Figure 3.
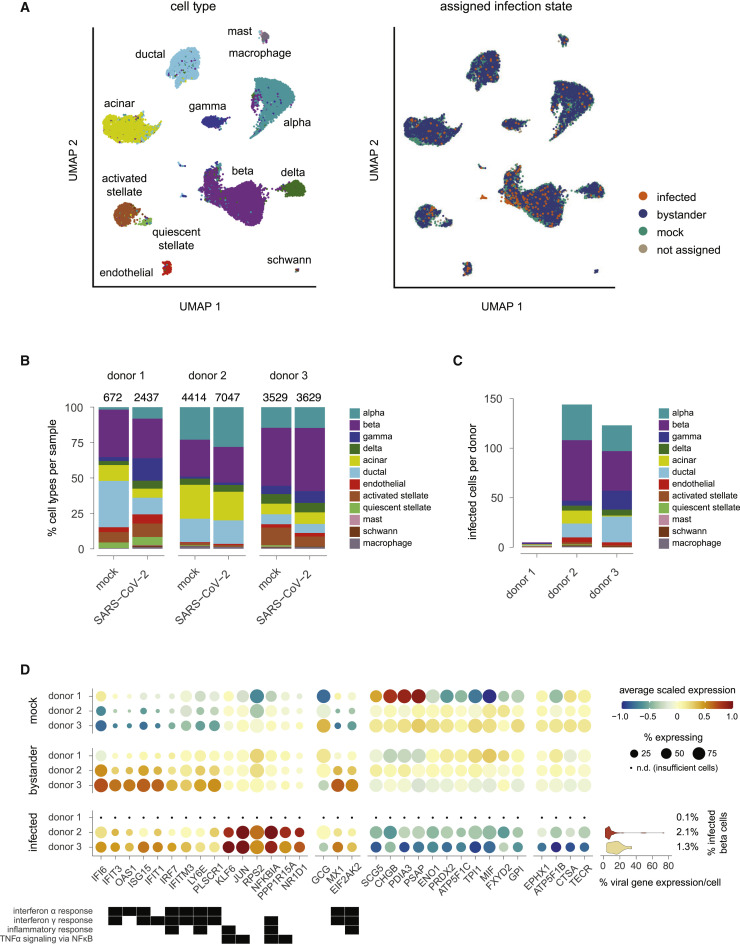
scRNA-seq analysis of pancreatic islets infected in vitro with SARS-CoV-2
(A) UMAP (Uniform Manifold Approximation and Projection) of 21,728 single cells integrated across samples from 3 donors, each infected or mock-infected with SARS-CoV-2. Points are colored by assigned cell type (left) or assigned infection state (right).
(B) Relative cell type composition of each sample, with total number of cells per sample indicated at the top.
(C) Number of infected cells per donor (SARS-CoV-2-infected samples only) by cell type.
(D) Differentially expressed genes in pairwise comparisons between infected, bystander, or mock-infected β cells; selected genes have an adjusted p value of less than 0.01 and absolute log2 fold change greater than log2(1.5) in any contrast within any donor. Dot size indicates the percentage of β cells in the indicated donor and infection state expressing the gene; dot color indicates the average expression of that gene in β cells scaled across infection state within each donor. Filled boxes (below) indicate gene membership in the indicated hallmark gene sets. For infected cells, additional annotation (right) indicates the percentage of β cells infected per donor, and the violin plot indicates percent of all detected transcripts identified as viral transcripts per cell. Because of insufficient numbers of infected β cells in donor 1, this sample was excluded from differential expression contrasts involving infected cells.