ABSTRACT
Background
The relationship between dietary n-3 PUFAs and the prevention of cardiometabolic diseases, including type 2 diabetes, is unresolved. Examination of the association between n-3 PUFAs and chronic low-grade inflammation in a population where many individuals have had an extremely high intake of marine mammals and fish throughout their lifespan may provide important clues regarding the impact of n-3 PUFAs on health.
Objectives
The aim of this study was to explore associations between concentrations of n-3 PUFAs resulting from habitual intake of natural food sources high in fish and marine mammals with immune biomarkers of metabolic inflammation and parameters of glucose regulation.
Methods
A total of 569 Yup'ik Alaska Native adults (18–87 years old) were enrolled in this cross-sectional study between December 2016 and November 2019. The RBC nitrogen isotope ratio (NIR; 15N/14N) was used as a validated measure of n-3 PUFA intake to select 165 participant samples from the first and fourth quartiles of n-3 PUFA intakes. Outcomes included 38 pro- and anti-inflammatory cytokines and 8 measures of glucose homeostasis associated with type 2 diabetes risks. These outcomes were evaluated for their associations with direct measurements of EPA, DHA, and arachidonic acid in RBCs.
Analysis
Linear regression was used to detect significant relationships with cytokines and n-3 PUFAs, adiposity, and glucose-related variables.
Results
The DHA concentration in RBC membranes was inversely associated with IL-6 (β = –0.0066; P < 0.001); EPA was inversely associated with TNFα (β = –0.4925; P < 0.001); and the NIR was inversely associated with Monocyte chemoattractant protein-1 (MCP-1) (β = –0.8345; P < 0.001) and IL-10 (β = –1.2868; P < 0.001).
Conclusions
Habitual intake of marine mammals and fish rich in n-3 PUFAs in this study population of Yup'ik Alaska Native adults is associated with reduced systemic inflammation, which may contribute to the low prevalence of diseases in which inflammation plays an important role.
Keywords: fish oil, omega fatty acids, cytokines, chemokines, immune biomarkers
Introduction
Inflammation often accompanies obesity and is a risk factor for chronic, noncommunicable diseases (1), including type 2 diabetes (T2D) (2). Obesity-associated inflammation is associated with insulin resistance and thought to be mediated in part through the production of TNFα in adipose tissue (3, 4). In Yup'ik Alaska Native people, the prevalence of overweight and obesity is similar to that of the general US population (5), but the prevalence of T2D is only ∼3%, less than one-fourth of that reported in the US general population (6, 7). We hypothesize that this discordance between the prevalence of obesity and T2D in Yup'ik people is, in part, due to their intake of a traditional subsistence diet highly enriched in anti-inflammatory n-3 PUFAs derived from marine mammals and fish (8).
Dietary intake of saturated fat leads to the activation of inflammatory cytokine gene expression and the inhibition of insulin signaling (9). In contrast, n-3 PUFA intake inhibits the production of proinflammatory cytokines (8, 10–12), even in the presence of obesity (8); helps to maintain insulin sensitivity (9, 13, 14); and has cardioprotective effects (9). Historically, Yup'ik people lived a very active lifestyle and relied exclusively on locally harvested foods, including fish, marine mammals, berries, game, and wild greens. This traditional Yup'ik diet includes amounts of EPA and DHA up to 20 times higher than those observed in the general US population (15). Yup'ik individuals consuming this traditional diet exhibit cardioprotective lipid profiles and elevated adiponectin (16), which may contribute to their low T2D prevalence, lower blood pressure measurements (17), and reduced platelet activation (18).
Metabolic inflammation resulting from recruitment of both innate and adaptive immune cells is associated with immune cell infiltration of several tissues, including adipose, pancreas, liver, and brain tissues (4, 19, 20). Proinflammatory M1-polarized macrophages, CD4 + Th1 cells, and CD8 + effector T cells commonly found in adipose tissues of obese individuals collectively contribute to the synthesis and release of a spectrum of proinflammatory cytokines and chemokines believed to play a significant role in the development of insulin resistance and impaired β-cell secretory function (21), the combination of which leads to T2D.
Although not commonly considered among the immunomodulatory therapies proposed to prevent and treat cardiometabolic diseases such as T2D (22, 23), n-3 PUFAs have been shown to have protective effects against insulin resistance in rodent models, but not in humans, where results have been mixed (24). Most randomized controlled trials of n-3 PUFA supplementation have resulted in no demonstrable benefits on insulin sensitivity (25). However, these studies are typically of limited duration and may not capture the effects of habitually high n-3 PUFA intakes, utilize n-3 PUFA doses sufficiently high enough to create meaningful blood concentration differences between the control and treatment groups (26), or incorporate the n-3 PUFAs into a whole foods dietary pattern (27).
To address several of these limitations in the current literature, we comprehensively investigated associations between EPA, DHA, and arachidonic acid (AA) in RBCs with markers of chronic low-grade inflammation in a population habituated to a diet high in natural sources of EPA and DHA. In addition, we examined associations between these n-3 and n-6 PUFAs with circulating adiponectin; high-sensitivity C-reactive protein (hsCRP); and regulated upon activation, normal T-cell expressed and secreted [Regulated on Activation, Normal T Cell Expressed and Secreted (RANTES)]; as well as with measures of insulin resistance (HOMA2-IR), insulin sensitivity (HOMA2-S), beta cell function (HOMA2-B), and insulin secretory capacity relative to insulin sensitivity [HOMA2-disposition index (HOMA2-DI)] (28).
Methods
Participants and data collection
For this study, from a cohort of 569 participants, we selected a sub-sample of 216 Yup'ik participants at the extremes of n-3 PUFA intakes using an objective biomarker of n-3 PUFA intake: the nitrogen isotope ratio (NIR; 15N/14N) in RBCs (29). Immune biomarkers were only measured on this sub-sample of 216 participants because of limited resources and the exploratory nature of this study. The sex distributions were unbalanced at the extremes, forcing us to include some individuals in the second and third quartiles of NIR to balance the sexes. However, in the experience of the core laboratory that runs our clinical assays, cytokine panels may provide higher values due to an increased background or cross-reactivity with other targets in the panel. Therefore, for the purpose of the current study, we chose to limit the statistical analyses to quartiles 1 and 4 of the NIR distribution; 17 participants in the second quartile and 4 in the third quartile were thus excluded, leaving 195 participants. From this participant sub-sample, 7 participants were excluded because they had an hsCRP concentration > 1 mg/dL, 15 were excluded because they had a BMI below 20 kg/m2, 5 were excluded because they had a BMI above 40 kg/m2, 4 were excluded because they had T2D, 1 were excluded due to taking an immunosuppressive medication, and 2 participants had 2 of the exclusion criteria, leaving 165 participants for statistical analyses. BMI cutoff criteria were used because we previously found (8) the relationship between BMI and several clinical biomarkers to be nonlinear at BMIs <20 or >40 kg/m2. The study data were collected and managed using REDCap electronic data capture tools hosted at Oregon Health and Science University (OHSU) (30, 31). This study was approved by the University of Alaska Fairbanks (UAF) and OHSU Institutional Review Boards, as well as by the Yukon-Kuskokwim Health Corporation Human Studies Committee and Executive Board of Directors.
Anthropometric measurements and blood sample collection
Anthropometric measures and a fasting blood sample were obtained as previously described (32). Anthropometric measurements included height, weight, waist circumference (WC), percentage body fat, and resting blood pressure. Two trained researchers collected measures from all participants using standard NHANES III Anthropometric Procedures Manual protocols (33), as previously described (34). Blood samples were obtained by venipuncture from individuals after a 12-hour fast, collected in EDTA tubes, and processed into plasma, lymphocyte, and RBC fractions in the rural communities using a portable centrifuge. Samples for glucose analysis were collected in sodium fluoride tubes, immediately placed in a cooler rack, and processed within 5 minutes. All samples were initially stored between −15°C to −20°C for up to a week (while in rural Alaska), before being shipped to the UAF or OHSU and stored at −80°C or in liquid nitrogen cryovats.
Dietary biomarker measurements
The NIRs of RBC samples collected from EDTA tubes were analyzed by the Alaska Stable Isotope Facility at the UAF by continuous-flow isotope ratio MS as previously described (29). Natural abundance stable isotope ratios are conventionally expressed as the per mille (‰) abundance of 15N relative to an internal standard:
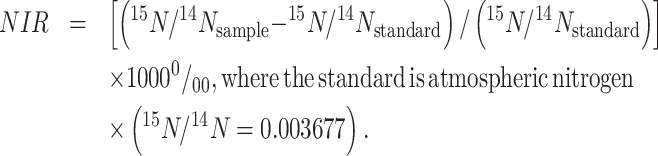 |
(1) |
The NIR has previously been validated as an objective biomarker of EPA and DHA intake in this population, and reflects the past 3–4 months of dietary intake (29, 35). As shown previously (29), there was a positive, linear correlation between EPA and NIR; a positive, nonlinear relationship between DHA and NIR; and an inverse, linear correlation between AA and NIR (Supplemental Figure 1).
Analysis of PUFAs from RBCs
The analytical method was adapted from a previously published protocol (36, 37). Briefly, a 10 μl aliquot of packed RBCs was diluted 1:100 with 0.9% NaCl in water, and deuterated EPA, DHA, AA internal standards, butylated hydroxytoluene, and triphenylphosphine were added. A 20 μL aliquot of the diluted sample was removed for further processing. An additional 50 μl aliquot was further diluted 1:4 with 0.9% NaCl in water for total protein quantitation using a bicinchoninic acid (BCA) kit. The 1:100 diluted sample was dried under a gentle stream of nitrogen, reconstituted in isopropanol, saponified with the addition of 200 μl 1M Potassium hydroxide (KOH), and then incubated in a water bath at 37°C for 45 minutes. The hydrolyzed sample was then neutralized with the addition of 200 μl 1M of HCl. Following neutralization, analytes of interest (AA, EPA, and DHA) were isolated by Biotage 96-well Evolute Express ABN SPE 30-mg plate solid-phase extraction, dried under a gentle stream of nitrogen, and derivatized with the addition of AMPP+ [N-(4-aminomethylphenyl)pyridinium]. The derivatized samples were then filtered and subjected to ultra performance liquid chromatography (UPLC)-MS/MS analysis using a Waters Xevo TQ-S-ESI tandem quadrupole MS operated in the positive ion mode and coupled to a BEH Shield RP18 chromatography column. A 10-mM formic acid/water/acetonitrile (ACN) mobile phase was used for analyte separation. The mass transitions for quantitation of the analytes were: 471.4 > 183.2 for AA, 479.4 > 183.2 for AA-d8, 469.4 > 183.2 for EPA, 474.4 > 183.2 for EPA-d5, 495.4 > 183.2 for DHA, and 501.4 > 183.2 for DHA-d5. Calibration curve samples were prepared from stock solutions of authentic EPA, DHA, and AA standards in ethyl acetate and processed in the same manner as diluted RBCs. Data were collected using MassLynx, and peak areas were integrated using QuanLynx software. The inter-day accuracy (percentage error) and precision (percentage deviation) across a range of AA, EPA, and DHA concentrations were all <8%. Intra-day assay precision (percentage error) and accuracy (percentage deviation) for replicate measurement of a range of AA, EPA, and DHA concentrations were all <15%.
Clinical laboratory measurements
Fasting and 2-hour oral glucose tolerance test samples were collected in sodium fluoride tubes, and plasma derived from these samples was analyzed on a Roche Hitachi 704 Chemistry Analyzer by the Lipoprotein Analytical Core (LAC) at OHSU or on a Siemens Dimension Xpand analyzer at the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) Phoenix Branch. Reagents used at the LAC were from Randox (Cat# GL1611), and the reagents used at the NIDDK included the Siemens Dimension Glucose Flex Cartridge (DF40-10444971) and the Siemens Chem I Calibrator (DC18C-10716280). The intra- and inter-assay coefficients from the LAC were reported by Randox as 3.7% and 5.6%, respectively. The intra- and inter-assay coefficients of variation from the NIDDK were 0.22% and 0.5%, respectively.
Plasma concentrations of insulin were measured by the Endocrine Technologies Core (ETC) at the Oregon National Primate Research Center (ONPRC) and by the University of California Davis (UCD). Insulin was measured using an RIA from Millipore Sigma at UCD, and the intra- and inter-assay CVs were 5.9% and 7.6%, respectively. Concentrations of insulin in human plasma were determined in the ETC using a chemiluminescence-based clinical automatic immunoassay platform (Roche Cobas e411, Roche Diagnostics). Intra- and inter-assay variations in the ETC were <7%. Using fasting glucose and insulin values, we then calculated the HOMA2-B, HOMA2-S, HOMA2-IR, and HOMA2-DI (the product of insulin sensitivity and β-cell function) were calculated by the HOMA2 method (28).
Plasma samples were analyzed for immune biomarkers in the ETC and the ONPRC using a 38-plex human cytokine Luminex panel (Millipore Sigma). Cytokines, chemokines, and growth factors included are shown in Supplemental Table 1. This panel included 15 proinflammatory cytokines, 10 proinflammatory chemokines, 6 anti-inflammatory cytokines, and 7 growth factors. Data were calculated using Milliplex Analyst software version 5.1 (EMD Millipore). Vendor-supplied quality control samples and an in-house human serum pool were analyzed to monitor assay variation. The percentages of samples below the lower limits of quantification (LLOQs) and inter-assay CVs are also shown in Supplemental Table 1. A total of 20 cytokines assayed were excluded from the statistical analysis because they either had more than 30% of the samples below the detection level threshold or an inter-assay CV > 15%.
Plasma RANTES concentrations were determined by ELISA by the ETC following the manufacturer's instructions (R&D Systems). Plasma was diluted at 1:100 for analysis. The assay range, when adjusted for dilution, was 3130 to 200,000 pg/ml. Intra- and inter-assay CVs for an in-house human serum pool was 3.4%.
Additionally, IL-6, hsCRP, and adiponectin were measured in sodium citrate plasma. IL-6 was measured with an ELISA kit (R&D Systems), and the range of the assay was 3.1 to 300 pg/ml. The IL-6 inter-assay CV was 10.6%, and the intra-assay CV ranged between 1.6% and 4.2%. A Randox assay was used to measure hsCRP, and the assay range was 0.477–10.0 mg/l, with a reported inter-assay CV range between 1.22% and 4.44% and an intra-assay CV range between 1.21% and 2.66%. A Randox assay was also used to measure adiponectin (Cat# AO8154). Randox reports the adiponectin intra- and inter-assay coefficients as 1.6% and 2.5%, respectively, and the assay range is 0.32 to 23.8 mg/ml.
Statistical analysis
All analyses were performed using the R statistical programming language, version 3.6.1 (R Studio). Outcome variables were transformed to approximate normality using either a power transform or a logarithmic transform. After applying the transform, values that were more than 2 times the IQR above the third quartile or below the first quartile were classified as outliers and dropped from the analysis. When appropriate, 1 of 3 transforms was used: no transformation (i.e., power = 1), square root transformation (i.e., power = 0.5), or a logarithmic transform (represented as power = 0). A different power was chosen only if it resulted in a variable with much better skewness and kurtosis than could be achieved by 1 of these 3 transforms, or if fewer transformed values were outliers or both. After transforming the variable, it was multiplied by a constant to make its variance close to 1. Supplemental Table 2 shows the powers used, the multipliers, and the number of resulting outliers, as well as the skewness, kurtosis, minimum, and maximum of the transformed data. Age, PUFA, adiposity, blood pressure, and physical activity variables were transformed using a rank-inverse-normal transformation. P values for comparisons of raw data in males compared with females and quartile 1 compared with quartile 4 of NIR were based on t-tests for the normalized variables (Table 1). We tested whether the proportion of data below the LLOQ differed between extremes of NIR using Fisher's exact test (Table 2). Continuous variables that were used only as predictors (age, measures of adiposity, measures of PUFA levels) in the linear models were not transformed, except for EPA levels, which were logarithm transformed because of their extreme skewness.
TABLE 1.
Demographic and health-related risk factors in Yup'ik participants by gender and by quartiles of n-3 PUFA intake1
| Variable | Male n = 92 | Female n = 73 | P value2 | Quartile 1 NIR n = 78 | Quartile 4 NIR n = 87 | P value |
|---|---|---|---|---|---|---|
| Age, y | 33.5 (32.2) | 40.0 (31.0) | 0.323 | 25.5 (13.0) | 54.0 (26.0) | <0.001 |
| NIR, ‰ | 7.4 (2.9) | 9.8 (3.4) | 7.1 (0.5) | 10.3 (1.1) | ||
| DHA, ng/mg protein | 225.5 (145.2) | 279.6 (129.1) | <0.001 | 167.9 (68.7) | 308.5 (75.6) | <0.001 |
| EPA, ng/mg protein | 45.7 (107.6) | 112.2 (128.9) | 0.012 | 26.1 (19.6) | 153.5 (85.5) | <0.001 |
| AA, ng/mg protein | 321.5 (79.8) | 342.9 (105.5) | 0.261 | 371.0 (83.1) | 302.8 (79.5) | <0.001 |
| DHA + EPA, ng/mg protein | 280.9 (274.8) | 420.2 (258.0) | <0.001 | 187.7 (94.4) | 467.7 (139.0) | <0.001 |
| BMI, kg/m2 | 23.9 (4.1) | 28.3 (7.8) | <0.001 | 24.1 (7.5) | 25.6 (6.5) | 0.450 |
| Waist circumference, cm | 82.7 (12.6) | 88.8 (21.4) | 0.053 | 82.3 (16.5) | 87.5 (16.5) | 0.038 |
| Percent body fat, % | 17.1 (8.3) | 35.8 (12.5) | <0.001 | 19.9 (18.8) | 27.0 (16.2) | 0.010 |
| Systolic blood pressure, mm Hg | 113.2 (16.2) | 100.3 (15.6) | <0.001 | 108.4 (17.6) | 104.0 (21.4) | 0.614 |
| Diastolic blood pressure, mm Hg | 67.7 (12.2) | 67.4 (12.4) | 0.921 | 67.6 (12.4) | 67.5 (12.2) | 0.158 |
| Glucose, mg/dL | 97.0 (10.6) | 97.6 (11.8) | 0.791 | 97.0 (10.1) | 98.7 (11.5) | 0.839 |
| HbA1c, % | 5.2 (0.3) | 5.2 (0.3) | 0.955 | 5.2 (0.4) | 5.2 (0.3) | 0.635 |
| 2-hour glucose, mg/dL | 65.8 (42.2) | 83.7 (36.3) | <0.001 | 74.8 (31.0) | 75.8 (36.2) | 0.713 |
| Insulin, μU/mL | 11.6 (7.4) | 14.9 (8.5) | 0.018 | 12.7 (9.1) | 13.6 (7.7) | 0.830 |
| Adiponectin,3 μg/mL | 8.0 (6.1) | 9.7 (5.3) | 0.054 | 8.2 (5.8) | 9.2 (5.8) | 0.605 |
| HOMA2-B | 104.3 (34.7) | 116.5 (34.9) | 0.012 | 109.4 (42.9) | 107.1 (36.0) | 0.557 |
| HOMA2-S | 75.4 (48.3) | 58.3 (33.1) | 0.015 | 66.7 (46.4) | 63.8 (36.0) | 0.968 |
| HOMA2-IR | 1.5 (1.0) | 2.0 (1.1) | 0.017 | 1.7 (1.2) | 1.8 (1.0) | 0.887 |
| HOMA2-DI | 7117.7 (2650.0) | 6634.5 (2231.5) | 0.107 | 6893.3 (2821.4) | 6692.2 (2131.8) | 0.746 |
| hsCRP,3 mg/L | 0.4 (1.3) | 0.4 (0.8) | 0.787 | 0.4 (0.9) | 0.4 (1.0) | 0.951 |
| IL-6,3 pg/mL | 1.1 (2.1) | 1.5 (2.2) | 0.459 | 1.4 (2.5) | 1.1 (1.9) | 0.596 |
| RANTES,3 ng/mL | 44.7 (36.6) | 31.5 (28.3) | 0.014 | 41.7 (37.9) | 36.9 (32.7) | 0.487 |
| Sedentary time (% awake wear time) | 16.1 (9.8) | 13.5 (10.0) | 0.295 | 13.9 (9.4) | 15.9 (10.9) | 0.330 |
| MVPA (% awake wear time) | 18.0 (7.8) | 14.0 (3.3) | <0.001 | 15.0 (7.2) | 15.0 (5.6) | 0.930 |
| Total Physical Activity (VMU/min) | 80.6 (24.3) | 73.4 (12.0) | <0.001 | 74.7 (22.0) | 76.3 (13.8) | 0.878 |
| Triglycerides, mg/dL | 72.5 (31.0) | 77.7 (46.2) | 0.082 | 74.0 (48.3) | 73.9 (32.2) | 0.625 |
| Total cholesterol, mg/dL | 192.2 (58.2) | 214.5 (57.7) | <0.001 | 179.7 (48.6) | 222.7 (45.7) | <0.001 |
| LDL cholesterol, mg/dL | 113.7 (50.7) | 133.3 (39.6) | 0.002 | 107.6 (42.2) | 137.1 (41.8) | <0.001 |
| HDL cholesterol, mg/dL | 58.1 (19.5) | 69.2 (24.0) | 0.024 | 55.5 (21.0) | 68.6 (21.8) | <0.001 |
| non-HDL cholesterol, mg/dL | 130.4 (49.4) | 148.7 (42.8) | 0.002 | 124.6 (48.2) | 154.2 (41.7) | <0.001 |
Values are reported as medians (IQRs). Blood-derived measures were assayed using plasma samples. AA, arachidonic acid; HbA1c, glycated hemoglobin; HOMA2-B, homeostasis model assessment of beta cell function; HOMA2-DI, homeostasis model assessment of disposition index; HOMA2-S, homeostasis model assessment of insulin sensitivity; hsCRP, high-sensitivity C-reactive protein; MVPA, moderate to vigorous physical activity; NIR, nitrogen isotope ratio; RANTES, regulated upon activation, normal T-cell expressed and secreted; VMU, vector magnitude units.
The P values from the t-test are based on transformed data. Significant differences between men and women or between the lowest or highest quartiles (quartiles 1 and 4, respectively) of the NIR, an objective biomarker of n-3 PUFA intake, are indicated.
Adipokines and cytokines were measured by ELISA. Age, PUFA, adiposity, blood pressure, and physical activity variables were transformed using a rank-inverse-normal transformation. All the outcome variables were transformed using Power Transformation.
TABLE 2.
Luminex panel characteristics: proportion of samples below the LLOQ overall and in the lowest and highest quartiles (quartiles 1 and 4, respectively) of n-3 PUFA intake among Yup'ik participants1
| % Sample < LLOQ3 | Quartile 1 (n = 78) | Quartile 4 (n = 87) | P value2 | |
|---|---|---|---|---|
| Proinflammatory Cytokines | ||||
| IFN-α2 | 56.97 | 39 (50.0) | 55 (63.2) | 0.115 |
| IFN-γ | 40.61 | 27 (34.6) | 40 (46.0) | 0.155 |
| IL-12p40 | 60.61 | 40 (51.3) | 60 (69.0) | 0.0262 |
| IL-12p70 | 55.76 | 35 (44.9) | 57 (65.5) | 0.0122 |
| IL-15 | 11.52 | 8 (10.3) | 11 (12.6) | 0.808 |
| sCD40L | 0.00 | 0 (0.0) | 0 (0.0) | 1.000 |
| IL-17A | 84.85 | 63 (80.8) | 77 (88.5) | 0.195 |
| IL-1α | 66.06 | 43 (55.1) | 66 (75.9) | 0.0052 |
| IL-1β | 74.55 | 54 (69.2) | 69 (79.3) | 0.155 |
| IL-2 | 97.58 | 77 (98.7) | 84 (96.6) | 0.623 |
| IL-3 | 100.00 | 78 (100.0) | 87 (100.0) | 1.000 |
| IL-6 | 60.00 | 38 (48.7) | 61 (70.1) | 0.0072 |
| IL-7 | 33.94 | 22 (28.2) | 34 (39.1) | 0.188 |
| TNFα | 0.00 | 0 (0.0) | 0 (0.0) | 1.000 |
| TNFβ | 52.12 | 31 (39.7) | 55 (63.2) | 0.0032 |
| Proinflammatory Chemokines | ||||
| Eotaxin-1 | 0.00 | 0 (0.0) | 0 (0.0) | 1.000 |
| GRO | 9.09 | 3 (3.8) | 12 (13.8) | 0.0312 |
| MCP-3 | 13.94 | 7 (9.0) | 16 (18.4) | 0.114 |
| IL-8 | 1.21 | 0 (0.0) | 2 (2.3) | 0.498 |
| MDC | 0.00 | 0 (0.0) | 0 (0.0) | 1.000 |
| IP-10 | 0.00 | 0 (0.0) | 0 (0.0) | 1.000 |
| MCP-1 | 0.00 | 0 (0.0) | 0 (0.0) | 1.000 |
| MIP-1α | 50.91 | 35 (44.9) | 49 (56.3) | 0.162 |
| MIP-1β | 0.00 | 0 (0.0) | 0 (0.0) | 1.000 |
| Fractalkine | 26.06 | 16 (20.5) | 27 (31.0) | 0.156 |
| Anti-Inflammatory Cytokines | ||||
| IL-10 | 15.15 | 7 (9.0) | 18 (20.7) | 0.0492 |
| IL-13 | 64.85 | 43 (55.1) | 64 (73.6) | 0.0152 |
| IL-1RA | 7.27 | 5 (6.4) | 7 (8.0) | 0.770 |
| IL-9 | 69.70 | 47 (60.3) | 68 (78.2) | 0.0172 |
| IL-4 | 18.79 | 11 (14.1) | 20 (23.0) | 0.166 |
| IL-5 | 34.5 | 24 (30.8) | 33 (37.9) | 0.413 |
| Growth Factors/Adipokines | ||||
| EGF | 0.60 | 0 (0.0) | 1 (1.1) | 1.000 |
| FGF-2 | 8.43 | 2 (2.6) | 11 (12.6) | 0.0202 |
| TGFα | 86.75 | 66 (84.6) | 77 (88.5) | 0.499 |
| G-CSF | 33.73 | 20 (25.6) | 36 (41.4) | 0.0482 |
| Flt-3L | 38.55 | 26 (33.3) | 37 (42.5) | 0.262 |
| GM-CSF | 22.29 | 12 (15.4) | 24 (27.6) | 0.062 |
| VEGF | 51.20 | 33 (42.3) | 51 (58.6) | 0.0432 |
Panel characteristics for 165 samples from the lower and upper quartiles of NIR (an objective biomarker of n-3 PUFA intake) with the number (%) of individuals below the LLOQ in each quartile. EGF, epidermal growth factor; FGF, fibroblast growth factor; Flt-3L, Fms-related tyrosine kinase 3 ligand; Fms, Feline McDonough Sarcoma; G-CSF, granulocyte colony-stimulating factor; GM-CSF, Granulocyte-macrophage colony-stimulating factor; GRO, growth related cytokine; IP, interferon γ-induced protein; LLOQ, lower limit of quantification; MCP, Monocyte chemoattractant protein-1; MDC, Macrophage-derived chemokine; MIP, macrophage inflammatory protein; NIR, nitrogen isotope ratio; sCD40L, soluble cluster of differentiation 40 ligand; TGF, transforming growth factor; VEGF, vascular endothelial growth factor.
Fisher exact test P values are shown, and significant differences between quartiles are indicated.
Number (%) samples below the LLOQ in each quartile for a given biomarker.
All linear regression models included sex (coded 0 for male and 1 for female), age, coastal (dichotomous variable indicating whether the community was coastal or inland), smoking (coded as 0 for no cigarette use and 1 for any cigarette use), and total physical activity (mean vector magnitude per minute) while awake from a triaxial accelerometer (ActiGraph GT3X, ActiGraph Corp, USA) worn for 2 days. We used community location as a dichotomous covariate to account for the population substructure between coastal and tundra communities, as previously described (38). In addition, when the cytokines were outcomes, we included statin, blood pressure, and anti-inflammatory medication use as covariates. NIR was coded as a dichotomous variable when used as a predictor in the models. Bonferroni's post hoc correction was used to adjust for multiple comparisons. Because of the correlation among variables, the effective number of tests was less than the actual number of tests. The method of Li and Ji (39) was used to derive the effective number of tests from the eigenvalues of the correlation matrix to determine the appropriate Bonferroni correction, 0.05/(effective number) (40). For the sensitivity analysis, we used permutation tests for all association tests.
Results
Demographic and health-related risk factors
Demographic characteristics of the 165 sub-sample participant group (upper and lower quartiles of RBC NIRs from a larger cohort) included in all subsequent analyses are shown in Table 1. Data are presented by gender, as well as by the NIR quartile, where quartile 1 represents a low n-3 PUFA intake and quartile 4 represents a high n-3 PUFA intake. Although the overall mean ages for male and female participants were similar, individuals in quartile 4 were significantly older than those in quartile 1. In quartile 1, there were 31 females (39.7%), compared to 42 females (48.3%) in quartile 4. As expected, all n-3 PUFA measures were greater in quartile 4 compared to quartile 1, whereas AA (a n-6 PUFA) values were greater in quartile 1 compared to quartile 4. Women had significantly higher DHA, EPA, total n-3 PUFA (DHA + EPA), and NIR values compared to men.
Women also had significantly higher BMIs and percentages of body fat, as well as higher insulin, adiponectin, HOMA2-B, and HOMA2-IR values, but lower HOMA2-S values, compared to men. Individuals in quartile 4 had higher WCs and percentages of body fat compared to individuals in quartile 1. Men were more active than women, and had higher systolic blood pressure measurements. Additionally, both women (compared to men) and individuals in quartile 4 (compared with quartile 1) had higher total cholesterol, LDL, HDL, and non-HDL cholesterol.
Luminex panel characteristics
Characteristics and descriptions of the 38 immune biomarkers included in the Luminex cytokine panel are shown in Supplemental Table 1. Biomarker data were not analyzed further if more than 30% of the samples assayed were below the LLOQ. This resulted in the elimination of 12 proinflammatory cytokines (IFN-α2, IFN-γ, IL-12p40, IL-12p70, IL-17A, IL-1α, IL-1β, IL-2, IL-3, IL-6, IL-7, and TNFβ); 1 proinflammatory chemokine [macrophage inflammatory protein (MIP)-1α], 3 anti-inflammatory cytokines (IL-13, IL-9, and IL-5), and 4 growth factors [transforming growth factor α, granulocyte colony-stimulating factor (G-CSF), FLT-3L, and vascular endothelial growth factor (VEGF)]. IL-6 was also measured using an ELISA assay, and only 69 of the 409 samples assayed were below the LLOQ, so IL-6 was included in further analyses.
For the remaining cytokines and chemokines subjected to statistical analysis, individual values below the LLOQ were excluded. To determine whether there were more inflammatory cytokines and chemokines eliminated from quartile 4 than quartile 1, we evaluated the number of samples below the LLOQ in each group (Table 2). In every case where the number of samples below the LLOQ was significantly different between quartile 1 and quartile 4 [IL-12p40, IL-12p70, IL-1α, IL-6, TNFβ, growth related cytokine (GRO), IL-10, IL-13, IL-9, fibroblast growth factor 2 (FGF-2), G-CSF, and VEGF], we noted more samples below the LLOQ in quartile 4.
Association of PUFAs with immune biomarkers
The associations between individual RBC PUFA levels, which derive from marine fats, and NIR, which derives from marine protein and inflammatory biomarkers, are shown in Table 3 and Supplemental Table 3. DHA was inversely associated with IL-6; EPA was inversely associated with TNFα; and the NIR was inversely associated with Monocyte chemoattractant protein-1 (MCP-1) and IL-10. All of these associations remained significant after Bonferroni correction. We observed several suggestive associations: DHA was inversely associated with Granulocyte-macrophage colony-stimulating factor (GM-CSF), TNFα, eotaxin-1, IL-8, Macrophage-derived chemokine (MDC), RANTES, IL-1RA, IL-10, and Epidermal growth factor (EGF); EPA was inversely associated with IL-10; the sum of EPA and DHA was inversely associated with IL-6, TNFα, MCP-1, MDC, RANTES, and IL-10; and the NIR was inversely associated with GM-CSF, IL-6, IL-15, TNFα, eotaxin-1, fractalkine, IL-8, MDC, and FGF-2. In comparison, AA was inversely associated with MDC and RANTES, and positively associated with MCP-1.
TABLE 3.
Association between immune biomarkers and PUFAs in Yup'ik participants1
| PUFA variable | β (SE) | P value2 | |
|---|---|---|---|
| Proinflammatory Cytokines | |||
| GM-CSF | DHA | –0.0034 (0.0014) | 0.019 |
| GM-CSF | NIR | –0.6856 (0.2511) | 0.007 |
| IL-6 | DHA | –0.0066 (0.0017) | <0.0012 |
| IL-6 | EPA + DHA | –0.0022 (0.0010) | 0.023 |
| IL-6 | NIR | –0.7580 (0.2907) | 0.010 |
| IL-15 | NIR | –0.6643 (0.2986) | 0.028 |
| TNFα | DHA | –0.0045 (0.0016) | 0.005 |
| TNFα | EPA | –0.4925 (0.1392) | <0.0012 |
| TNFα | EPA + DHA | –0.0027 (0.0009) | 0.003 |
| TNFα | NIR | –0.7104 (0.2797) | 0.012 |
| Proinflammatory Chemokines | |||
| Eotaxin-1 | DHA | –0.0071 (0.0024) | 0.003 |
| Eotaxin-1 | NIR | –1.0782 (0.4268) | 0.013 |
| Fractalkine | NIR | –1.0558 (0.4381) | 0.017 |
| IL-8 | DHA | –0.0056 (0.0018) | 0.002 |
| IL-8 | NIR | –0.7381 (0.3255) | 0.025 |
| MCP-1 | AA | 0.0043 (0.0015) | 0.004 |
| MCP-1 | EPA + DHA | –0.0017 (0.0008) | 0.036 |
| MCP-1 | NIR | –0.8345 (0.2410) | <0.0012 |
| MDC | DHA | –0.0073 (0.0023) | 0.001 |
| MDC | AA | –0.0051 (0.0024) | 0.037 |
| MDC | EPA + DHA | –0.0034 (0.0013) | 0.010 |
| MDC | NIR | –1.0710 (0.3935) | 0.007 |
| RANTES | DHA | –0.0048 (0.0017) | 0.004 |
| RANTES | AA | –0.0043 (0.0018) | 0.017 |
| RANTES | EPA + DHA | –0.0020 (0.0010) | 0.041 |
| Anti-Inflammatory Cytokines | |||
| IL-1RA | DHA | –0.0031 (0.0014) | 0.028 |
| IL-10 | DHA | –0.0056 (0.0019) | 0.003 |
| IL-10 | EPA | –0.4537 (0.1684) | 0.008 |
| IL-10 | EPA + DHA | –0.0034 (0.0011) | 0.002 |
| IL-10 | NIR | –1.2868 (0.3360) | <0.0012 |
| Growth Factors | |||
| EGF | DHA | –0.0050 (0.0023) | 0.029 |
| FGF-2 | NIR | –0.9633 (0.4397) | 0.030 |
Twenty immune biomarkers in the Luminex panel were excluded from this analysis because >30% of the values were below the LLOQ (see Supplemental Table 1). All linear regression models included sex, age, smoking/tobacco use, physical activity, coastal compared with inland location, statin, blood pressure, and inflammatory medication as covariates. Quartiles 1 and 4 included 78 and 87 participants, respectively, corresponding to the lowest NIR and highest quartile of NIR measurement from the overall sample of 569 participants. Column 2 indicates the PUFA associated with a particular cytokine. AA, arachidonic acid; EGF, epidermal growth factor; FGF, fibroblast growth factor; GM-CSF, Granulocyte-macrophage colony-stimulating factor; LLOQ, lower limit of quantification; MCP, Monocyte chemoattractant protein-1; MDC, Macrophage-derived chemokine; NIR, nitrogen isotope ratio; RANTES, regulated upon activation, normal T-cell expressed and secreted.
P values were statistically significant after Bonferroni adjustment for multiple testing [Bonferroni threshold for significance: 0.05/(3 × 16) = 0.00104]. The effective number of independent PUFA variables equaled 3, and the effective number of independent immune biomarkers determined by spectral decomposition (39) equaled 16. β values are slopes of the regression models.
In addition, we performed permutation tests, and the empirical P values are provided in Supplemental Table 3. Note that the empirical P values were lower than the regression P values, with some exceptions. For example, IL-8 and MDC associations with DHA became significant with Bonferroni correction in the permutation tests. However, the empirical P value of the association test between NIR and MCP-1 was no longer significant with the Bonferroni correction. In general, the empirical P values were lower than the regression P values.
Correlations of immune biomarkers and clinical measures
All of the adiposity variables (BMI, WC, and percentage body fat) were positively correlated with glucose, insulin, 2-hour glucose, HOMA2-B, and HOMA2-IR, and negatively correlated with HOMA2-DI (Figure 1). Some of the adiposity variables were also positively correlated with immune biomarkers, including IL-6, hsCRP, interferon γ-induced protein (IP)-10, MCP-1, MIP-1β, and IL-1RA, and negatively correlated with IL-15, soluble cluster of differentiation 40 ligand (sCD40L), eotaxin-1, MCP-3, IL-8, MDC, fractalkine, RANTES, IL-10, IL-4, EGF, FGF-2, and GM-CSF. Adiponectin was only negatively correlated with WC.
FIGURE 1.
Spearman correlation coefficients between PUFAs, diabetes, and cardiovascular risk factors, and inflammatory immune biomarkers in Yup'ik participants. The red and blue colors indicate the positive and negative correlation coefficient value intensities, respectively. *P < 0.05; **statistically significant after Bonferroni adjustment for multiple testing. Abbreviations for all the immune biomarkers are listed in Supplemental Table 1.
Similarly, 1 or more of the n-3 PUFAs (EPA, DHA, EPA + DHA, and NIR) were negatively correlated with proinflammatory cytokines (IL-15 and sCD40L) and proinflammatory chemokines (eotaxin-1, MCP-3, MDC, fractalkine, and RANTES); however, n-3 PUFAs were positively correlated with IP-10 and adiponectin.
Patterns of correlations between immune biomarkers and glucose parameters representing insulin resistance and a predisposition to T2D (glucose, insulin, HbA1c, 2-hour glucose, HOMA2-B, and HOMA2-IR) were generally weak and also inconsistent. Exceptions included IL-6, hsCRP, IP-10, and MCP-1, which positively correlated with 1 or more of the prediabetes parameters. Similarly, IL-15, MCP-3, MDC, fractalkine, IL-10, and IL-4 were negatively correlated with 1 or more prediabetes parameters.
Association between WC and parameters of glucose metabolism
WC was strongly associated with several glucose parameters, including fasting glucose, insulin, and all HOMA2 measures (Table 4). A suggestive positive association was observed between WC and HbA1c, but not 2-hour glucose. Similar associations were observed between BMI and glucose parameters (data not shown). A similar trend of the empirical P values was observed with the permutation test.
TABLE 4.
Association between parameters of glucose metabolism and waist circumference in Yup'ik participants
| Glucose parameter (outcome) | Adiposity variable (predictor) | β (SE) | P value1 | Empirical P values2 |
|---|---|---|---|---|
| Glucose | WC | –0.02183 (0.0061) | <0.0011 | <0.0011 |
| Insulin | WC | 0.0434 (0.0089) | <0.0011 | <0.0011 |
| HbA1c | WC | –0.01443 (0.0056) | 0.011 | 0.018 |
| 2-hour glucose | WC | 0.0012 (0.0031) | 0.688 | 0.357 |
| HOMA2-B | WC | 0.0371 (0.0102) | <0.0011 | <0.0011 |
| HOMA2-S | WC | –0.0284 (0.0059) | <0.0011 | <0.0011 |
| HOMA2-IR | WC | 0.0297 (0.0060) | <0.0011 | <0.0011 |
| HOMA2-DI | WC | –0.0354 (0.0078) | <0.0011 | <0.0011 |
Association remained significant after multiple testing adjustment (Bonferroni threshold for significance: 0.05/5 = 0.01). The effective number of independent parameters of glucose metabolism, determined by spectral decomposition (39), equaled 5. All linear regression models included sex, age, smoking/tobacco use, physical activity, and coastal compared with inland location as covariates. HbA1c, glycated hemoglobin; HOMA2-B, homeostasis model assessment of beta cell function; HOMA2-DI, homeostasis model assessment of disposition index; HOMA2-S, homeostasis model assessment of insulin sensitivity; WC, waist circumference.
Empirical P values of the permutation test. β values are slopes of the regression models.
Transformations used to normalize glucose and HbA1c were decreasing functions, resulting in beta values with the opposite sign.
Association of WC with immune biomarkers
None of the immune biomarkers was significantly associated with WC after Bonferroni correction. Several suggestive positive associations were observed between WC and hsCRP and IP-10, and WC was inversely associated with adiponectin (Table 5). Similar associations were observed between BMI and immune biomarkers (data not shown). A similar trend of the empirical P values was observed with the permutation test, with the exception of the association between adiponectin and WC, which became significant with an empirical P value of 0.0027 after Bonferroni correction.
TABLE 5.
Association between immune biomarkers and waist circumference in Yup'ik participants
| Adiposity variable (predictor) | β (SE) | P value1 | Empirical P values2 | |
|---|---|---|---|---|
| Proinflammatory Cytokines & Biomarkers (Outcome) | ||||
| hsCRP | WC | 0.0188 (0.0082) | 0.0243 | 0.0093 |
| GM-CSF | WC | –0.0043 (0.0081) | 0.595 | 0.313 |
| IL-6 | WC | 0.0085 (0.0106) | 0.423 | 0.209 |
| IL-15 | WC | –0.0005 (0.0095) | 0.955 | 0.478 |
| TNFα | WC | 0.0035 (0.0089) | 0.695 | 0.361 |
| sCD40L | WC | –0.0059 (0.0106) | 0.577 | 0.295 |
| Proinflammatory Chemokines | ||||
| Eotaxin-1 | WC | –0.0151 (0.0136) | 0.270 | 0.158 |
| Fractalkine | WC | –0.0059 (0.0140) | 0.674 | 0.339 |
| GRO | WC | 0.0099 (0.0139) | 0.476 | 0.237 |
| IL-8 | WC | –0.0077 (0.0104) | 0.459 | 0.235 |
| IP-10 | WC | 0.0285 (0.0118) | 0.0173 | 0.0183 |
| MCP-1 | WC | 0.0116 (0.0078) | 0.141 | 0.084 |
| MCP-3 | WC | –0.0003 (0.0085) | 0.971 | 0.496 |
| MDC | WC | 0.0001 (0.0126) | 0.996 | 0.491 |
| MIP-1β | WC | 0.0086 (0.0067) | 0.205 | 0.105 |
| RANTES | WC | –0.0012 (0.0093) | 0.898 | 0.454 |
| Anti-Inflammatory Hormones & Cytokines | ||||
| Adiponectin | WC | –0.0967 (0.0367) | 0.0093 | 0.0031 |
| IL-1RA | WC | 0.0162 (0.0080) | 0.0453 | 0.0193 |
| IL-4 | WC | –0.0050 (0.0122) | 0.682 | 0.350 |
| IL-10 | WC | –0.0001 (0.0110) | 0.992 | 0.491 |
| Growth Factors | ||||
| EGF | WC | –0.0140 (0.0127) | 0.272 | 0.140 |
| FGF-2 | WC | –0.0272 (0.0140) | 0.053 | 0.0323 |
No associations remained significant after Bonferroni multiple testing correction. Bonferroni threshold for significance: 0.05/16 = 0.003125. The effective number of independent immune biomarkers, determined by spectral decomposition (39), equaled 16. EGF, epidermal growth factor; FGF, fibroblast growth factor; GM-CSF, Granulocyte-macrophage colony-stimulating factor; GRO, growth related cytokine; hsCRP, high-sensitivity C-reactive protein; IP, inflammatory protein; MCP, Monocyte chemoattractant protein-1; MIP, macrophage inflammatory protein; RANTES, regulated upon activation, normal T-cell expressed and secreted; sCD40L, soluble cluster of differentiation 40 ligand; WC, waist circumference.
Empirical P values of the permutation test. β values are slopes of the regression models.
P values < 0.05 but greater than the Bonferroni threshold for significance.
Association between immune biomarkers and parameters of glucose metabolism
None of the immune biomarkers were significantly associated with any glucose parameter after Bonferroni correction for multiple testing (Supplemental Table 4). However, suggestive positive associations were observed between glucose with fractalkine and MDC; HbA1c with IL-15, IL-4, and TNFα; and 2-hour glucose with MDC. Insulin and HOMA2-IR were positively associated with IP-10; HOMA2-S was inversely associated with IP10; and HOMA2-DI was positively associated with fractalkine and MDC in permutation testing.
Association between PUFAs and parameters of glucose metabolism
There were no significant associations between PUFAs and any of the glucose parameters, with or without Bonferroni correction (Supplemental Table 5).
Discussion
We report that the NIR (an objective blood biomarker of n-3 PUFA intake) and individually measured n-3 PUFAs (EPA and DHA) in RBC membranes were inversely associated with several proinflammatory cytokines and chemokines. Macrophage-activating CC-chemokines are associated with chronic inflammation (41). Since the NIR and DHA were inversely associated with several proinflammatory macrophage-activating CC-chemokines (MCP-1, MDC, RANTES, and eotaxin), our findings suggest that n-3 PUFAs may reduce monocyte recruitment and the accumulation of proinflammatory M1 macrophages and T cells in peripheral tissues (42–44). Individuals with high n-3 PUFA intakes also had correspondingly reduced AAs. Several different factors may explain this finding, including metabolic differences between individuals in quartile 1 compared with quartile 4 and differences in dietary patterns, and it is possible that high n-3 PUFA intakes may replace AAs in RBC membranes to further reduce inflammation by altering the pattern of eicosanoid production (45–47).
M1-activated macrophages, along with CD8 + T cells and CD4 + Th1 cells, secrete several proinflammatory cytokines, including TNFα, IL-6, IFN-γ, and IL-1β, resulting in systemic inflammation (42, 43). Decreased production of IL-6, TNFα, and IL-1β has been consistently observed following n-3 PUFA supplementation (48). We similarly observed significant inverse associations between EPA and TNFα and between DHA and IL-6. However, more than 30% of the samples assayed had IFN-γ and IL-1β below the LLOQ (Table 1); thus, these 2 cytokines were excluded from further analyses. Interestingly, we didn't observe an association between the n-3 PUFA intake and hsCRP, even though IL-6 was inversely associated with the n-3 PUFA intake and IL-6 regulates C-reactive protein synthesis in the liver. This may have resulted from our study population being only slightly overweight (Table 1), because we previously observed reductions in hsCRP in obese individuals consuming a diet rich in n-3 PUFAs (8).
Although IL-10 is considered an anti-inflammatory cytokine (49), the inverse association of NIR with IL-10 and weak inverse associations of EPA and DHA with IL-10 may be explained because n-3 PUFAs activate the β-arrestin 2 signaling pathway (9), which inhibits the release of NF-κB, a known activator of toll-like receptor-dependent IL-10 synthesis (50). Moreover, deacetylation of NF-kB by n-3 PUFAs downregulates its signaling, leading to suppression of downstream NF-kB-dependent gene expression (51). Additionally, activation of peroxisome proliferator–activated receptor-γ by n-3 PUFAs interferes with NF-κB translocation to the nucleus (48), further altering activation of cytokine gene expression. Our observation that several inflammatory cytokines and chemokines were inversely associated with the n-3 PUFA intake is consistent with the fact that they are known target genes of NF-κB (52, 53).
Despite our modest sample size and a study population that was only slightly overweight, many of the observed immune biomarker associations remained significant after Bonferroni correction for multiple testing, as well as multivariate adjustment for several potential confounders, including age, sex, smoking, physical activity, and coastal compared with inland location, which we have previously shown to be associated with n-3 PUFA intakes (54). The direction and magnitude of these associations were not modified by physical activity or smoking. Moreover, while we observed some significant associations specific to the NIR that were not significant for EPA and DHA (or vice versa), we frequently observed weak associations between EPA, DHA, or the NIR and a given immune biomarker with similar directionality. We have previously observed differences in chronic disease risk factor associations between the NIR and individual n-3 PUFAs (EPA and DHA), despite the fact that the NIR and n-3 PUFAs are highly correlated (16, 29). This may be due to the NIR capturing unmeasured nutrients associated with the lean and lipid portions of marine foods, whereas direct measures of EPA and DHA represent only the lipid component of the diet. Because the NIR is associated with the Yup'ik traditional dietary pattern, which contains a number of subsistence foods in addition to marine mammals and fish rich in n-3 PUFAs (55), the reduced inflammatory cytokines associated with the NIR may also be derived from unidentified food synergies (27) present in the subsistence diet. Because the NIR is a surrogate for n-3 PUFA intake over the past 3–4 months (35), it is possible that we are not capturing seasonal variations in n-3 PUFA intakes. However, previous studies measuring NIR in a segmental analysis of hair revealed that while the NIR in hair segments fluctuates seasonally in some individuals, the seasonal variations in traditional food intake were modest relative to the interindividual variation (56).
Since n-3 PUFAs were inversely associated with inflammation, and inflammation has been strongly linked to both insulin resistance and islet cell dysfunction (14), we hypothesized that n-3 PUFAs would be associated with improved glycemic parameters. While central adiposity was significantly associated with multiple adverse parameters of glucose metabolism, we only observed a small number of weak associations between WC and immune biomarkers, and we did not find any evidence that n-3 PUFA consumption was associated with parameters of glucose metabolism.
There are several strengths to this study, including the evaluation of n-3 PUFA consumption from whole foods rather than supplements, which allowed for the evaluation of food synergies (27) associated with habitual intake of marine mammals and fish, including micronutrients. Yup'ik people regularly consume a large and diverse array of fish, fish eggs, and marine mammals that provide a rich source of food-derived n-3 PUFAs (57–61). Moreover, we have reported on factors that characterize dietary patterns among Yup'ik people in our study communities (55, 60, 62). We were also able to select individuals at the extremes of n-3 PUFA intakes using a surrogate measure of n-3 PUFA intake (NIR), and then directly measure n-3 PUFAs in RBC membranes. Moreover, while the mean n-3 PUFA intake in this study population is more than 20 times higher than that in the US general population (15), the range in n-3 PUFA intakes is still large (16, 29). Our approach also allowed us an opportunity to compare associations between the lean (protein) component of foods rich in n-3 PUFAs (using the NIR) and the lipid component of these foods (using EPA, DHA, and AA measures) with a comprehensive array of cytokines, chemokines, adipokines, growth factors, and glucose parameters in the same population. Finally, we included information on important potential confounders (physical activity, smoking) in a population not taking n-3 PUFA supplements. While it is possible that other unmeasured dietary, behavioral, or lifestyle factors may have confounded our findings, an advantage to working in small Yup'ik communities is the homogenous nature of their surroundings: there are only 1–2 small markets per community, the variety of available market foods is extremely limited, and, for the most part, community members have similar socioeconomic statuses, educational levels, and housing within and between these small communities (see Supplemental Table 6). Limitations to this study include the lack of diet recall data, the modest sample size, limited sensitivity of the Luminex panel to detect low levels of circulating immune biomarkers, and our sample selection criteria based on the extremes of n-3 PUFA intakes without regard to obesity status, resulting in few obese participants. Although samples were initially stored at −20°C for up to a week because of our rural research locations, and might possibly lead to some oxidative degradation, it didn't affect the relationships between NIRs and n-3 PUFA levels in RBCs, which were very similar to those previously reported (29). Since the NIR measurement is not sensitive to alterations in molecular structure, this suggests that PUFA oxidation was minimal, random, and not likely to affect the relationships with other biomarkers that we report. Additionally, while male and female participants were age matched, participants in quartile 1 were significantly younger than participants in quartile 4. Finally, although the consistency of inverse associations across several inflammatory biomarkers is suggestive of causality, we were nevertheless limited in making causal conclusions by the cross-sectional study design.
In conclusion, in this population of Yup'ik Alaska Native adults consuming a diet rich in marine mammals and fish, a higher n-3 PUFA intake is associated with lower systemic inflammation in healthy adults, but not with better glycemic status, which was strongly influenced by body composition. Nevertheless, chronic low-grade inflammation is an important component of many different diseases and conditions (48). Our comprehensive evaluation of 41 pro- and anti-inflammatory cytokines, chemokines, adipokines, and growth factors expands the list of immune biomarkers potentially influenced by n-3 PUFAs. Additionally, evaluation of n-3 PUFA consumption using an objective biomarker (NIR) implies that the protein component of the traditional diet may contain important micronutrients not found in the lipid component of the diet, as previously observed (16). The results from this study may help to resolve some of the inconsistencies previously observed in randomized controlled trials conducted with pharmaceutical grade n-3 PUFAs or dietary supplements regarding the potential clinical benefit of consumption of foods rich in n-3 PUFAs to prevent inflammatory diseases (45, 48, 63). Future studies focused on the identification of n-3 PUFA–derived lipid mediators resulting from dietary patterns rich in marine oils may help to illuminate protective physiological mechanisms and lead to promising therapeutic dietary recommendations to prevent inflammatory diseases.
Supplementary Material
ACKNOWLEDGEMENTS
We thank Eliza Orr and Samantha Louey for their help with data collection. The authors’ responsibilities were as follows – BBB, HKT, SEH, KET: designed the research; BBB, HKT, SEH, KET, TA, DMO: conducted the research; HKT, HWW, JJP, CXZ: analyzed the data; BBB, HKT, HWW, TA, CXZ: drafted the paper; BBB: had primary responsibility for the final content; and all authors: contributed to, read, and approved the final manuscript.
Notes
This study was funded by grants from the NIH (R01DK104347, R01DK112358, R01HL146549, P01GM116691, P20RR016430, P30GM103325, and UL1TR002369). The Endocrine Technologies Core at the Oregon National Primate Research Center is supported by NIH Grant P51OD011092.
Author disclosures: The authors report no conflicts of interest.
Supplemental Figure 1 and Supplemental Tables 1–6 are available from the “Supplementary data” link in the online posting of the article and from the same link in the online table of contents at https://academic.oup.com/jn/.
Abbreviations used: AA, arachidonic acid; EGF, epidermal growth factor; ETC, Endocrine Technologies Core; FGF, fibroblast growth factor; GM-CSF, Granulocyte-macrophage colony-stimulating factor; HOMA2-B, homeostasis model assessment of beta cell function; HOMA2-DI, homeostasis model assessment of disposition index; HOMA2-S, homeostasis model assessment of insulin sensitivity; hsCRP, high-sensitivity C-reactive protein; IP, interferon γ-induced protein; LAC, Lipoprotein Analytical Core; LLOQ, lower limit of quantification; MCP, Monocyte chemoattractant protein-1; MIP, macrophage inflammatory protein; NIDDK, National Institute of Diabetes and Digestive and Kidney Diseases; NIR, nitrogen isotope ratio; OHSU, Oregon Health and Science University; ONPRC, Oregon National Primate Research Center; RANTES, regulated upon activation, normal T-cell expressed and secreted; sCD40L, soluble cluster of differentiation 40 ligand; T2D, type 2 diabetes; UAF, University of Alaska Fairbanks; UCD, University of California Davis; VEGF, vascular endothelial growth factor; WC, waist circumference.
Contributor Information
Bert B Boyer, Department of Obstetrics and Gynecology, Oregon Health & Science University, Portland, OR, USA.
Scarlett E Hopkins, Department of Obstetrics and Gynecology, Oregon Health & Science University, Portland, OR, USA.
Howard W Wiener, Department of Statistics, University of Alabama at Birmingham, Birmingham, AL, USA.
Jonathan Q Purnell, Department of Medicine, Oregon Health & Science University, Portland, OR, USA.
Diane M O'Brien, Department of Biology and Wildlife, University of Alaska Fairbanks, Fairbanks, AK, USA.
Cindy X Zhang, Department of Pharmaceutics, University of Washington, Seattle, WA, USA.
Joseph E Aslan, Department of Medicine, Oregon Health & Science University, Portland, OR, USA.
Theresa Aliwarga, Department of Pharmaceutics, University of Washington, Seattle, WA, USA.
Jeremy J Pomeroy, Clinical Research Center, Marshfield Clinic, Marshfield, WI, USA.
Kenneth E Thummel, Department of Pharmaceutics, University of Washington, Seattle, WA, USA.
Hemant K Tiwari, Department of Statistics, University of Alabama at Birmingham, Birmingham, AL, USA.
Data Availability
Data described in the manuscript, codebook, and analytic code will be made available upon request pending Tribal approval by the Yukon-Kuskokwim Health Corporation Executive Board.
References
- 1. Phillips CM, Chen LW, Heude B, Bernard JY, Harvey NC, Duijts L, Mensink-Bout SM, Polanska K, Mancano G, Suderman Met al. Dietary inflammatory index and non-communicable disease risk: A narrative review. Nutrients. 2019;11(8):1873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Hotamisligil GS. Inflammation and metabolic disorders. Nature. 2006;444(7121):860–7. [DOI] [PubMed] [Google Scholar]
- 3. Hotamisligil GS, Shargill NS, Spiegelman BM. Adipose expression of tumor necrosis factor-alpha: Direct role in obesity-linked insulin resistance. Science. 1993;259(5091):87–91. [DOI] [PubMed] [Google Scholar]
- 4. Hotamisligil GS. Inflammation, metaflammation and immunometabolic disorders. Nature. 2017;542(7640):177–85. [DOI] [PubMed] [Google Scholar]
- 5. Hopkins SE, Austin MA, Metzger JS, Koller KR, Umans JG, Kaufmann C, Wolfe AW, Howard BV, Boyer BB. Sex differences in obesity prevalence and cardiometabolic factors among Western Alaska native people. Nutr Metab Cardiovasc Dis. 2015;25(3):312–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Koller KR, Metzger JS, Jolly SE, Umans JG, Hopkins SE, Kaufmann C, Wilson AS, Ebbesson SO, Raymer TW, Austin MAet al. Cardiometabolic correlates of low type 2 diabetes incidence in Western Alaska native people–The WATCH study. Diabetes Res Clin Pract. 2015;108(3):423–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Wang L, Li X, Wang Z, Bancks MP, Carnethon MR, Greenland P, Feng YQ, Wang H, Zhong VW.. Trends in prevalence of diabetes and control of risk factors in diabetes among US adults, 1999–2018. JAMA. 2021;326(8):1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Makhoul Z, Kristal AR, Gulati R, Luick B, Bersamin A, O'Brien D, Hopkins SE, Stephensen CB, Stanhope KL, Havel PJet al. Associations of obesity with triglycerides and C-reactive protein are attenuated in adults with high red blood cell eicosapentaenoic and docosahexaenoic acids. Eur J Clin Nutr. 2011;65(7):808–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Yang Q, Vijayakumar A, Kahn BB. Metabolites as regulators of insulin sensitivity and metabolism. Nat Rev Mol Cell Biol. 2018;19(10):654–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Calder PC. Omega-3 fatty acids and inflammatory processes: From molecules to man. Biochem Soc Trans. 2017;45(5):1105–15. [DOI] [PubMed] [Google Scholar]
- 11. Ferrucci L, Cherubini A, Bandinelli S, Bartali B, Corsi A, Lauretani F, Martin A, Andres-Lacueva C, Senin U, Guralnik JM. Relationship of plasma polyunsaturated fatty acids to circulating inflammatory markers. J Clin Endocrinol Metab. 2006;91(2):439–46. [DOI] [PubMed] [Google Scholar]
- 12. So J, Wu D, Lichtenstein AH, Tai AK, Matthan NR, Maddipati KR, Lamon-Fava S. EPA and DHA differentially modulate monocyte inflammatory response in subjects with chronic inflammation in part via plasma specialized pro-resolving lipid mediators: A randomized, double-blind, crossover study. Atherosclerosis. 2021;316:90–8. [DOI] [PubMed] [Google Scholar]
- 13. Feskens EJ, Bowles CH, Kromhout D. Inverse association between fish intake and risk of glucose intolerance in normoglycemic elderly men and women. Diabetes Care. 1991;14(11):935–41. [DOI] [PubMed] [Google Scholar]
- 14. Lombardo YB, Chicco AG. Effects of dietary polyunsaturated n-3 fatty acids on dyslipidemia and insulin resistance in rodents and humans. A review. J Nutr Biochem. 2006;17(1):1–13. [DOI] [PubMed] [Google Scholar]
- 15. Makhoul Z, Kristal AR, Gulati R, Luick B, Bersamin A, Boyer B, Mohatt GV. Associations of very high intakes of eicosapentaenoic and docosahexaenoic acids with biomarkers of chronic disease risk among Yup'ik Eskimos. Am J Clin Nutr. 2010;91(3):777–85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. O'Brien DM, Kristal AR, Nash SH, Hopkins SE, Luick BR, Stanhope KL, Havel PJ, Boyer BB. A stable isotope biomarker of marine food intake captures associations between n-3 fatty acid intake and chronic disease risk in a Yup'ik study population, and detects new associations with blood pressure and adiponectin. J Nutr. 2014;144(5):706–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Beaulieu-Jones BR, O'Brien DM, Hopkins SE, Moore JH, Boyer BB, Gilbert-Diamond D. Sex, adiposity, and hypertension status modify the inverse effect of marine food intake on blood pressure in Alaska native (Yup'ik) people. J Nutr. 2015;145(5):931–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Au NT, Reyes M, Boyer BB, Hopkins SE, Black J, O'Brien D, Fohner AE, Yracheta J, Thornton T, Austin MAet al. Dietary and genetic influences on hemostasis in a Yup'ik Alaska native population. PLoS One. 2017;12(4):e0173616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Weisberg SP, McCann D, Desai M, Rosenbaum M, Leibel RL, Ferrante AW Jr. Obesity is associated with macrophage accumulation in adipose tissue. J Clin Invest. 2003;112(12):1796–808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Zatterale F, Longo M, Naderi J, Raciti GA, Desiderio A, Miele C, Beguinot F. Chronic adipose tissue inflammation linking obesity to insulin resistance and type 2 diabetes. Front Physiol. 2019;10:1607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Donath MY, Dinarello CA, Mandrup-Poulsen T. Targeting innate immune mediators in type 1 and type 2 diabetes. Nat Rev Immunol. 2019;19(12):734–46. [DOI] [PubMed] [Google Scholar]
- 22. Donath MY, Shoelson SE. Type 2 diabetes as an inflammatory disease. Nat Rev Immunol. 2011;11(2):98–107. [DOI] [PubMed] [Google Scholar]
- 23. Donath MY, Meier DT, Böni-Schnetzler M. Inflammation in the pathophysiology and therapy of cardiometabolic disease. Endocr Rev. 2019;40(4):1080–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Xun P, He K. Fish consumption and incidence of diabetes: Meta-analysis of data from 438,000 individuals in 12 independent prospective cohorts with an average 11-year follow-up. Diabetes Care. 2012;35(4):930–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Akinkuolie AO, Ngwa JS, Meigs JB, Djoussé L. Omega-3 polyunsaturated fatty acid and insulin sensitivity: A meta-analysis of randomized controlled trials. Clin Nutr. 2011;30(6):702–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Wu JHY, Marklund M, Imamura F, Tintle N, Ardisson Korat AV, de Goede J, Zhou X, Yang WS, de Oliveira Otto MC, Kröger Jet al. Omega-6 fatty acid biomarkers and incident type 2 diabetes: Pooled analysis of individual-level data for 39 740 adults from 20 prospective cohort studies. Lancet Diabetes Endocrinol. 2017;5(12):965–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Jacobs DR Jr., Gross MD, Tapsell LC. Food synergy: An operational concept for understanding nutrition. Am J Clin Nutr. 2009;89(5):1543s–8s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Caumo A, Perseghin G, Brunani A, Luzi L. New insights on the simultaneous assessment of insulin sensitivity and beta-cell function with the HOMA2 method. Diabetes Care. 2006;29(12):2733–4. [DOI] [PubMed] [Google Scholar]
- 29. O'Brien DM, Kristal AR, Jeannet MA, Wilkinson MJ, Bersamin A, Luick B. Red blood cell delta15N: A novel biomarker of dietary eicosapentaenoic acid and docosahexaenoic acid intake. Am J Clin Nutr. 2009;89(3):913–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Harris PA, Taylor R, Minor BL, Elliott V, Fernandez M, O'Neal L, McLeod L, Delacqua G, Delacqua F, Kirby Jet al. The REDCap consortium: Building an international community of software platform partners. J Biomed Inform. 2019;95:103208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Harris PA, Taylor R, Thielke R, Payne J, Gonzalez N, Conde JG. Research electronic data capture (REDCap)–A metadata-driven methodology and workflow process for providing translational research informatics support. J Biomed Inform. 2009;42(2):377–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Bray M, Pomeroy J, Knowler WC, Hopkins S, Boyer BB. Objective measurement of physical activity in Yup'ik Eskimos using a combined heart-rate movement monitor [abstract]. In: International Journal of Circumpolar Health in 2013: Proceedings of the 15th International Congress on Circumpolar Health; 2012; 72(1):22447. Fairbanks, AK. DOI: 10.3402/ijch.v72i0.22447 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Lohman TG, Roche AF, Martorell R. Anthropometric Standardization Reference Manual. Champaign, IL: Human Kinetics Books; 1988. [Google Scholar]
- 34. Boyer BB, Mohatt GV, Plaetke R, Herron J, Stanhope KL, Stephensen C, Havel PJ. Metabolic syndrome in Yup'ik Eskimos: The Center for Alaska Native Health Research (CANHR) study. Obesity. 2007;15(11):2535–40. [DOI] [PubMed] [Google Scholar]
- 35. Votruba SB, Shaw PA, Oh EJ, Venti CA, Bonfiglio S, Krakoff J, O'Brien DM. Associations of plasma, RBCs, and hair carbon and nitrogen isotope ratios with fish, meat, and sugar-sweetened beverage intake in a 12-wk inpatient feeding study. Am J Clin Nutr. 2019;110(6):1306–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Zeigler M, Whittington D, Sotoodehnia N, Lemaitre RN, Totah RA. A sensitive and improved throughput UPLC-MS/MS quantitation method of total cytochrome P450 mediated arachidonic acid metabolites that can separate regio-isomers and cis/trans-EETs from human plasma. Chem Phys Lipids. 2018;216:162–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Bollinger JG, Thompson W, Lai Y, Oslund RC, Hallstrand TS, Sadilek M, Turecek F, Gelb MH. Improved sensitivity mass spectrometric detection of eicosanoids by charge reversal derivatization. Anal Chem. 2010;82(16):6790–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Aslibekyan S, Vaughan LK, Wiener HW, Lemas DJ, Klimentidis YC, Havel PJ, Stanhope KL, O'Brien D M, Hopkins SE, Boyer BBet al. Evidence for novel genetic loci associated with metabolic traits in Yup'ik people. Am J Hum Biol. 2013;25(5):673–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Li J, Ji L. Adjusting multiple testing in multilocus analyses using the eigenvalues of a correlation matrix. Heredity. 2005;95(3):221–7. [DOI] [PubMed] [Google Scholar]
- 40. Nyholt DR. A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am J Hum Genet. 2004;74(4):765–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Matsukawa A, Hogaboam CM, Lukacs NW, Lincoln PM, Evanoff HL, Kunkel SL. Pivotal role of the CC chemokine, macrophage-derived chemokine, in the innate immune response. J Immunol. 2000;164(10):5362–8. [DOI] [PubMed] [Google Scholar]
- 42. Guzik TJ, Skiba DS, Touyz RM, Harrison DG. The role of infiltrating immune cells in dysfunctional adipose tissue. Cardiovasc Res. 2017;113(9):1009–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Osborn O, Olefsky JM. The cellular and signaling networks linking the immune system and metabolism in disease. Nat Med. 2012;18(3):363–74. [DOI] [PubMed] [Google Scholar]
- 44. Vasudevan AR, Wu H, Xydakis AM, Jones PH, Smith EO, Sweeney JF, Corry DB, Ballantyne CM. Eotaxin and obesity. J Clin Endocrinol Metab. 2006;91(1):256–61. [DOI] [PubMed] [Google Scholar]
- 45. Innes JK, Calder PC. The differential effects of eicosapentaenoic acid and docosahexaenoic acid on cardiometabolic risk factors: A systematic review. Int J Mol Sci. 2018;19(2):532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Calder PC. Fatty acids and inflammation: The cutting edge between food and pharma. Eur J Pharmacol. 2011;668:S50–8. [DOI] [PubMed] [Google Scholar]
- 47. Calder PC. Mechanisms of action of (n-3) fatty acids. J Nutr. 2012;142(3):592s–9s. [DOI] [PubMed] [Google Scholar]
- 48. Calder PC. Marine omega-3 fatty acids and inflammatory processes: Effects, mechanisms and clinical relevance. Biochim Biophys Acta. 2015;1851(4):469–84. [DOI] [PubMed] [Google Scholar]
- 49. Saraiva M, Vieira P, O'Garra A. Biology and therapeutic potential of interleukin-10. J Exp Med. 2020;217(1):e20190418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Saraiva M, O'Garra A. The regulation of IL-10 production by immune cells. Nat Rev Immunol. 2010;10(3):170–81. [DOI] [PubMed] [Google Scholar]
- 51. Xue B, Yang Z, Wang X, Shi H. Omega-3 polyunsaturated fatty acids antagonize macrophage inflammation via activation of AMPK/SIRT1 pathway. PLoS One. 2012;7(10):e45990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Berin MC, Dwinell MB, Eckmann L, Kagnoff MF. Production of MDC/CCL22 by human intestinal epithelial cells. Am J Physiol Gastroint Liver Physiol. 2001;280(6):G1217–26. [DOI] [PubMed] [Google Scholar]
- 53. Liu T, Zhang L, Joo D, Sun SC. NF-κB signaling in inflammation. Signal Transduct Target Ther. 2017;2(1):17023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Aslibekyan S, Wiener HW, Havel PJ, Stanhope KL, O'Brien DM, Hopkins SE, Absher DM, Tiwari HK, Boyer BB. DNA methylation patterns are associated with n-3 fatty acid intake in Yup'ik people. J Nutr. 2014;144(4):425–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Ryman TK, Austin MA, Hopkins S, Philip J, O'Brien D, Thummel K, Boyer BB. Using exploratory factor analysis of FFQ data to identify dietary patterns among Yup'ik people. Public Health Nutr. 2014;17(3):510–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Choy K, Nash SH, Hill C, Bersamin A, Hopkins SE, Boyer BB, O'Brien DM. The nitrogen isotope ratio is a biomarker of Yup'ik traditional food intake and reflects dietary seasonality in segmental hair analyses. J Nutr. 2019;149(11):1960–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Fall JA. The division of subsistence of the Alaska Department of Fish and Game: An overview of its research program and findings: 1980–1990. Arct Anthropol. 1990;27(2):68–92. [Google Scholar]
- 58. Johnson JS, Nobmann ED, Asay E, Lanier AP. Dietary intake of Alaska native people in two regions and implications for health: The Alaska native dietary and subsistence food assessment project. Int J Circumpolar Health. 2009;68(2):109–22. [DOI] [PubMed] [Google Scholar]
- 59. Luick B, Bersamin A, Stern JS. Locally harvested foods support serum 25-hydroxyvitamin D sufficiency in an indigenous population of Western Alaska. Int J Circumpolar Health. 2014;73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Nash SH, Bersamin A, Kristal AR, Hopkins SE, Church RS, Pasker RL, Luick BR, Mohatt GV, Boyer BB, O'Brien DM. Stable nitrogen and carbon isotope ratios indicate traditional and market food intake in an indigenous circumpolar population. J Nutr. 2012;142(1):84–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Schichnes J, Chythlook M. Use of fish and wildlife in Manokotak, Alaska. Dillingham, AK: Alaska Department of Fish and Game, Division of Subsistence; 1988. [Google Scholar]
- 62. Ryman TK, Boyer BB, Hopkins S, Philip J, Beresford SA, Thompson B, Heagerty PJ, Pomeroy JJ, Thummel KE, Austin MA. Associations between diet and cardiometabolic risk among Yup'ik Alaska native people using food frequency questionnaire dietary patterns. Nutr Metab Cardiovasc Dis. 2015;25(12):1140–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Rangel-Huerta OD, Aguilera CM, Mesa MD, Gil A. Omega-3 long-chain polyunsaturated fatty acids supplementation on inflammatory biomarkers: A systematic review of randomised clinical trials. Br J Nutr. 2012;107(Suppl 2):S159–70. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data described in the manuscript, codebook, and analytic code will be made available upon request pending Tribal approval by the Yukon-Kuskokwim Health Corporation Executive Board.



