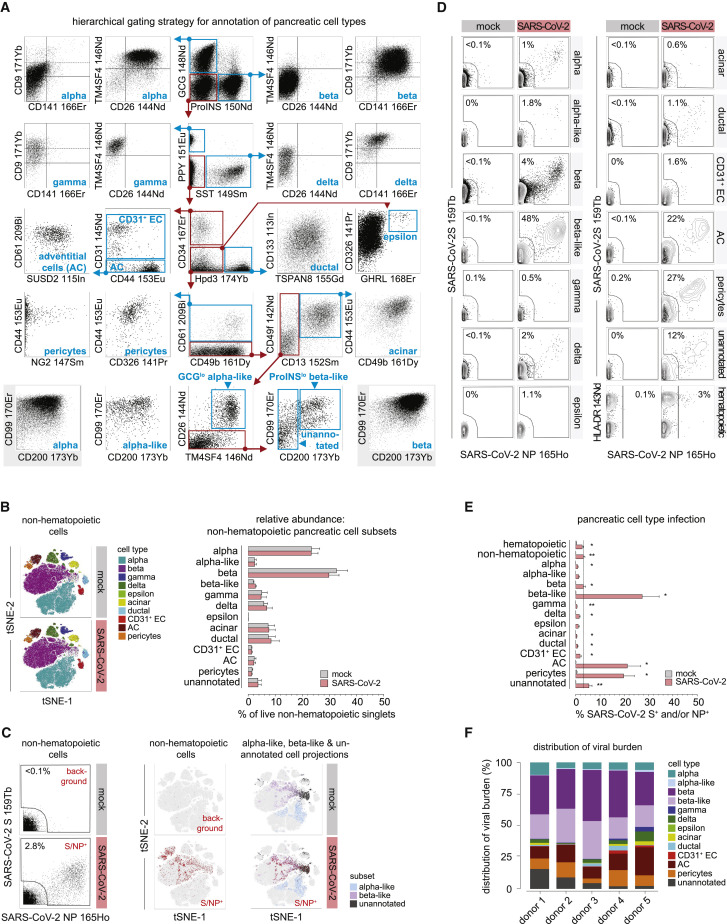
Figure 4.
MC-based pancreatic cell annotation and quantification of SARS-CoV-2 infection
(A) Hierarchical gating strategy for identification of major pancreatic cell subsets. Starting with a distinction of α (GCG+) and β (proinsulin [proINS]+) cells (top row, center), the panel represents a flow chart where demarcated regions and associated arrows connect successive plots to designate and characterize pancreatic cell populations (PPY, pancreatic polypeptide; SST, somatostatin; GHRL, ghrelin). Regions, arrows, and cell type names rendered in blue highlight final cell annotations. Red regions/arrows refer to subsets selected for further phenotypic stratification. The bottom left and right plots with a gray background feature comparative α and β cell properties.
(B) Left: tSNE visualization of pancreatic cell populations in 48-h mock- and SARS-CoV-2-infected samples. Right: relative abundance of non-hematopoietic pancreatic cell subsets in mock- and SARS-CoV-2-infected samples (n = 5 donors).
(C) Left: co-detection of SARS-CoV-2-S/NP in live non-hematopoietic cells. Center: distribution of viral burden visualized by projection onto tSNE plots. Right: α-like, β-like, and unannotated cell projections onto tSNE clusters and associated viral burden (viral infection of other cell types are excluded here).
(D) Representative SARS-CoV-2-S and -NP expression in mock- and SARS-CoV-2-infected pancreatic cell types (48-h culture).
(E) Extent of SARS-CoV-2 infection in all pancreatic cell types (dotted line, average infection extent for all non-hematopoietic cells; n = 5 donors; asterisks indicate statistical differences calculated between mock- and SARS-CoV-2-infected samples by paired t test; ∗, p < 0.05; ∗∗, p < 0.01).
(F) Relative distribution of the viral burden across pancreatic cell subsets.