the misjudgment of orthologous sequences, genetic redundancy [42], and the false positives introduced by misjudgment of disruptive mutations, genome sequencing/assembly errors, and annotation errors in reference genome. To address these issues and to obtain high-confidence gene losses for each species, we implemented a series of filtering steps (Additional file 1: Fig. S1; see "Methods" for details). Briefly, we first established the unitary status of each putative pseudogenic loci by (i) removing loci hit by predictively annotated proteins, intronless cDNA/expressed sequences, or genes belonging to large gene families such as olfactory receptors, zinc finger proteins, and vomeronasal receptors; (ii) removing genes with highly similar sequences at more than one locus in the genome; (iii) removing genes without conserved genomic position between query species and mouse. Secondly, we removed false disruptive mutations generated by (i) GeneWise alignment errors; (ii) the sequencing and assembly errors; (iii) potential annotation errors that resulted from the reference proteins with low transcript supporting level, or from downstream compensatory mutations that rescues the ORF disruption. As a result, 211, 211, 617, and 389 high-confidence unitary pseudogenes were identified from the NMRs, BMRs, guinea pigs, and rats, respectively. To make our results more conservative, we further removed those pseudogenes without signatures of relaxed selection (Additional file 1: Fig. S1; see "Methods") because gene dispensability and the resulting relaxed selective constraint is considered the hallmark of gene losses [30]. Finally, we obtained 167, 139, 341, and 112 gene losses in NMRs, BMRs, guinea pigs, and rats, respectively (Fig. 1, Table 1 and Additional file 2: Table S1). In order to assess the validity of our methodology, we further validated our filtering procedures with BUSCO gene sets, which are generally assumed to be conserved and universally presented in most species [43]. Using 9150 mammalian conserved BUSCO genes, we identified 20 in BMRs and 22 in NMRs as pseudogenes, suggesting ~ 0.24% putative false positive ratio, which is comparable to previous pseudogene screening studies (e.g., < 0.3% [33]). The use of 13,650 glires conserved genes showed similar results (0.30% = 41/13,650 in BMRs and 0.31% = 42/13,650 in NMRs, respectively). Thus, our pipeline produces minimal false positives.
Fig. 1.

Gene losses identified in each species. The numbers of gene losses were labelled above each branch of the phylogenetic tree of species investigated
Table 1.
Statistics of each step of identifying gene loss events in NMRs, BMRs, guinea pigs, and rats
| Species | Overlapping best hits | Putative pseudogenic loci | Unitary pseudogenic loci | False positives removed | Selective constraints relaxed |
|---|---|---|---|---|---|
| NMRs | 14,254 | 1362 | 883 | 211 | 167 |
| BMRs | 16,546 | 1609 | 1023 | 211 | 139 |
| Guinean pigs | 15,969 | 2695 | 2217 | 617a | 341 |
| Rats | 17,396 | 2821 | 1701 | 389a | 112 |
aSequencing errors were not examined
Functional bias and convergence of gene losses in subterranean lineages
Next, we examined what kinds of genes were preferentially lost in NMRs and BMRs, which are more likely to be associated with subterranean adaptations. Using mouse orthologs as surrogates, we firstly obtained all associated GO terms for all pseudogenes in each species. Functionally cross-related GO terms were clustered into functional groups, and a subset of GO terms from each group that are overrepresented in subterranean lineage pseudogenes compared to subterranean lineage non-pseudogenes or control lineage pseudogenes (NMRs vs. guinea pigs and BMRs vs. rats) were identified as an enriched functional group (EFGs) (see "Methods"). Among the 24 clustered functional groups, a total of 10 EFGs were identified, with 5 in the NMR pseudogenes and 5 in the BMR pseudogenes (Fig. 2 and Additional file 3: Table S2). For each EFG, the proportion of pseudogenes from the subterranean lineage is significantly higher than that of pseudogenes from the corresponding control lineage and/or the proportion of functional genes from its own genome (Fig. 2 and Additional file 3: Table S2), indicating the functional loss of genes in these EFGs likely to be related to the subterranean lifestyle. Genes in two EFGs might be associated with species-specific traits. For example, the EFG for “lipid metabolism” might be associated with the alternating membrane phospholipid composition in NMRs [44]. The EFG “neural processes” might be associated with the enlarged brain volume in BMRs [20, 21]. In contrast, four EFGs, namely, those for the “visual system,” “reproduction,” “DNA damage response,” and “proteolysis,” were shared between the pseudogene lists from the two subterranean species. These repeatedly observed EFGs support a notion that subterranean adaptations are relatively restricted to where certain physiological process was recurrently modified to cope with stressful environments.
Fig. 2.
Functional groups enrichment analysis. Lineage-specific and shared EFGs identified in NMR (A) and BMR (B) pseudogenes. The proportions of pseudogenes and functional genes background from each species in each EFG are shown and proportional differences in each EFG were tested with Fisher’s exact test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001
The hypoxic and hypercapnic burrow environments cause a significant oxidative stress, which can cause DNA damage, especially DNA double-stranded breaks (DSBs). We observed significant enrichments of NMR and BMR pseudogenes in “DNA damage response” EFG when compared to their own functional gene background (FDR corrected P = 0.0081 for NMRs and P = 0.0024 for BMRs, Fisher’s exact test) (Additional file 3: Table S2), while guinea pig and rat pseudogenes show no significance compared to their respective functional gene background (Additional file 3: Table S2). The genes in this EFG involve DNA damage checkpoint and DNA repair regulation, including D7ERTD443E, NUDT16L1, and EME2 in NMRs and NEK11, D330045A20RIK, and EME2 in BMRs (Fig. 3 and Additional file 3: Table S2 and Additional file 4: Table S3). Among them, D7ERTD443E (FATS) and NEK11 are checkpoint related genes that affect cell cycles and apoptosis by regulating p53/p21 and CDC25A, respectively [45–48]. NUDT16L1(TIRR) and D330045A20RIK (RADX) are negative DNA repair regulators [49, 50]. EME2 is an endonuclease component gene that is capable of causing DNA damage under replication stress [51–53] (Fig. 3). Thus, the functional loss of these DNA damage response (DDR)-related genes in NMRs and BMRs might enhance DNA repair and reduce stress-induced apoptosis, which is in consistence with the similar direction of alteration of DNA damage response observed in NMRs and BMRs [8, 54, 55]. Therefore, these convergent DDR-related gene losses are probably adaptive in response to stressful subterranean environments.
Fig. 3.
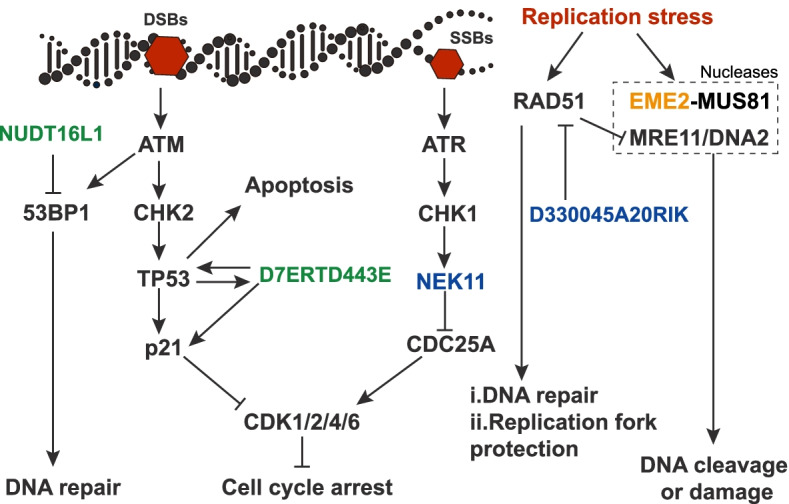
Functional convergences of gene losses between NMRs and BMRs in DDR pathway. A part of DNA damage response pathway is represented. Genes highlighted in green and blue are gene losses from NMRs and BMRs, respectively, and the gene highlighted in orange indicates shared gene loss between NMRs and BMRs
Interestingly, we also noticed that some pseudogenes in a functional group named “cardiovascular system” also show a similar pattern of functional convergence, although no significant enrichments were observed (Additional file 3: Table S2). For example, genes involved in blood pressure and heart rate regulation were detected as pseudogenes in NMRs (TNNI3K [56] and RNPEP) and BMRs (PDE5A) (Additional file 4: Table S3). Inhibition of RNPEP was shown to suppress hypertension in spontaneously hypertensive rats [57], and inhibition of TNNI3K and PDE5A was shown to be associated with suppressed contractility [58, 59]. Even though the cardiomyocyte contractility in BMRs has yet to be determined, the heart rate of BMRs was observed to be lower than rat under both normoxia and hypoxia conditions [60], which might be caused by weakened cardiac contractility. Therefore, functional convergence among these pseudogenes might be associated with the low blood pressure in NMRs and low heart rate in BMRs that caused by changes in cardiac contraction [22, 60–62]. In addition, inhibition of TNNI3K or PDE5A was shown to be cardioprotective against several cardiomyopathies [63–67]. Moreover, TNNI3K inhibition was also shown to increase the frequency of the mononuclear diploid cardiomyocyte (MNDCM) population and to elevate cardiomyocyte proliferation after injury [56, 68]. Taken together, the association of distinct pseudogenes with convergent adaptive traits highly suggests that gene losses could play a positive role in subterranean adaptations.
Potential roles of convergent gene losses (CGLs) in subterranean adaptations
It has been shown recently that CGLs contribute to repeatedly evolved adaptations [33, 37, 69]. To reveal subterranean adaptations that were contributed by CGLs, we compared the pseudogene lists from NMRs and BMRs. Twenty CGLs were observed between NMRs and BMRs (Additional file 4: Table S3 and Additional file 5: Table S4). The hypergeometric test shows that these CGLs significantly outnumbered the random expectation (1.42, P = 1.25 × 10−17, Fig. 4A). To examine the extent to which the number of CGLs is affected by similarity of ecological niches, we further paired the four species according to the similarities and differences in their living environments. Among 4 pairs of species, NMRs-BMRs (N-B) and guinea pigs-rats (G-R) live in similar environment (homotypic pair), while NMRs-rats (N-R) and BMRs-guinea pigs (B-G) live in different environment (heterotypic pair). Comparison of pseudogene lists between paired species shows that homotypic N-B pair is the most prominent with the highest number of CGLs (Fig. 4B). In contrast, the number of CGLs from homotypic G-R pair is significantly lower than N-B pair (5/453 vs. 20/306, P < 0.0001, Fisher’s exact test) (Fig. 4B). The CGLs number from the homotypic N-B pair is significantly higher than that from heterotypic B-G pair (20/306 vs. 7/480, P = 0.0002), while there is no significant difference between the CGLs number from the homotypic G-R pair and that from the heterotypic N-R pair (Fig. 4B). In line with the functional convergence among distinct genes, the CGLs between NMRs and BMRs directly reflect their similar responses to subterranean environments and likely contributed to the evolution of shared traits.
Fig. 4.
Convergent gene losses in species pairs from similar and distinct environments. A Venn diagram showing the one-to-one orthologous genes (large box) and overlapping pseudogenes in NMRs-BMRs pair. The numbers in parentheses represent the percentages of each category in the total number of pseudogenes in NMRs and BMRs (286 in total). P-value from a hypergeometric test indicates that gene losses in NMRs and BMRs are not independent from each other. B Proportional difference of overlapping pseudogenes among 4 species pairs including NMRs-BMRs, guinea pigs-rats, NMRs-rats, and BMRs-guinea pigs. The numbers of overlapping and total pseudogene from each pair are labelled above the bars. P-values are from two-tailed Fisher’s exact test
To associate overlapping pseudogenes with subterranean traits, we first leveraged knockout mouse phenotypic information (MPI) from the literature or MGI database. Degenerated visual capacity is one of the most significant traits that shared between NMRs and BMRs. We found that 5 convergent gene losses, including GUCY2F, ABCB5, RP1L1, CRB1, and ARR3 [70], are vision related and knockout mice of each gene show visual abnormality (Additional file 6: Table S5). It is worth noting that we also identified many other vision-related genes that lost specifically in each species, including genes reported by previously studies [4, 15, 33, 39–41], and new vision-related pseudogenes, such as CRB1 in BMRs (Additional file 7: Table S6). Thus, these gene losses collectively contribute to the eye degeneration in NMRs and BMRs. We then searched for the significant genetic associations (P < 1 × 10−6) with all overlapping genes that curated in the NHGRI-EBI GWAS catalog [71]. Interestingly, we found that variants in 5 genes, including ABCB5, LETM2, INMT, RP1L1, and CRB1, are significantly associated with cardiovascular traits including PR interval, pulse pressure, and systolic blood pressure [72–76] (Additional file 8: Table S7). Although further investigations are needed, these data imply contributions of losing such genes to cardiovascular adaptations in both NMRs and BMRs, especially to blood pressure control under hypoxia.
Previous functional studies also provide clues regarding the potential roles of gene losses in subterranean adaptations. For example, loss of EME2 could probably alleviate DNA damages induced by oxidative stress under hypoxic condition [51–53]. Another identified pseudogene, TRIM17, is more intriguing. It has been suggested that Trim17 is involved in trophic factor withdrawal-induced neuronal apoptosis [77]. Considering that both NMRs and BMRs are highly tolerant to hypoxia-induced brain injury [7, 13], it is possible that independent loss of TRIM17 in NMRs and BMRs contribute to protecting neural cells against hypoxia-induced apoptosis. However, no direct links have been established between hypoxic stress and the actions of TRIM17 in in vivo or in vitro. It is also possible that uncharacterized functions of TRIM17 are associated with other subterranean traits. For example, Trim17 knockout mice show physiological traits that are not involved in nerve system, including increased circulating alkaline phosphatase level (MP: 0002968) and increased circulating thyroxine level (MP: 0005477) (Additional file 6: Table S5). Therefore, functional investigations for these less well-characterized genes would provide insights into the roles of their functional losses in subterranean adaptations.
Functional loss of TRIM17 contributes to neuroprotection under hypoxia in vitro
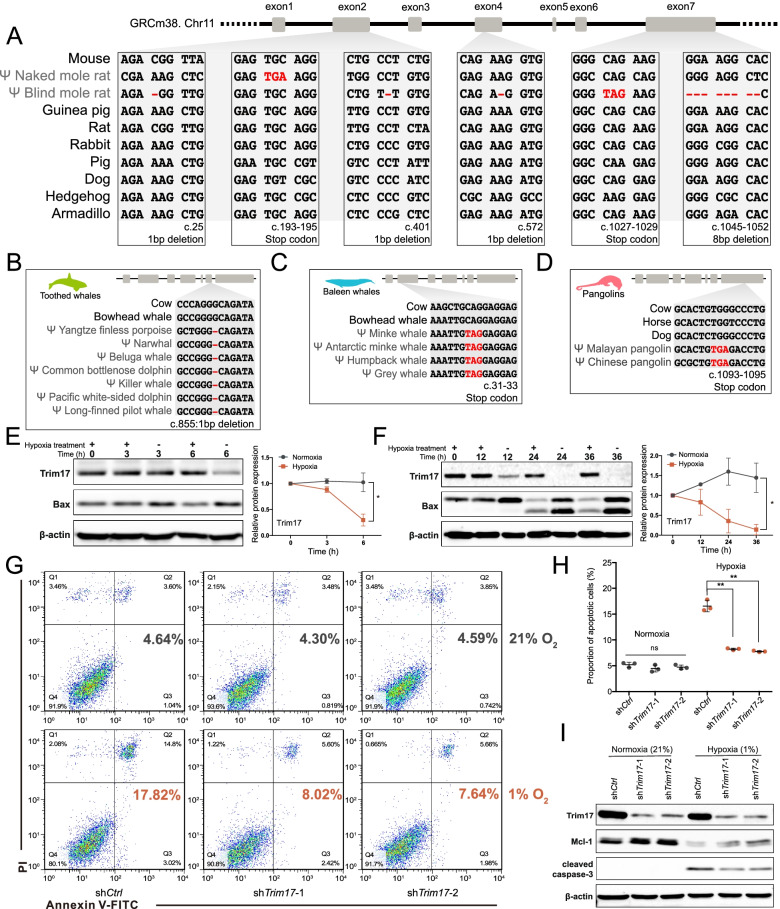
We hence chose TRIM17 to perform functional survey to examine whether loss of TRIM17 is involved in neuronal protection under hypoxia, which is a critical theme in hypoxic environment adaptation [78]. Disruptive mutations were confirmed by manual inspection of the alignment, showing that TRIM17 is inactivated by one premature stop codon in NMRs and 1 stop codon/4 indels in BMRs (Fig. 5A). The RING finger domain of TRIM17 is indispensable for its E3 ligase function [77] but is disrupted in both species, hinting the functional loss of this gene. We also examined the frequency of disruptive mutations in 11 BMR individuals [79], and no segregation was found. In NMRs, the premature stop codon was also verified by the available transcriptomic reads and genomic sequence of another individual (Additional file 4: Table S3). Furthermore, TRIM17 shows signature of relaxed selective constraint in both species, though not significant (NMRs: k = 0.69, P = 0.1338; BMRs: k = 0.72, P = 0.1214) (Additional file 2: Table S1). These results together confirmed the inactivation of TRIM17 in both NMRs and BMRs. Interestingly, we noticed that TRIM17 was independently lost at least two times in cetaceans, with a shared 1 bp deletion in toothed whales (Monodontidae, Phocoenidae, and Delphinidae) (Fig. 5B) and a shared premature stop codon in baleen whales (Balaenopteridae) (Fig. 5C), in consistent with a previous report [33]. Inactivating mutations were also detected in several species in other families including right whale (Eubalaena japonica), pygmy sperm whale (Kogia breviceps), and boutu (Inia geoffrensis) (Additional file 1: Fig. S2). Moreover, TRIM17 was also lost in two pangolins (Manis pentadactyla and Manis javanica) due to a shared premature stop codon (Fig. 5D) [33]. Like NMRs and BMRs, cetacean and pangolins experience hypoxic stress during their lifetime [78, 80, 81]. Therefore, independent losses of TRIM17 in distinct hypoxia-tolerant taxa suggest a relevance of TRIM17 losses in hypoxia adaptation.
Fig. 5.
Functional tests of TRIM17 for its role in hypoxia adaptation. A Coding sequence alignment of TRIM17 orthologous genes from mouse, NMR, BMR, and 7 other mammals. One stop codon in NMRs and one stop codon/four frameshifts in BMRs were observed that resulted in disrupted coding potential of TRIM17 gene. B–D Examples of inactivating mutations in toothed whales (B), baleen whales (C), and pangolins (D). The positions of all mutations shown in this figure are labelled under the corresponding alignment using mouse Trim17 coding sequence as reference. E–F Immunoblot analyses for Trim17 protein level changes during hypoxia treatments in primary cortical neurons (E) and N2a cell lines (F). “+” indicates hypoxia treatment and “−” indicates normoxia treatment. Quantification and relative fold change analysis of protein level showed a significant decrease of Trim17 protein induced by hypoxia in both primary cortical neurons and N2a cells. Data from three independent assays were used to perform paired two-tailed t-test, *P < 0.05. G–I Knockdown of Trim17 expression by shRNAs (shTrim17-1 and shTrim17-2) provided neuroprotection under hypoxia (1% O2) compared to control (shCtrl). Apoptotic cells were detected by staining cells with PI and Annexin V-FITC followed by flowmetry analysis. Cells with Trim17 knockdown showed significantly lower apoptosis rate under hypoxia compared to control, while no significant changes were observed under nomorxia (G, H), paired two-tail t-test, *P < 0.05, **P < 0.01. Immunoblot analysis for corresponding cells showed elevated Mcl-1 protein level and decreased activated caspase-3 protein level under hypoxia in cells with Trim17 knockdown (I)
To illustrate this idea, we next examined the changes in Trim17 expression in response to hypoxia in both Neuro-2a (N2a) cells and mouse primary cortical neurons. An immunoblot analysis showed that in contrast to trophic factor withdrawal, the Trim17 protein level was significantly decreased in both N2a cells and primary neurons after treating with hypoxic stress (1% O2) (Fig. 5E–F and Additional file 1: Fig. S3), suggesting a hypoxia-responsive regulatory mechanism for Trim17 protein. We further tested whether Trim17 knockdown confers a neuroprotective effect under hypoxia. N2a cell lines stably expressing shRNAs that targeted Trim17 and control shRNA were constructed and treated under hypoxic condition (1% O2) for 48 h, followed by an apoptosis analysis via flow cytometry. In contrast to the increased apoptotic rate in the control cells (shCtrl), the Trim17 knockdown cells (shTrim17-1, 2) showed significantly reduced apoptosis under hypoxia compared with normoxia (21% O2) (Fig. 5G–H). As Trim17 was shown to initiate apoptosis through the proteasome-dependent degradation of Mcl-1 [82], we examined whether this is also the case under hypoxia. An immunoblot analysis revealed elevated Mcl-1 protein level and consequently decreased active caspase-3 protein under hypoxia when compared to the control (Fig. 5I and Additional file 1: Fig. S3). Taken together, these results suggested that by preventing Mcl-1 protein from degradation, eliminating Trim17 function confers neuroprotection against hypoxia. Further investigations addressing the linkages among the components of phenotype-genotype-fitness continuum would provide us a better understanding of the contribution of gene loss to animal adaptation.
Gene losses and PSGs cooperatively contribute to subterranean adaptations
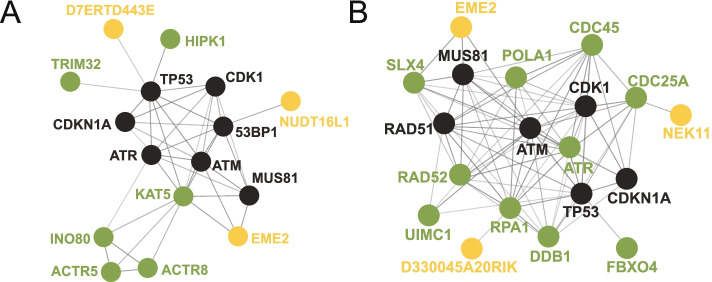
Complex physiological adaptations, such as modifications of DNA damage response and cardiovascular system, are expected to be involving multiple genes. We noticed that many genomic studies had uncovered PSGs that are also involved in DDR and heart functions in NMRs and BMRs [4, 15, 25, 29]. Unlike gene losses, PSGs show a signature of selection that can be easily detected, and thus could be assigned as adaptive. To examine whether PSGs also show enrichments in functional groups that similar to EFGs in pseudogenes, we collected PSGs from previous genomic studies and compiled a PSG set for NMRs [4, 29] and BMRs [15, 25], respectively (see “Methods”). Interestingly, for some of the EFGs in pseudogenes, we also observed similar functional enrichments in their respective PSG sets, including functional groups related to “DNA damage response,” “lipid metabolism,” and “neural processes” (Additional file 9: Table S8). Moreover, in consistent with the phenotypic convergence in heart contractile function, PSGs in functional group of “cardiovascular system” also show significant enrichment (Additional file 9: Table S8). In fact, several genes involved in cardiac muscle contraction in both species were under positive selection. These observations highlight the collaborative contributions of not only different genes, but also different types of genetic makeups to physiological adaptations. To illustrate this, we showed that in both NMRs and BMRs, protein-protein interaction network (STRING) involving DNA damage response contain both pseudogenes and PSGs, and both of them interact with other critical mediators such as ATM, ATR, or TP53 (Fig. 6). Such simultaneous gain-of-function and lost-of-function changes in multiple genes among a network reflect a strong rewiring process in response to drastic environmental changes, supporting an adaptive role of gene losses during subterranean adaptations.
Fig. 6.
Interacting network of DDR-related proteins. Different genes with different types of genetic changes are interacting with each other in both NMRs (A) and BMRs (B). PSGs are highlighted in green and gene losses are highlighted in yellow, and core mediators of DDR are highlighted in black. Interaction information was derived from STRING database (https://string-db.org/, Version 11.0)
Discussion
Challenges in genome-wide identification of adaptive gene losses
In this study, we systematically identified gene losses in two well-adapted subterranean rodential lineages, NMRs and BMRs, to examine the contribution of gene losses to subterranean adaptations after they split from mouse. By far, the identification of pseudogenes remains a challenging work because many factors can affect the final repertoires, including not only the sequencing, assembling, annotating qualities, but also the reference selection. Even though several genomic studies had identified pseudogenes in BMRs and NMRs (e.g., [4, 15]), to the best of our knowledge, this is the first study to identify gene losses using mouse rather than human as reference. For this reason, the repertoires of pseudogenes identified in our study would differ from others. For example, many pseudogenes identified using human as reference were generated in the ancestor of rodents and are not included in our pseudogene list. Meanwhile, it is not possible to identify loss of rodent-specific genes in NMRs and BMRs using human reference but this could be achieved using mouse reference. Our work thus provides an opportunity to survey the lineage-specific pseudogenization within rodents. Additionally, differences in the identification strategy and filtering criteria can generate different pseudogene repertoires as well. For example, the pseudogene lists of previous studies contain not only unitary pseudogenes, but also pseudogenes with potential functionally redundant paralogs. Here, we applied stringent criteria to maximumly reduce false positives (see “Methods”), e.g., the additional application of filtering signature of relaxed selection (RELAX) provides a most conservative result. The validation using BUSCO genes proved that our pipeline produces minimal false positives, although some real pseudogenes might be neglected due to the stringent filtering criteria. For example, RIPK3, a critical component in the necroptosis pathway, was recently reported to be lost in NMRs [83]. Our approaches did detect the disruptive mutations in RIPK3 sequence in both NMRs and BMRs. However, they were removed from our final list due to the poor GeneWise alignment (NMRs), or no signature of relaxed constraint (BMRs). In this way, we tried to guarantee the pseudogenes on our list are most reliable. Despite this, previously reported pseudogenes such as FASTK [4], MTNR1A [4], MTNR1B [4], TNNI3K [56], and AWAT2 [84] in NMRs and many vision-related pseudogenes in both NMRs and BMRs (Additional file 7: Table S6) are identified in our study, validating the effectiveness of our approaches. Collectively, our results provide a new perspective on subterranean adaptation at a smaller evolutionary scale and with higher resolution and give the community more insights into the molecular changes in response to subterranean environments.
Gene losses were generally thought to be deleterious and many pseudogenes in mammalian genome are the consequences of relaxed selective constraints. It is not an easy task to claim adaptive gene losses. Population genetic analysis could provide some evidences for such adaptation. As exampled by the CASPASE12 gene in human populations [85], the complete or nearly complete fixation of a null allele, in contrast to the high nucleotide diversity of the sequences adjacent to this gene, is the signature of positive selection and thus indicates an adaptive gene loss. However, population genetics approaches can only detect recent positive selection signatures (< 250,000 years) [86], and thus limit our capability of detecting ancient adaptive gene loss events. Unfortunately, the subterranean adaptations of both NMRs and BMRs occurred more than 30–40 Myrs ago [15], far beyond the detecting range of population genetics approaches. The sequence of ancient pseudogenes, even though they were adaptive and underwent positive selection at the early stage, usually evolves neutrally afterwards, making it difficult to detect the signal of positive selection. The direct evidence for an adaptive gene loss is the increased fitness of individuals with gene function loss compared to individuals with normal gene function in a population. However, it is a challenge to perform such tests for most of the non-model mammals. Previous genomic studies usually claim adaptive gene losses by associating them with adaptive traits, leveraging the available gene functional information or extrapolating phenotypic information from model organisms [33–37]. In this study, we also tried to link pseudogenes with adaptive traits of NMRs and BMRs based on currently available resources. We conducted genomic survey and functional group enrichment analyses to highlight several cases of gene losses to support the notion of adaptive gene losses, including genes involving DNA repair and cardiac contraction. More importantly, we tried to evidence their adaptive roles by showing the functional convergence of different gene losses in different species, in vitro functional assays of a convergent gene loss (TRIM17), and the interaction between pseudogenes and PSGs. Although further studies are still necessary to add more details in this process, our work provides a primary overview for the adaptive gene losses in subterranean lifestyle of NMRs and BMRs.
The adaptive role of losing TRIM17 in subterranean mole-rats
Despite the fact that current studies generally associate pseudogenes to the adaptive traits through information integration and speculation, the experimental evidence, either in vitro or in vivo, is more persuasive. In this study, we used TRIM17, one of the 20 convergently lost genes, as an example to demonstrate the beneficial role of gene inactivation in the subterranean adaptation. Although Trim17 has been reported to regulate neuronal apoptosis [77], its relationship with hypoxic insults is still not clear. For this reason, we provided functional evidence from an in vitro assay for the involvement of Trim17 in the response to hypoxia, and demonstrated the beneficial effects of losing Trim17 in neuroprotection under hypoxia. Therefore, by directly linking Trim17 with hypoxia, our results support the positive roles of inactivation of TRIM17 in both NMRs and BMRs in hypoxia tolerance during their transition to subterranean niches.
It is notable that Trim17 is also highly expressed in mouse testis, which usually implies an important reproductive related function. Unexpectedly, the Trim17 knockout mice show no abnormality of composition, quantity, and morphology of spermatogenic cells; the morphology and motility of the mature spermatozoa are not affected; and the fecundity of knockout males is comparable to that of the wild type [87]. This is probably due to the non-essentiality of Trim17 protein to fertility and the inactivation of Trim17 in testis might be compensated by other TRIM family members such as Trim58 or Trim69 [88]. In contrast, this compensation was not observed in brain/neuronal cells [77], in consistent with our observation of differentiated cell death between wild type and Trim17 knockdown N2a cells (Fig. 5G, H). A plausible explanation is that subterranean NMRs and BMRs may utilize some kind of tissue-specific mechanism to maintain the beneficial neuroprotective role of inactivated TRIM17 while not to dampen the fertility. Nevertheless, further investigations are necessary to address this issue in the future.
The interaction of PSGs and pseudogenes implies adaptive gene losses
For decades of years, the PSGs and pseudogenes are rarely discussed together, due to the traditional understanding of their distinct phenotypic effects. To the best of our knowledge, this study, for the first time, examined the interaction of PSGs and gene losses and highlighted a scenario in which PSGs and pseudogenes work together to promote adaptations. The occurrence of PSGs in a network containing pseudogenes suggests an evolution direction of the network that is not likely to be regressive, but adaptive. Therefore, this observation highly suggests the adaptive roles of gene losses in this network. Meanwhile, it also raises a question of how gene losses and PSGs work together to achieve a functional benefit. One can easily speculate that gene losses and PSGs may have similar functional effects that together result in an additive effect on a trait. For example, in the case of NMR DDR network, two genes, a PSG TRIM32 and a pseudogene D7ERTD443E (FATS) interact directly with TP53 (Fig. 6A). Both of them are ubiquitin ligases, but they regulate protein stability of TP53 in opposite directions, with a ubiquitination-mediated stabilization by FATS and a ubiquitination-mediated degradation by TRIM32 [48, 89]. Pseudogenization of FATS eliminates one layer of positive control of TP53 protein stability and alternatively facilitates negative regulation together with TRIM32, or other negative regulators. There might be other possibilities including the “sign-epistatic” interaction [90] and the compensation hypothesis that pseudogenes serve as a buffer for deleterious changes [91], although both await further investigations for verification in the future. Nevertheless, it is more likely that all these possible interactions exert their phenotypic effects simultaneously, due to the rewiring of a complex genetic network underlying physiological adaptations, especially to hypoxia. Our results highlight the complexity of underlying genetic mechanisms of subterranean adaptations, and considering the effects of different types of genetic makeups simultaneously could help us get a better understanding of adaptive evolution to subterranean niches or other extreme environments.
Conclusions
With combination of comparative genomic approaches and in vitro experimental validations, our study, for the first time, uncovered that gene losses might contribute to the evolution of specialized physiological traits in both NMRs and BMRs, including altered DNA damage response, specialized cardiovascular system, and neuroprotection. We also highlighted that the genetic mechanisms underlying physiological adaptations to subterranean niches are complex, involving not only multiple genes but also different types of genetic makeups. Taken together, our study provides new insights into the molecular underpinnings of subterranean adaptations, highlighting the important roles of gene losses. Several pseudogenes identified in BMRs and NMRs might have implications for medical researches involving aging and longevity, neuroprotection in ischemic/hypoxic injury, and cardiovascular diseases. Moreover, our study suggests an integration of different genetic makeups in future studies and also has implications for the comprehensive understanding of subterranean adaptation.
Methods
Genomic data
The genome sequences of each species analyzed in this study were retrieved from the Ensembl database [92] (mouse: GRCm38.p2, release 75; rat: Rnor_6.0; guinea pig: Cavpor 3.0, release 89) and NCBI database [93] (NMR: HetGla_female_1.0; BMR: S.galili_v1.0) together with its annotated coding sequences (CDS) and protein sequences. The CDS and protein sequences from rabbit, which were used to estimate the evolutionary rate, were retrieved from the Ensembl database (OryCun2.0).
Identification of gene losses
Basically, the gene losses in each species are defined as genes harboring ORF-disrupting mutations, including premature stop codons and/or frameshifts with intact 1:1 orthologs in the mouse genome. Orthologous relationships were first established by mapping the longest protein sequence for each mouse gene to each query species by simultaneously using BLAT [94] and genBlastA [95]. The genomic regions identified by both BLAT and genBlastA were considered as orthologous sequences and were extracted with 5000 bp downstream and upstream flanking sequences to predict the gene structure and ORF by GeneWise [96]. Alignments containing premature stop codons and/or frameshifts were identified, and the corresponding genomic regions were defined as putative pseudogenic loci. To establish the unitary status of each putative pseudogenic locus, several stringent filtering steps were executed as follows: (1) Loci hit by proteins belonging to large gene families including olfactory receptors, zinc finger protein, and vomeronasal receptors were removed because of their high sequence similarity. (2) Loci hit by predicted proteins or intronless cDNA/expressed sequences were removed because they are unlikely to be real or unitary [97]. (3) To remove the potential functional redundancy, e.g., gene duplication or retrotransposition, BLAT mapping hits from the initial step were reanalyzed to remove loci with homologous sequences across the genome. In detail, for each locus, we firstly removed hits with E-value higher than 1 × 10−3, and then we assembled the remaining hits into “copies” of this pseudogene along the mouse protein coordinates. Each copy includes non-overlapping hits that together cover the mouse protein as long as possible. Simply, for each “copy” with total protein length longer than 80% of the corresponding mouse protein, we considered it as a potential functional redundancy and therefore removed it. (4) We examined the conserved genomic position for each locus to ensure the orthologous relationship. Pairwise genomic alignments between mouse and each query species were downloaded from Ensembl database. For each locus, the genomic coordinates identified by us were compared to the coordinates inferred from a pairwise genomic alignment. Only pseudogenic loci with conserved genomic position were retained for further consideration. Furthermore, for genes discussed in the main text, we also manually examined the conserved gene order according to mouse genome annotation in Ensembl. To generate high-confidence pseudogene sets, false positives with disruptive mutations introduced by GeneWise, sequencing errors, or annotation errors were removed through the following steps: (a) in-house Perl script filtering in combination with manual inspection was performed to remove false positives introduced by GeneWise. Pseudogenes with a single disruptive mutation located at the end of the coding region (which resulted in truncated sequences longer than 90% of predicted intact protein [98], at the intron and exon boundaries or in regions with low similarity (30 bp up- and downstream the mutation, < 40% identity) in GeneWise alignments were removed. Pseudogenes with multiple disruptions were manually inspected to remove short or low-quality GeneWise alignments. For the genes discussed in the main text, we realigned the mutated exons on mouse genome sequence using CESAR [99] to remove spurious disruptive mutations caused by evolutionary splice site shifts. (b) To remove the sequencing and assembly errors, we retrieved the raw sequencing reads from NMRs and BMRs to check the validity of each disruption in the corresponding reads. For genes discussed in the main text, we further checked their validity using additional genomic/transcriptomic resources, i.e., raw transcriptomic reads of different tissues [100] and genomic sequences of a male individual for NMRs (HetGla_1.0) [101]; resequencing reads of 11 individuals for BMRs [102]. (c) The coding potential of the corresponding genes was examined by checking the Ensembl transcriptional support level (TSL, level 1/2) and CCDS assignment, and those satisfying any of the criteria were retained. (d) Pseudogenes with only two compensatory frameshifts were manually examined and removed because it provided little evidence for gene loss.
Detection of relaxed selection
Genes with disruptive mutations on coding sequence do not necessarily indicate gene dispensability. Such disruptive mutations might be rare genetic variation or sequencing/assembly errors. To address this issue, we detected signature of relaxed selection for each pseudogene by RELAX [103] and using a phylogenetic tree with rabbits as the outgroup. In detail, the coding sequences (CDS) and their translations (with disruptive mutations removed) of each pseudogene were extracted from the GeneWise alignments. The protein sequences were used to identify 1:1 orthologous gene in the rest of the species for each pseudogene by reciprocal best BLAST hit (RBBH) method, and then, the results were clustered into orthologous groups. For each orthologous group, CDS were used to construct codon-based multiple sequence alignment with a phylogeny-aware alignment algorithm as implemented in the PRANK program [104]. To obtain reliable CDS alignments, we set up rigorous filtering criteria including: (i) removing gaps and ambiguous bases together with 6 bp flanking sequences with Gblocks [105] using the default parameters; (ii) removing alignment fragments with low quality (Gblocks, default parameters); and (iii) removing alignments with lengths of less than 150 bp. Alignments containing at least two sequences were assigned to a specific tree file. In each alignment, the species in which this gene loss occurred was set as “Foreground” branch and the rest of species were set as “Background” branch. Only pseudogenes with a higher dN/dS in foreground and a selection intensity parameter k < 1 (k > 1: intensified selection; k < 1: relaxed selection) were retained.
Gene ontology
Orthologous mouse genes were used as surrogates to infer the functions of the pseudogenes or PSGs in corresponding species. Only pseudogenes and PSGs with orthologous relationship with mouse (established by BLAT and genBlastA mapping) were further considered. To identify the EFGs from pseudogenes, the associated GO terms for each gene were extracted from go.obo (released February 20, 2018), and the mouse gene annotation files (Mus musculus, released September 29, 2017) were downloaded from the Gene Ontology Consortium (http://geneontology.org/). NMR and BMR genes with GO terms involved in certain biological processes and systems were manually clustered into functional groups based on the keywords. For each functional group, we then compared the proportional differences of genes from subterranean lineage pseudogene lists, subterranean lineage non-pseudogene lists (with one2one mouse orthologs), control lineage pseudogene lists, and control lineage non-pseudogene lists (with one2one mouse orthologs). For example, a subset of GO terms with a significantly higher proportion of NMR pseudogenes compared to that of NMR non-pseudogenes and/or that of guinea pig pseudogenes was defined as an EFG. For PSGs, EFGs were defined only when significantly higher proportion of NMR (or BMR) PSGs in a subset of GO terms was observed compared to that of NMR (or BMR) non-PSG background (with one2one mouse orthologs). Fisher’s exact test was used to test the significances of differences in proportion and FDR-adjusted P-values for each multiple test were reported using p.adjust() function in R.
Cell culture
Primary cortical neurons were prepared from E16-E18 embryos of Kunming mice. Specifically, embryonic cortices were dissected in cold DMEM (Gibco) supplemented with 10% fetal bovine serum (Gibco) and then digested using 2 mg/ml papain (CHI). The resulting cell suspension was centrifuged at 1500 rpm for 5 min. The cells were gently resuspended in DMEM supplemented with 10% FBS and gentamycin (50 μg/ml, Solarbio) and filtered through a 70-μm cell strainer (Corning Falcon). Then, the cells were seeded at a density of 1.2 × 106 cells/ml in culture dishes coated with 20 μg/ml poly-D-lysine (Sigma). The neurons were cultured at 37 °C in a humidified incubator with 5% CO2/95% air, and the medium was replaced 6 h afterwards with neurobasal medium (Gibco) supplemented with B27 supplements (Gibco) and L-glutamine (Gibco). After 6 days in vitro (DIV 6), the neural cultures were subjected to hypoxia treatment and protein extraction. N2a cells were purchased from the Cell Bank of Type Culture Collection of the Chinese Academy of Sciences (Shanghai; Institution Code: CBTCCCAS) and were cultured at 37 °C in a humidified incubator with 5% CO2/95% air with MEM (Gibco) supplemented with 1.5 mg/ml NaHCO3, 0.11 mg/ml sodium pyruvate (Gibco), and 10% FBS.
Hypoxia treatment paradigms
The hypoxia (1% oxygen concentration) for the cell culture treatment was achieved using a humidified hypoxia chamber (Proox, model C21, BioSpherix) set at 37 °C, 1% O2, and 5% CO2 and maintained with a N2 supply. The primary cortical neuron cultures and N2a cells (including the stable cell lines) were treated at 1% hypoxia for various periods of time. As controls, cells were cultured at 37 °C in a humidified incubator with 5% CO2 and 95% air.
Immunoblot analysis
Primary cortical neuron and N2a cells (including stable cell lines) under both hypoxic and normoxic conditions were collected and lysed using RIPA buffer (Biomiga) supplemented with a protease inhibitor phenylmethanesulfonyl fluoride (PMSF) and cocktail. Immunoblotting experiments were performed according to a previous method [106, 107]. Briefly, proteins were separated by 10% SDS-PAGE gel followed by electrotransfer to PVDF membranes (Millipore). The membranes were blocked with TBST supplemented with 10% skim milk for 30 min at room temperature and further probed with primary antibodies at 4 °C overnight including Trim17 (1:500, Sigma); Mcl-1 (1:1000, CST); Bax (1:1000, CST); and cleaved caspase 3 (1:500, CST) with β-actin (1:500, GeneTex) as the loading control. The membranes were subsequently incubated with HRP-conjugated secondary antibodies, and the protein bands were detected by ECL (Thermo Scientific). The quantifications of the protein expression levels were conducted by the gray scanning of the target proteins with Image-Pro Plus 6.0 software. Three independent experiments were performed to test the expression differences.
shRNA-lentiviral infection and apoptosis detection of N2a cells
The HIV-derived lentiviral vector pLKO.1 containing shRNAs (TRIM17 MISSION shRNA Bacterial Glycerol Stock, Sigma) that targeted Trim17 (shTrim17-1: CCATCTGCCTTGACTACTTTA; shTrim17-2: CTGTTACCCAATTCCACTCTA) and control shRNA (shCtrl: GACACTGGGTGTGCCACAGTT) together with lentiviral packaging plasmids pCMVΔ8.9 and pMD2.G were co-transfected into HEK293T cells to generate lentiviruses. Supernatants containing different lentiviruses were collected 48 h and 72 h post-transfection, respectively. N2a cells were infected with supernatants in the presence of 6 μg/ml polybrene for 48 h, and 2 μg/ml puromycin was used to select the stable cell lines. Three stable cell lines expressing shRNAs were cultured simultaneously under 1% hypoxia and 21% normoxia for 48 h and collected for FITC-conjugated annexin V (BD Pharmingen) staining followed by apoptotic cell number analysis by flow cytometry according to a method described previously [108]. Three independent experiments were performed for apoptosis detection to test the differences.
Statistical analysis
Two-tailed Fisher’s exact tests were used to detect the proportional differences in pseudogenes for each functional group. Paired t-tests were used to analyze differences in Trim17 protein levels and apoptosis rates. Hypergeometric tests (http://systems.crump.ucla.edu/hypergeometric/index.php) were used to assess the significance of the overlapping pseudogenes.
Supplementary Information
Additional file 1: Fig. S1. Flowchart for identifying gene loss events in NMRs, BMRs, guinea pigs and rats. Fig. S2. Inactivating mutations of TRIM17 in another three cetaceans. Fig. S3. Original western blot images.
Additional file 2: Table S1. Gene losses identified in each species.
Additional file 3: Table S2. Results of functional group enrichment analysis.
Additional file 4: Table S3. Mutations of pseudogenes discussed in the main text.
Additional file 5: Table S4. Independent gene losses between NMRs and BMRs.
Additional file 6: Table S5. MGI MP information for overlapping pseudogenes.
Additional file 7: Table S6. Vision related pseudogenes.
Additional file 8: Table S7. Genetic associations of pseudogenes with cardiovascular traits.
Additional file 9: Table S8. Functional group enrichment analysis in PSGs in NMRs and BMRs.
Acknowledgements
We thank Dr. Xudong Zhao, Dr. Cuiping Yang, Dr. Dong Yang, and Dr. Tao Zhang for their technical support, Dr. Huabin Zhao for providing BMR tissue samples. We also thank Dr. Michael Hiller, Dr. Vera Gorbunova, Dr. Andrei Seluanov, and three anonymous reviewers for their valuable comments.
Abbreviations
- BMRs
Blind mole-rats
- CDS
Coding sequences
- CGLs
Convergent gene losses
- DDR
DNA damage response
- DSBs
Double-stranded breaks
- EBI
The European Bioinformatics Institute
- EFGs
Enriched functional groups
- GO
Gene Ontology
- GWAS
Genome-Wide Association Studies
- HRP
Horseradish peroxidase
- MGI
Mouse Genome Informatics
- MNDCM
Mononuclear diploid cardiomyocyte
- MPI
Mice phenotypic information
- NHGRI
National Human Genome Research Institute
- NMRs
Naked mole-rats
- ORF
Open reading frame
- PSGs
Positively selected genes
Authors’ contributions
PS and ZZ conceived the project and designed the research. ZZ and RH performed the research. ZZ, HY, PS, and GX analyzed data. ZZ, HY, and PS wrote the manuscript with input from all authors. All authors have read and approved the final version of the manuscript.
Funding
This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (grant no. XDB13020400), the National Natural Science Foundation of China (grant no. 31871277), and Yunling Scholar Project, Ten-Thousand Talents Plan of Yunnan Province (to PS).
Availability of data and materials
The datasets supporting the conclusions of this article are included within the article and its additional files. The Perl scripts used for pseudogene screening and identification are available on GitHub (https://github.com/zzheng24/Gene-losses).
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Zhizhong Zheng and Rong Hua contributed equally to this work.
Contributor Information
Hui Yang, Email: yanghui@mail.kiz.ac.cn.
Peng Shi, Email: ship@mail.kiz.ac.cn.
References
- 1.Nevo E. Adaptive convergence and divergence of subterranean mammals. Annu Rev Ecol Syst. 1979;10(1):269–308. [Google Scholar]
- 2.Perez VI, Buffenstein R, Masamsetti V, Leonard S, Salmon AB, Mele J, Andziak B, Yang T, Edrey Y, Friguet B, et al. Protein stability and resistance to oxidative stress are determinants of longevity in the longest-living rodent, the naked mole-rat. Proc Natl Acad Sci U S A. 2009;106(9):3059–3064. doi: 10.1073/pnas.0809620106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Seluanov A, Hine C, Azpurua J, Feigenson M, Bozzella M, Mao Z, Catania KC, Gorbunova V. Hypersensitivity to contact inhibition provides a clue to cancer resistance of naked mole-rat. Proc Natl Acad Sci U S A. 2009;106(46):19352–19357. doi: 10.1073/pnas.0905252106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kim EB, Fang X, Fushan AA, Huang Z, Lobanov AV, Han L, Marino SM, Sun X, Turanov AA, Yang P, et al. Genome sequencing reveals insights into physiology and longevity of the naked mole rat. Nature. 2011;479(7372):223–227. doi: 10.1038/nature10533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Smith ES, Omerbasic D, Lechner SG, Anirudhan G, Lapatsina L, Lewin GR. The molecular basis of acid insensitivity in the African naked mole-rat. Science. 2011;334(6062):1557–1560. doi: 10.1126/science.1213760. [DOI] [PubMed] [Google Scholar]
- 6.Tian X, Azpurua J, Hine C, Vaidya A, Myakishev-Rempel M, Ablaeva J, Mao Z, Nevo E, Gorbunova V, Seluanov A. High-molecular-mass hyaluronan mediates the cancer resistance of the naked mole rat. Nature. 2013;499(7458):346–349. doi: 10.1038/nature12234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Park TJ, Reznick J, Peterson BL, Blass G, Omerbasic D, Bennett NC, Kuich P, Zasada C, Browe BM, Hamann W, et al. Fructose-driven glycolysis supports anoxia resistance in the naked mole-rat. Science. 2017;356(6335):307–311. doi: 10.1126/science.aab3896. [DOI] [PubMed] [Google Scholar]
- 8.Zhao Y, Tyshkovskiy A, Munoz-Espin D, Tian X, Serrano M, de Magalhaes JP, Nevo E, Gladyshev VN, Seluanov A, Gorbunova V. Naked mole rats can undergo developmental, oncogene-induced and DNA damage-induced cellular senescence. Proc Natl Acad Sci U S A. 2018;115(8):1801–1806. doi: 10.1073/pnas.1721160115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Avivi A, Albrecht U, Oster H, Joel A, Beiles A, Nevo E. Biological clock in total darkness: the Clock/MOP3 circadian system of the blind subterranean mole rat. Proc Natl Acad Sci U S A. 2001;98(24):13751–13756. doi: 10.1073/pnas.181484498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ashur-Fabian O, Avivi A, Trakhtenbrot L, Adamsky K, Cohen M, Kajakaro G, Joel A, Amariglio N, Nevo E, Rechavi G. Evolution of p53 in hypoxia-stressed Spalax mimics human tumor mutation. Proc Natl Acad Sci U S A. 2004;101(33):12236–12241. doi: 10.1073/pnas.0404998101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Shams I, Avivi A, Nevo E. Hypoxic stress tolerance of the blind subterranean mole rat: expression of erythropoietin and hypoxia-inducible factor 1 alpha. Proc Natl Acad Sci U S A. 2004;101(26):9698–9703. doi: 10.1073/pnas.0403540101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Nasser NJ, Nevo E, Shafat I, Ilan N, Vlodavsky I, Avivi A. Adaptive evolution of heparanase in hypoxia-tolerant Spalax: gene cloning and identification of a unique splice variant. Proc Natl Acad Sci U S A. 2005;102(42):15161–15166. doi: 10.1073/pnas.0507279102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Avivi A, Gerlach F, Joel A, Reuss S, Burmester T, Nevo E, Hankeln T. Neuroglobin, cytoglobin, and myoglobin contribute to hypoxia adaptation of the subterranean mole rat Spalax. Proc Natl Acad Sci U S A. 2010;107(50):21570–21575. doi: 10.1073/pnas.1015379107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gorbunova V, Hine C, Tian X, Ablaeva J, Gudkov AV, Nevo E, Seluanov A. Cancer resistance in the blind mole rat is mediated by concerted necrotic cell death mechanism. Proc Natl Acad Sci U S A. 2012;109(47):19392–19396. doi: 10.1073/pnas.1217211109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fang X, Nevo E, Han L, Levanon EY, Zhao J, Avivi A, Larkin D, Jiang X, Feranchuk S, Zhu Y, et al. Genome-wide adaptive complexes to underground stresses in blind mole rats Spalax. Nat Commun. 2014;5:3966. doi: 10.1038/ncomms4966. [DOI] [PubMed] [Google Scholar]
- 16.Delaney MA, Ward JM, Walsh TF, Chinnadurai SK, Kerns K, Kinsel MJ, Treuting PM. Initial case reports of cancer in naked mole-rats (Heterocephalus glaber) Vet Pathol. 2016;53(3):691–696. doi: 10.1177/0300985816630796. [DOI] [PubMed] [Google Scholar]
- 17.Jarvis JU. Eusociality in a mammal: cooperative breeding in naked mole-rat colonies. Science. 1981;212(4494):571–573. doi: 10.1126/science.7209555. [DOI] [PubMed] [Google Scholar]
- 18.Buffenstein R, Woodley R, Thomadakis C, Daly TJ, Gray DA. Cold-induced changes in thyroid function in a poikilothermic mammal, the naked mole-rat. Am J Physiol Regul Integr Comp Physiol. 2001;280(1):R149–R155. doi: 10.1152/ajpregu.2001.280.1.R149. [DOI] [PubMed] [Google Scholar]
- 19.Davis-Walton J, Sherman PW. Sleep arrhythmia in the eusocial naked mole-rat. Naturwissenschaften. 1994;81(6):272–275. doi: 10.1007/BF01131581. [DOI] [PubMed] [Google Scholar]
- 20.Mann MD, Rehkamper G, Reinke H, Frahm HD, Necker R, Nevo E. Size of somatosensory cortex and of somatosensory thalamic nuclei of the naturally blind mole rat, Spalax ehrenbergi. J Hirnforsch. 1997;38(1):47–59. [PubMed] [Google Scholar]
- 21.Kimchi T, Terkel J. Magnetic compass orientation in the blind mole rat Spalax ehrenbergi. J Exp Biol. 2001;204(Pt 4):751–758. doi: 10.1242/jeb.204.4.751. [DOI] [PubMed] [Google Scholar]
- 22.Widmer HR, Hoppeler H, Nevo E, Taylor CR, Weibel ER. Working underground: respiratory adaptations in the blind mole rat. Proc Natl Acad Sci U S A. 1997;94(5):2062–2067. doi: 10.1073/pnas.94.5.2062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Du K, Yang L, He S. Phylogenomic analyses reveal a molecular signature linked to subterranean adaptation in rodents. BMC Evol Biol. 2015;15:287. doi: 10.1186/s12862-015-0564-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Shao Y, Li JX, Ge RL, Zhong L, Irwin DM, Murphy RW, Zhang YP. Genetic adaptations of the plateau zokor in high-elevation burrows. Sci Rep. 2015;5:17262. doi: 10.1038/srep17262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Davies KTJ, Bennett NC, Faulkes CG, Rossiter SJ. Limited evidence for parallel molecular adaptations associated with the subterranean niche in mammals: a comparative study of three superorders. Mol Biol Evol. 2018;35(10):2544–2559. doi: 10.1093/molbev/msy161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Liu Z, Wang W, Zhang TZ, Li GH, He K, Huang JF, Jiang XL, Murphy RW, Shi P. Repeated functional convergent effects of NaV1.7 on acid insensitivity in hibernating mammals. Proc Biol Sci. 2014;281(1776):20132950. doi: 10.1098/rspb.2013.2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Malik A, Korol A, Hubner S, Hernandez AG, Thimmapuram J, Ali S, Glaser F, Paz A, Avivi A, Band M. Transcriptome sequencing of the blind subterranean mole rat, Spalax galili: utility and potential for the discovery of novel evolutionary patterns. Plos One. 2011;6(8):e21227. doi: 10.1371/journal.pone.0021227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Malik A, Korol A, Weber M, Hankeln T, Avivi A, Band M. Transcriptome analysis of the spalax hypoxia survival response includes suppression of apoptosis and tight control of angiogenesis. BMC Genomics. 2012;13:615. doi: 10.1186/1471-2164-13-615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Fang X, Seim I, Huang Z, Gerashchenko MV, Xiong Z, Turanov AA, Zhu Y, Lobanov AV, Fan D, Yim SH, et al. Adaptations to a subterranean environment and longevity revealed by the analysis of mole rat genomes. Cell Rep. 2014;8(5):1354–1364. doi: 10.1016/j.celrep.2014.07.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Albalat R, Canestro C. Evolution by gene loss. Nat Rev Genet. 2016;17(7):379–391. doi: 10.1038/nrg.2016.39. [DOI] [PubMed] [Google Scholar]
- 31.Wittkopp PJ, Kalay G. Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence. Nat Rev Genet. 2012;13(1):59–69. doi: 10.1038/nrg3095. [DOI] [PubMed] [Google Scholar]
- 32.Bonduriansky R, Day T. Nongenetic inheritance and its evolutionary implications. Annu Rev Ecol Evol Syst. 2009;40:103–125. [Google Scholar]
- 33.Sharma V, Hecker N, Roscito JG, Foerster L, Langer BE, Hiller M. A genomics approach reveals insights into the importance of gene losses for mammalian adaptations. Nat Commun. 2018;9(1):1215. doi: 10.1038/s41467-018-03667-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Huelsmann M, Hecker N, Springer MS, Gatesy J, Sharma V, Hiller M. Genes lost during the transition from land to water in cetaceans highlight genomic changes associated with aquatic adaptations. Sci Adv. 2019;5(9):eaaw6671. doi: 10.1126/sciadv.aaw6671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lachner J, Mlitz V, Tschachler E, Eckhart L. Epidermal cornification is preceded by the expression of a keratinocyte-specific set of pyroptosis-related genes. Sci Rep. 2017;7(1):17446. doi: 10.1038/s41598-017-17782-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Espregueira Themudo G, Alves LQ, Machado AM, Lopes-Marques M, da Fonseca RR, Fonseca M, Ruivo R, Castro LFC. Losing genes: the evolutionary remodeling of cetacea skin. Front Mar Sci. 2020;7:592375. [Google Scholar]
- 37.Hecker N, Sharma V, Hiller M. Convergent gene losses illuminate metabolic and physiological changes in herbivores and carnivores. Proc Natl Acad Sci U S A. 2019;116(8):3036–3041. doi: 10.1073/pnas.1818504116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Olson MV. When less is more: gene loss as an engine of evolutionary change. Am J Hum Genet. 1999;64(1):18–23. doi: 10.1086/302219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Emerling CA, Springer MS. Eyes underground: regression of visual protein networks in subterranean mammals. Mol Phylogenet Evol. 2014;78:260–270. doi: 10.1016/j.ympev.2014.05.016. [DOI] [PubMed] [Google Scholar]
- 40.Prudent X, Parra G, Schwede P, Roscito JG, Hiller M. Controlling for phylogenetic relatedness and evolutionary rates improves the discovery of associations between species’ phenotypic and genomic differences. Mol Biol Evol. 2016;33(8):2135–2150. doi: 10.1093/molbev/msw098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Emerling CA. Regressed but not gone: patterns of vision gene loss and retention in subterranean mammals. Integr Comp Biol. 2018;58(3):441–451. doi: 10.1093/icb/icy004. [DOI] [PubMed] [Google Scholar]
- 42.Vavouri T, Semple JI, Lehner B. Widespread conservation of genetic redundancy during a billion years of eukaryotic evolution. Trends Genet. 2008;24(10):485–488. doi: 10.1016/j.tig.2008.08.005. [DOI] [PubMed] [Google Scholar]
- 43.Manni M, Berkeley MR, Seppey M, Simao FA, Zdobnov EM. BUSCO update: Novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol Biol Evol. 2021;38(10):4647–4654. doi: 10.1093/molbev/msab199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mitchell TW, Buffenstein R, Hulbert AJ. Membrane phospholipid composition may contribute to exceptional longevity of the naked mole-rat (Heterocephalus glaber): a comparative study using shotgun lipidomics. Exp Gerontol. 2007;42(11):1053–1062. doi: 10.1016/j.exger.2007.09.004. [DOI] [PubMed] [Google Scholar]
- 45.Melixetian M, Klein DK, Sorensen CS, Helin K. NEK11 regulates CDC25A degradation and the IR-induced G2/M checkpoint. Nat Cell Biol. 2009;11(10):1247–1253. doi: 10.1038/ncb1969. [DOI] [PubMed] [Google Scholar]
- 46.Li Z, Zhang Q, Mao JH, Weise A, Mrasek K, Fan X, Zhang X, Liehr T, Lu KH, Balmain A, et al. An HDAC1-binding domain within FATS bridges p21 turnover to radiation-induced tumorigenesis. Oncogene. 2010;29(18):2659–2671. doi: 10.1038/onc.2010.19. [DOI] [PubMed] [Google Scholar]
- 47.Zhang X, Zhang Q, Zhang J, Qiu L, Yan SS, Feng J, Sun Y, Huang X, Lu KH, Li Z. FATS is a transcriptional target of p53 and associated with antitumor activity. Mol Cancer. 2010;9:244. doi: 10.1186/1476-4598-9-244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Yan S, Qiu L, Ma K, Zhang X, Zhao Y, Zhang J, Li X, Hao X, Li Z. FATS is an E2-independent ubiquitin ligase that stabilizes p53 and promotes its activation in response to DNA damage. Oncogene. 2014;33(47):5424–5433. doi: 10.1038/onc.2013.494. [DOI] [PubMed] [Google Scholar]
- 49.Bhat KP, Krishnamoorthy A, Dungrawala H, Garcin EB, Modesti M, Cortez D. RADX modulates RAD51 activity to control replication fork protection. Cell Rep. 2018;24(3):538–545. doi: 10.1016/j.celrep.2018.06.061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Drane P, Brault ME, Cui G, Meghani K, Chaubey S, Detappe A, Parnandi N, He Y, Zheng XF, Botuyan MV, et al. TIRR regulates 53BP1 by masking its histone methyl-lysine binding function. Nature. 2017;543(7644):211–216. doi: 10.1038/nature21358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Amangyeld T, Shin YK, Lee M, Kwon B, Seo YS. Human MUS81-EME2 can cleave a variety of DNA structures including intact Holliday junction and nicked duplex. Nucleic Acids Res. 2014;42(9):5846–5862. doi: 10.1093/nar/gku237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Pepe A, West SC. Substrate specificity of the MUS81-EME2 structure selective endonuclease. Nucleic Acids Res. 2014;42(6):3833–3845. doi: 10.1093/nar/gkt1333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Techer H, Koundrioukoff S, Carignon S, Wilhelm T, Millot GA, Lopez BS, Brison O, Debatisse M. Signaling from Mus81-Eme2-dependent DNA damage elicited by Chk1 deficiency modulates replication fork speed and origin usage. Cell Rep. 2016;14(5):1114–1127. doi: 10.1016/j.celrep.2015.12.093. [DOI] [PubMed] [Google Scholar]
- 54.Petruseva IO, Evdokimov AN, Lavrik OI. Genome stability maintenance in naked mole-rat. Acta Naturae. 2017;9(4):31–41. [PMC free article] [PubMed] [Google Scholar]
- 55.Domankevich V, Eddini H, Odeh A, Shams I. Resistance to DNA damage and enhanced DNA repair capacity in the hypoxia-tolerant blind mole rat Spalax carmeli. J Exp Biol. 2018;221(8):jeb174540. doi: 10.1242/jeb.174540. [DOI] [PubMed] [Google Scholar]
- 56.Gan P, Patterson M, Velasquez A, Wang K, Tian D, Windle JJ, Tao G, Judge DP, Makita T, Park TJ, et al. Tnni3k alleles influence ventricular mononuclear diploid cardiomyocyte frequency. Plos Genet. 2019;15(10):e1008354. doi: 10.1371/journal.pgen.1008354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Aoyagi T, Wada T, Kojima F, Nagai M, Harada S, Hachisu M, Murata S, Umezawa H. Inhibitors of aminopeptidase B suppress the development of hypertension in spontaneously hypertensive rats. Chem Pharm Bull (Tokyo) 1986;34(11):4852–4854. doi: 10.1248/cpb.34.4852. [DOI] [PubMed] [Google Scholar]
- 58.Borlaug BA, Melenovsky V, Marhin T, Fitzgerald P, Kass DA. Sildenafil inhibits beta-adrenergic-stimulated cardiac contractility in humans. Circulation. 2005;112(17):2642–2649. doi: 10.1161/CIRCULATIONAHA.105.540500. [DOI] [PubMed] [Google Scholar]
- 59.Wang H, Wang L, Song L, Zhang YW, Ye J, Xu RX, Shi N, Meng XM. TNNI3K is a novel mediator of myofilament function and phosphorylates cardiac troponin I. Braz J Med Biol Res. 2013;46(2):128–137. doi: 10.1590/1414-431X20122515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Arieli R, Ar A. Heart-rate responses of the mole rat (Spalax-Ehrenbergi) in hypercapnic, hypoxic, and cold conditions. Physiol Zool. 1981;54(1):14–21. [Google Scholar]
- 61.Grimes KM, Voorhees A, Chiao YA, Han HC, Lindsey ML, Buffenstein R. Cardiac function of the naked mole-rat: ecophysiological responses to working underground. Am J Physiol Heart Circ Physiol. 2014;306(5):H730–H737. doi: 10.1152/ajpheart.00831.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Grimes KM, Reddy AK, Lindsey ML, Buffenstein R. And the beat goes on: maintained cardiovascular function during aging in the longest-lived rodent, the naked mole-rat. Am J Physiol Heart Circ Physiol. 2014;307(3):H284–H291. doi: 10.1152/ajpheart.00305.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Takimoto E, Champion HC, Li M, Belardi D, Ren S, Rodriguez ER, Bedja D, Gabrielson KL, Wang Y, Kass DA. Chronic inhibition of cyclic GMP phosphodiesterase 5A prevents and reverses cardiac hypertrophy. Nat Med. 2005;11(2):214–222. doi: 10.1038/nm1175. [DOI] [PubMed] [Google Scholar]
- 64.Perez NG, Piaggio MR, Ennis IL, Garciarena CD, Morales C, Escudero EM, Cingolani OH. Chiappe de Cingolani G, Yang XP, Cingolani HE: Phosphodiesterase 5A inhibition induces Na+/H+ exchanger blockade and protection against myocardial infarction. Hypertension. 2007;49(5):1095–1103. doi: 10.1161/HYPERTENSIONAHA.107.087759. [DOI] [PubMed] [Google Scholar]
- 65.Tedford RJ, Hemnes AR, Russell SD, Wittstein IS, Mahmud M, Zaiman AL, Mathai SC, Thiemann DR, Hassoun PM, Girgis RE, et al. PDE5A inhibitor treatment of persistent pulmonary hypertension after mechanical circulatory support. Circ Heart Fail. 2008;1(4):213–219. doi: 10.1161/CIRCHEARTFAILURE.108.796789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Giannetta E, Isidori AM, Galea N, Carbone I, Mandosi E, Vizza CD, Naro F, Morano S, Fedele F, Lenzi A. Chronic inhibition of cGMP phosphodiesterase 5A improves diabetic cardiomyopathy: A randomized, controlled clinical trial using magnetic resonance imaging with myocardial tagging. Circulation. 2012;125(19):2323–2333. doi: 10.1161/CIRCULATIONAHA.111.063412. [DOI] [PubMed] [Google Scholar]
- 67.Vagnozzi RJ, Gatto GJ, Jr, Kallander LS, Hoffman NE, Mallilankaraman K, Ballard VL, Lawhorn BG, Stoy P, Philp J, Graves AP, et al. Inhibition of the cardiomyocyte-specific kinase TNNI3K limits oxidative stress, injury, and adverse remodeling in the ischemic heart. Sci Transl Med. 2013;5(207):207ra141. doi: 10.1126/scitranslmed.3006479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Patterson M, Barske L, Van Handel B, Rau CD, Gan P, Sharma A, Parikh S, Denholtz M, Huang Y, Yamaguchi Y, et al. Frequency of mononuclear diploid cardiomyocytes underlies natural variation in heart regeneration. Nat Genet. 2017;49(9):1346–1353. doi: 10.1038/ng.3929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Meyer WK, Jamison J, Richter R, Woods SE, Partha R, Kowalczyk A, Kronk C, Chikina M, Bonde RK, Crocker DE, et al. Ancient convergent losses of Paraoxonase 1 yield potential risks for modern marine mammals. Science. 2018;361(6402):591–594. doi: 10.1126/science.aap7714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Deming JD, Pak JS, Brown BM, Kim MK, Aung MH, Eom YS, Shin JA, Lee EJ, Pardue MT, Craft CM. Visual cone arrestin 4 contributes to visual function and cone health. Invest Ophthalmol Vis Sci. 2015;56(9):5407–5416. doi: 10.1167/iovs.15-16647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.MacArthur J, Bowler E, Cerezo M, Gil L, Hall P, Hastings E, Junkins H, McMahon A, Milano A, Morales J. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog) Nucleic Acids Res. 2017;45(D1):D896–D901. doi: 10.1093/nar/gkw1133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Deng X, Sabino EC, Cunha-Neto E, Ribeiro AL, Ianni B, Mady C, Busch MP, Seielstad M. Genome wide association study (GWAS) of Chagas cardiomyopathy in Trypanosoma cruzi seropositive subjects. Plos One. 2013;8(11):e79629. doi: 10.1371/journal.pone.0079629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Evangelou E, Warren HR, Mosen-Ansorena D, Mifsud B, Pazoki R, Gao H, Ntritsos G, Dimou N, Cabrera CP, Karaman I. Genetic analysis of over 1 million people identifies 535 new loci associated with blood pressure traits. Nat genet. 2018;50(10):1412–1425. doi: 10.1038/s41588-018-0205-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Feitosa MF, Kraja AT, Chasman DI, Sung YJ, Winkler TW, Ntalla I, Guo X, Franceschini N, Cheng C-Y, Sim X. Novel genetic associations for blood pressure identified via gene-alcohol interaction in up to 570K individuals across multiple ancestries. Plos One. 2018;13(6):e0198166. doi: 10.1371/journal.pone.0198166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Ramírez J, van Duijvenboden S, Ntalla I, Mifsud B, Warren HR, Tzanis E, Orini M, Tinker A, Lambiase PD, Munroe PB. Thirty loci identified for heart rate response to exercise and recovery implicate autonomic nervous system. Nat Commun. 2018;9(1):1–10. doi: 10.1038/s41467-018-04148-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Giri A, Hellwege JN, Keaton JM, Park J, Qiu C, Warren HR, Torstenson ES, Kovesdy CP, Sun YV, Wilson OD. Trans-ethnic association study of blood pressure determinants in over 750,000 individuals. Nat Genet. 2019;51(1):51–62. doi: 10.1038/s41588-018-0303-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Lassot I, Robbins I, Kristiansen M, Rahmeh R, Jaudon F, Magiera MM, Mora S, Vanhille L, Lipkin A, Pettmann B, et al. Trim17, a novel E3 ubiquitin-ligase, initiates neuronal apoptosis. Cell Death Differ. 2010;17(12):1928–1941. doi: 10.1038/cdd.2010.73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Ramirez JM, Folkow LP, Blix AS. Hypoxia tolerance in mammals and birds: from the wilderness to the clinic. Annu Rev Physiol. 2007;69(1):113–143. doi: 10.1146/annurev.physiol.69.031905.163111. [DOI] [PubMed] [Google Scholar]
- 79.Li K, Hong W, Jiao H, Wang GD, Rodriguez KA, Buffenstein R, Zhao Y, Nevo E, Zhao H. Sympatric speciation revealed by genome-wide divergence in the blind mole rat Spalax. Proc Natl Acad Sci U S A. 2015;112(38):11905–11910. doi: 10.1073/pnas.1514896112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Weber RE, Heath ME, White FN. Oxygen binding functions of blood and hemoglobin from the Chinese pangolin, Manis pentadactyla: possible implications of burrowing and low body temperature. Respir Physiol. 1986;64(1):103–112. doi: 10.1016/0034-5687(86)90064-2. [DOI] [PubMed] [Google Scholar]
- 81.Choo SW, Rayko M, Tan TK, Hari R, Komissarov A, Wee WY, Yurchenko AA, Kliver S, Tamazian G, Antunes A, et al. Pangolin genomes and the evolution of mammalian scales and immunity. Genome Res. 2016;26(10):1312–1322. doi: 10.1101/gr.203521.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Magiera MM, Mora S, Mojsa B, Robbins I, Lassot I, Desagher S. Trim17-mediated ubiquitination and degradation of Mcl-1 initiate apoptosis in neurons. Cell Death Differ. 2013;20(2):281–292. doi: 10.1038/cdd.2012.124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Agueda-Pinto A, Alves LQ, Neves F, McFadden G, Jacobs BL, Castro LFC, Rahman MM, Esteves PJ. Convergent loss of the necroptosis pathway in disparate mammalian lineages shapes viruses countermeasures. Front Immunol. 2021;12:747737. doi: 10.3389/fimmu.2021.747737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Lopes-Marques M, Machado AM, Alves LQ, Fonseca MM, Barbosa S, Sinding M-HS, Rasmussen MH, Iversen MR, Frost Bertelsen M, Campos PF, et al. Complete inactivation of sebum-producing genes parallels the loss of sebaceous glands in Cetacea. Mol Biol Evol. 2019;36(6):1270–1280. doi: 10.1093/molbev/msz068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Wang X, Grus WE, Zhang J. Gene losses during human origins. Plos Biol. 2006;4(3):e52. doi: 10.1371/journal.pbio.0040052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Sabeti PC, Schaffner SF, Fry B, Lohmueller J, Varilly P, Shamovsky O, Palma A, Mikkelsen T, Altshuler D, Lander E. Positive natural selection in the human lineage. Science. 2006;312(5780):1614–1620. doi: 10.1126/science.1124309. [DOI] [PubMed] [Google Scholar]
- 87.Lu Y, Oura S, Matsumura T, Oji A, Sakurai N, Fujihara Y, Shimada K, Miyata H, Tobita T, Noda T, et al. CRISPR/Cas9-mediated genome editing reveals 30 testis-enriched genes dispensable for male fertility in mice. Biol Reprod. 2019;101(2):501–511. doi: 10.1093/biolre/ioz103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.He X, Xie W, Li H, Cui Y, Wang Y, Guo X, Sha J. The testis-specifically expressed gene Trim69 is not essential for fertility in mice. J Biomed Res. 2021;35(1):47–60. doi: 10.7555/JBR.34.20200069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Liu J, Zhang C, Wang XL, Ly P, Belyi V, Xu-Monette ZY, Young KH, Hu W, Feng Z. E3 ubiquitin ligase TRIM32 negatively regulates tumor suppressor p53 to promote tumorigenesis. Cell Death Differ. 2014;21(11):1792–1804. doi: 10.1038/cdd.2014.121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Weinreich DM, Delaney NF, Depristo MA, Hartl DL. Darwinian evolution can follow only very few mutational paths to fitter proteins. Science. 2006;312(5770):111–114. doi: 10.1126/science.1123539. [DOI] [PubMed] [Google Scholar]
- 91.Szamecz B, Boross G, Kalapis D, Kovacs K, Fekete G, Farkas Z, Lazar V, Hrtyan M, Kemmeren P, Groot Koerkamp MJ, et al. The genomic landscape of compensatory evolution. Plos Biol. 2014;12(8):e1001935. doi: 10.1371/journal.pbio.1001935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.The Ensembl database. http://useast.ensembl.org/index.html. Accessed Oct 2014.
- 93.The NCBI database. https://www.ncbi.nlm.nih.gov. Accessed Oct 2014
- 94.Kent WJ. BLAT--the BLAST-like alignment tool. Genome Res. 2002;12(4):656–664. doi: 10.1101/gr.229202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.She R, Chu JS, Wang K, Pei J, Chen N. GenBlastA: enabling BLAST to identify homologous gene sequences. Genome Res. 2009;19(1):143–149. doi: 10.1101/gr.082081.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Birney E, Clamp M, Durbin R. GeneWise and Genomewise. Genome Res. 2004;14(5):988–995. doi: 10.1101/gr.1865504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Zhang ZD, Frankish A, Hunt T, Harrow J, Gerstein M. Identification and analysis of unitary pseudogenes: historic and contemporary gene losses in humans and other primates. Genome Biol. 2010;11(3):R26. doi: 10.1186/gb-2010-11-3-r26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.MacArthur DG, Balasubramanian S, Frankish A, Huang N, Morris J, Walter K, Jostins L, Habegger L, Pickrell JK, Montgomery SB, et al. A systematic survey of loss-of-function variants in human protein-coding genes. Science. 2012;335(6070):823–828. doi: 10.1126/science.1215040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Sharma V, Schwede P, Hiller M. CESAR 2.0 substantially improves speed and accuracy of comparative gene annotation. Bioinformatics. 2017;33(24):3985–3987. doi: 10.1093/bioinformatics/btx527. [DOI] [PubMed] [Google Scholar]
- 100.Kim EB, Fang X, Fushan AA, Huang Z, Lobanov AV, Han L, et al. Genome sequencing reveals insights into physiology and longevity of the naked mole rat. Bioproject. 2011; https://identifiers.org/GEO/GSE30337. [DOI] [PMC free article] [PubMed]
- 101.Kim EB, Fang X, Fushan AA, Huang Z, Lobanov AV, Han L, et al. Genome sequencing reveals insights into physiology and longevity of the naked mole rat. Bioproject. 2011; https://identifiers.org/bioproject/PRJNA68323. [DOI] [PMC free article] [PubMed]
- 102.Li K-X, Hong W, Jiao H-W, Wang G-D, Rodriguez KA, Buffenstein R, et al. Sympatric speciation revealed by genome-wide divergence in the blind mole rat Spalax. Bioproject https://identifiers.org/bioproject/PRJNA285131. 2015. [DOI] [PMC free article] [PubMed]
- 103.Wertheim JO, Murrell B, Smith MD, Kosakovsky Pond SL, Scheffler K. RELAX: detecting relaxed selection in a phylogenetic framework. Mol Biol Evol. 2015;32(3):820–832. doi: 10.1093/molbev/msu400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Loytynoja A, Goldman N. An algorithm for progressive multiple alignment of sequences with insertions. Proc Natl Acad Sci U S A. 2005;102(30):10557–10562. doi: 10.1073/pnas.0409137102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 2000;17(4):540–552. doi: 10.1093/oxfordjournals.molbev.a026334. [DOI] [PubMed] [Google Scholar]
- 106.Guo DK, Zhu Y, Sun HY, Xu XY, Zhang S, Hao ZB, Wang GH, Mu CC, Ren HG. Pharmacological activation of REV-ERBα represses LPS-induced microglial activation through the NF-κB pathway. Acta Pharmacol Sin. 2019;40(1):26–34. doi: 10.1038/s41401-018-0064-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Yu W, Wang B, Zhou L, Xu G. Endoplasmic reticulum stress-mediated p62 downregulation inhibits apoptosis via c-Jun upregulation. Biomol Ther (Seoul) 2021;29(2):195–204. doi: 10.4062/biomolther.2020.089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Shen X, Zhao YF, Xu SQ, Wang L, Cao HM, Cao Y, Zhu Y, Wang Y, Liang ZQ. Cathepsin L induced PC-12 cell apoptosis via activation of B-Myb and regulation of cell cycle proteins. Acta Pharmacol Sin. 2019;40(11):1394–1403. doi: 10.1038/s41401-019-0286-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Fig. S1. Flowchart for identifying gene loss events in NMRs, BMRs, guinea pigs and rats. Fig. S2. Inactivating mutations of TRIM17 in another three cetaceans. Fig. S3. Original western blot images.
Additional file 2: Table S1. Gene losses identified in each species.
Additional file 3: Table S2. Results of functional group enrichment analysis.
Additional file 4: Table S3. Mutations of pseudogenes discussed in the main text.
Additional file 5: Table S4. Independent gene losses between NMRs and BMRs.
Additional file 6: Table S5. MGI MP information for overlapping pseudogenes.
Additional file 7: Table S6. Vision related pseudogenes.
Additional file 8: Table S7. Genetic associations of pseudogenes with cardiovascular traits.
Additional file 9: Table S8. Functional group enrichment analysis in PSGs in NMRs and BMRs.
Data Availability Statement
The datasets supporting the conclusions of this article are included within the article and its additional files. The Perl scripts used for pseudogene screening and identification are available on GitHub (https://github.com/zzheng24/Gene-losses).