Fig. 1 |. Updated classification of class 1 CRISPR–Cas systems.
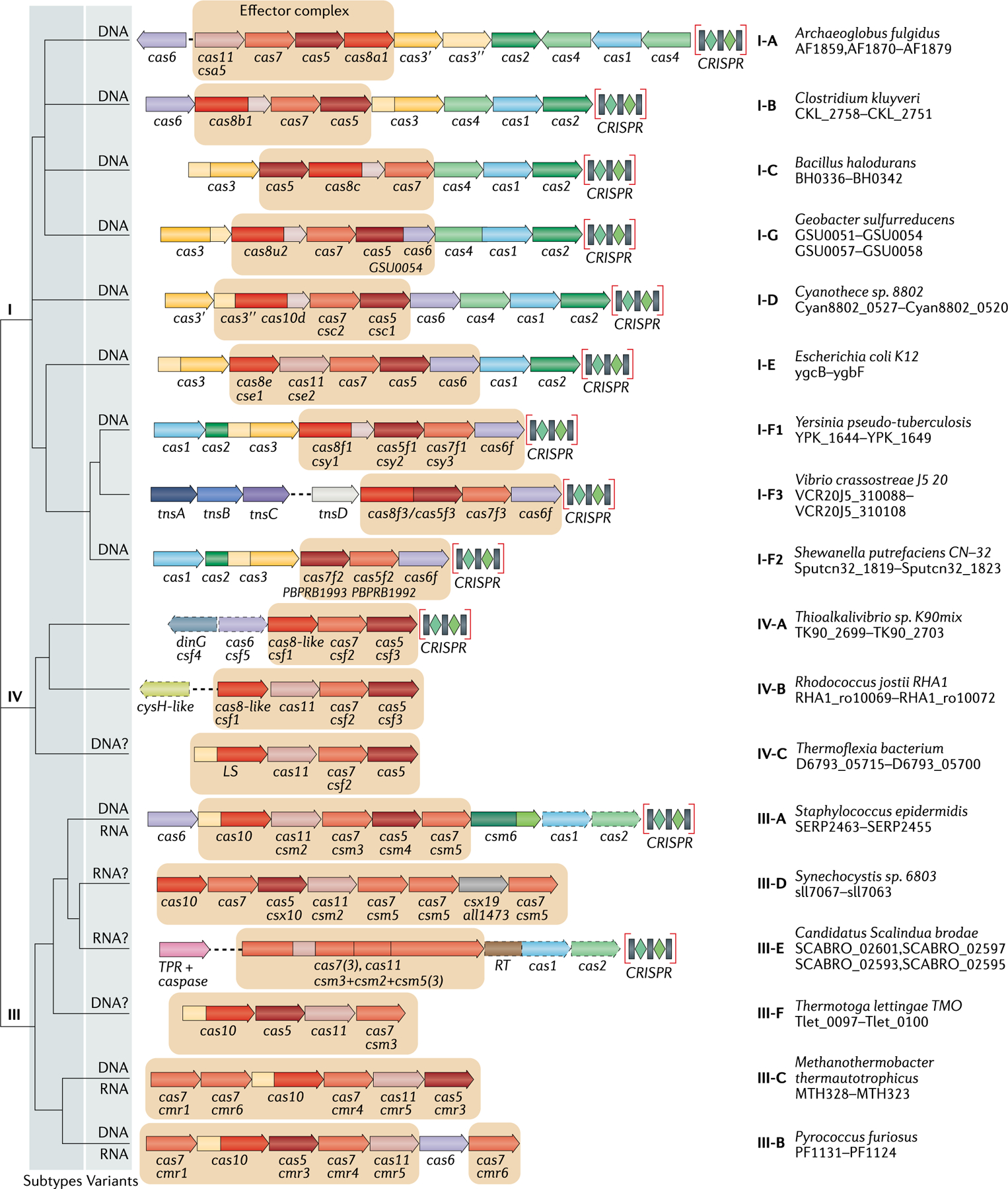
The figure schematically shows representative (typical) CRISPR–cas loci of each class 1 subtype and of selected distinct variants, with the dendrogram on the left showing the likely evolutionary relationships between the types and subtypes. The column on the right indicates the organism and the corresponding gene range. Homologous genes are colour-coded and identified by a family name. The gene names follow the previous classification18. Where both a systematic name and a legacy name are commonly used, the legacy name is given under the systematic name. The small subunit is encoded by csm2, cmr5, cse2, csa5 and several additional families of homologous genes that are collectively denoted cas11. The adaptation module genes cas1 and cas2 are dispensable in subtypes III-A and III-E (dashed lines). Gene regions coloured cream represent the HD nuclease domain; the HD domain in Cas10 is distinct from that in Cas3 and Cas3″. Functionally uncharacterized genes are shown in grey. The tan shading shows the effector module. The grey shading of different hues shows the two levels of classification: subtypes and variants. Most of the subtype III-B, III-C, III-E and III-F loci, as well as IV-B and IV-C loci, lack CRISPR arrays and are shown accordingly, although for each of the type III subtypes exceptions have been detected. CHAT, protease domain of the caspase family; RT, reverse transcriptase; TPR, tetratricopeptide repeat.
