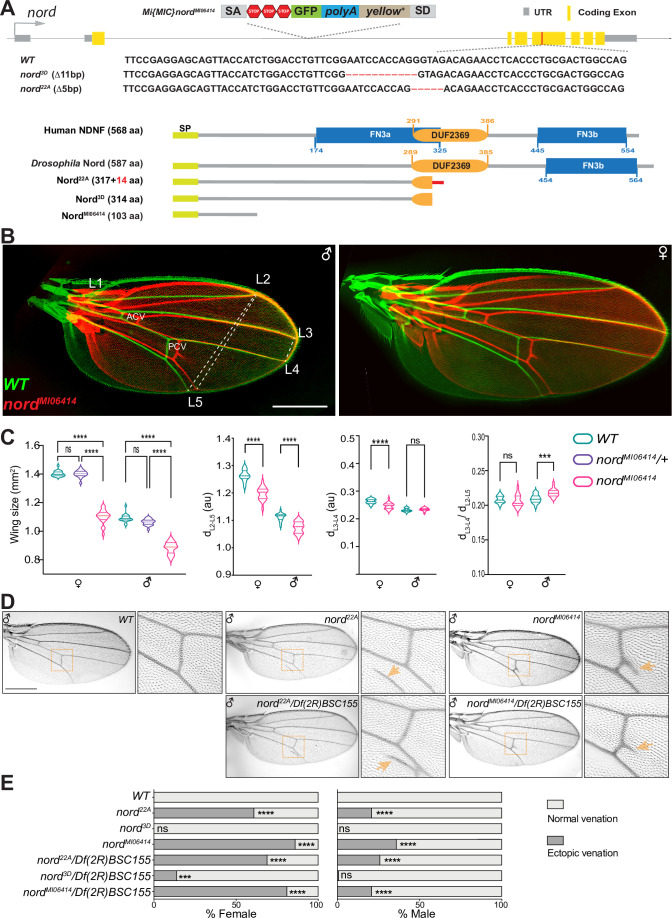
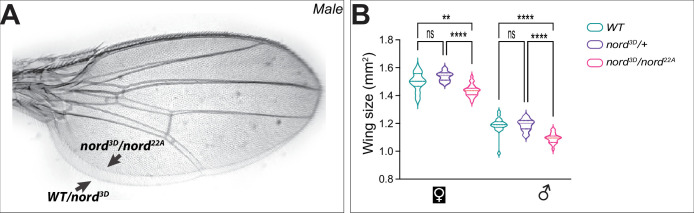
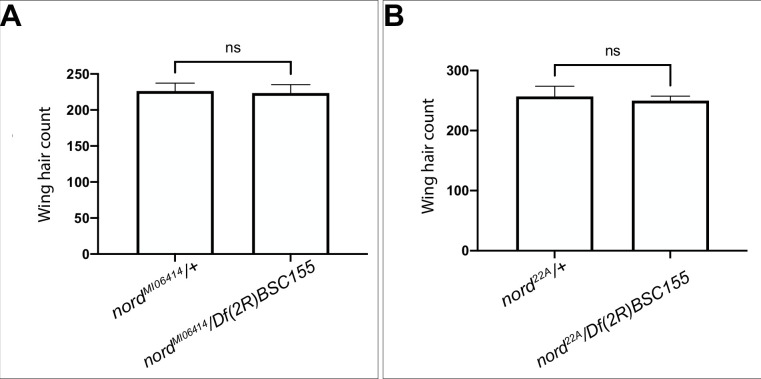
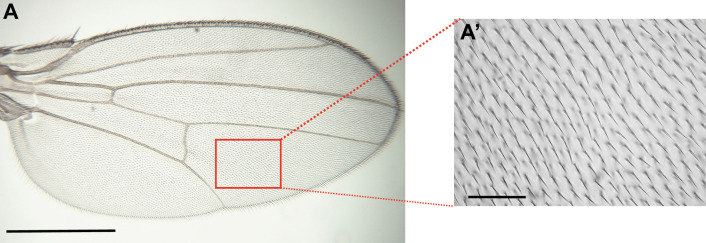
Figure 4. Nord is required for proper crossvein patterning and growth of the Drosophila wing.
(A) Upper panel: schematic diagram of the wild-type nord locus, the gene-trap, and the CRISPR alleles of nord. A Minos-Mediated Integration Cassette (MiMIC) cassette consisting of a splice acceptor site followed by stop codons in all three reading frames was inserted into the first coding intron of nord in the nord gene-trap allele nordMI0641. Detailed view of the deleted regions in the nord mutant alleles generated by the CRISPR/Cas9 system. Lower panel: schematic diagram of the human Neuron-Derived Neurotrophic Factor (NDNF) protein, Drosophila Nord protein, and the predicted polypeptide products from the indicated nord mutant alleles. (B) Adult wings obtained from male or female flies with indicated genotypes were pseudo-colored and overlapped to show the size difference. ACV, anterior crossvein; PCV, posterior crossvein; LV, longitudinal veins (L1–L5). (C) Quantification of wing size, distance between distal ends of LVs L2 and L5 (dL2–L5), L3 and L4 (dL3–L4), and the ratio of dL3–L4 /dL2–L5. Each bar shows the mean ± SD from n = 20 wings. All flies were grown at 25°C. One-way ANOVA followed by Sidak’s multiple comparison test or unpaired two-tailed t-test was used for statistical analysis. ***p<0.001, ****p<0.0001, ns, not significant; au, arbitrary units. (D) Adult wings of flies with the indicated genotypes. Yellow arrowhead indicates ectopic vein near posterior crossvein (PCV) or L5. (E) Quantification of the ectopic venation phenotype in adult wings from flies with the indicated genotypes. n > 50. Two-sided Fisher’s exact tests were used for statistical analysis. ***p<0.001, ****p<0.0001, ns, not significant. Scale bar, 500 μm.