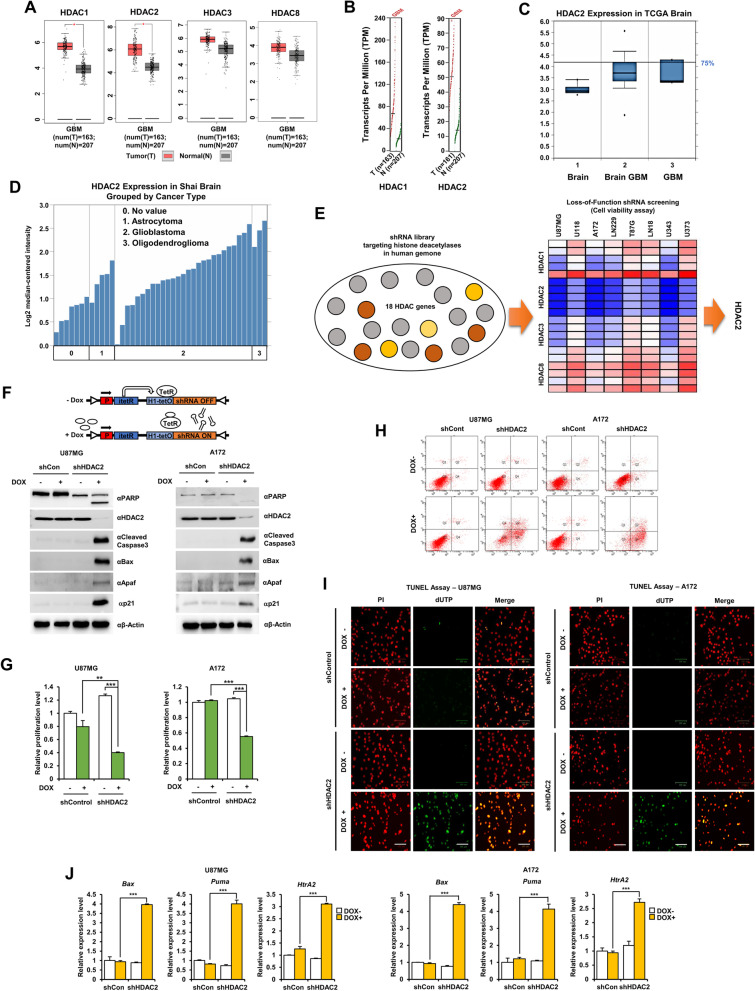
Fig. 1.
HDAC2 is highly expressed in GBM. A Expression of HDAC1, 2, 3 and 8 proteins. 163 GBM tissues (T) and 207 normal tissues (N) were analyzed from the GEPIA Database. *p < 0.05. B HDAC1 and HDAC2 expression in GBM tissues from the GEPIA Database. Axis units are Log2(TPM + 1) (TPM: Transcripts Per Million). C Box plots derived from gene expression data from the Oncomine database comparing expression of HDAC2. D HDAC2 expression levels from the Oncomine database. E Screening of a shRNA library systematically targeting HDAC genes in GBM cells. Cell viability was measured by using the MTT assay. HDAC knockdown was performed using the lentiviral system of shRNA library (five unique shRNA of HDACs) targeting 18 types of HDACs in GBM cells (U87MG, U118, A172, LN229, T98G, LN18, U343, and U373). Red: High-cell viability, Blue: Low-cell viability. F Model of lentiviral expression of DOX-inducible shRNA targeting human HDAC2. G Cell proliferation in DOX-inducible shHDAC2 GBM cells. H FACS analysis in DOX-inducible shHDAC2 GBM cells. I TUNEL assay in DOX-inducible shHDAC2 GBM cells. TUNEL-positive cells were stained (green), and nuclei were counterstained with PI (red). J mRNA expression of apoptotic cell death markers (Bax, Puma and HtrA2) using qPCR in DOX-inducible shHDAC2 GBM cells with doxycycline. Scale bar: 100 μm. All data are expressed as the mean ± SD from three independent experiments, each performed in triplicate. *p < 0.05, **p < 0.01, ***p < 0.001