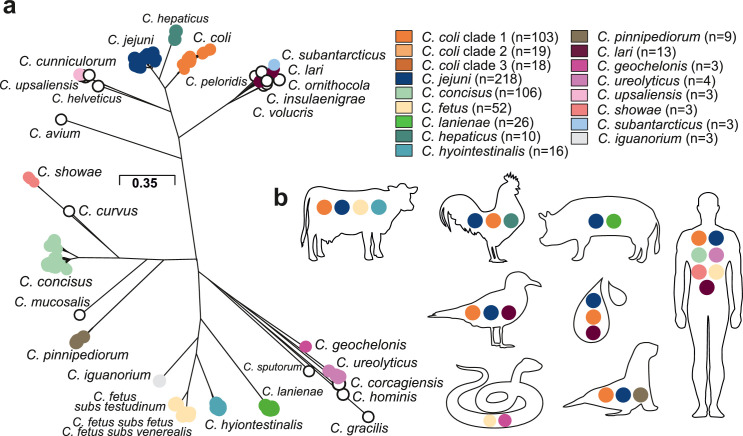
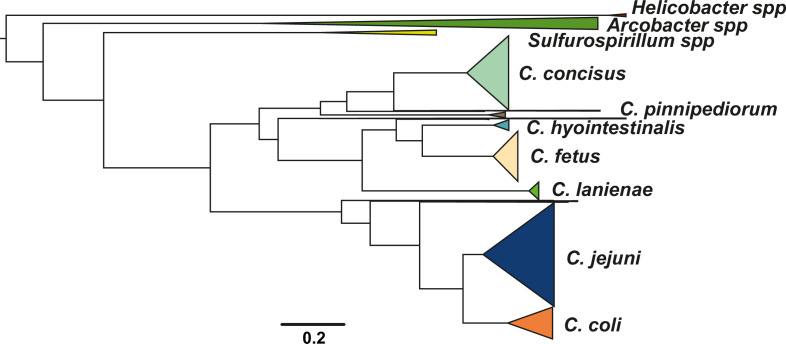
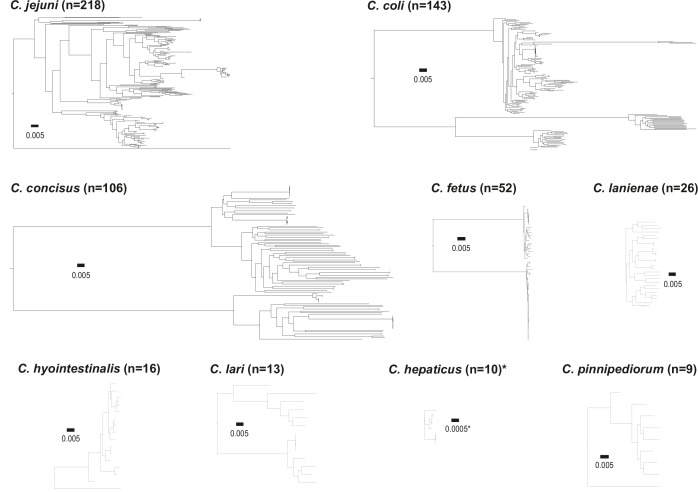
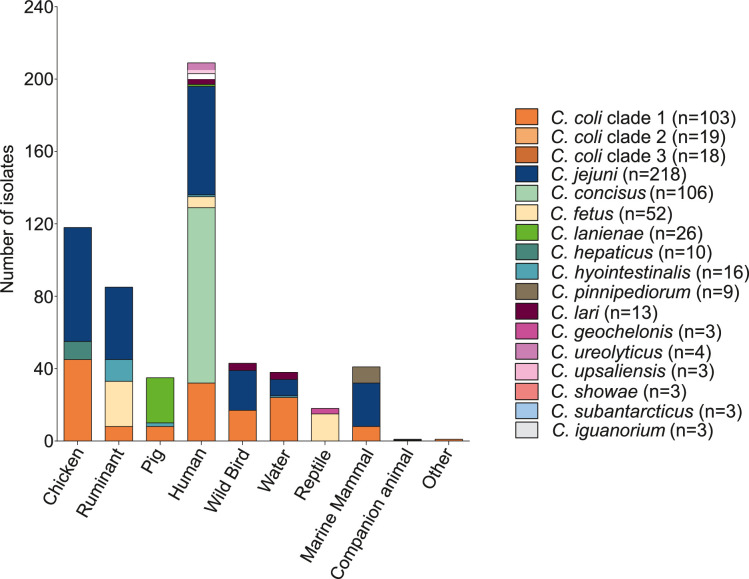
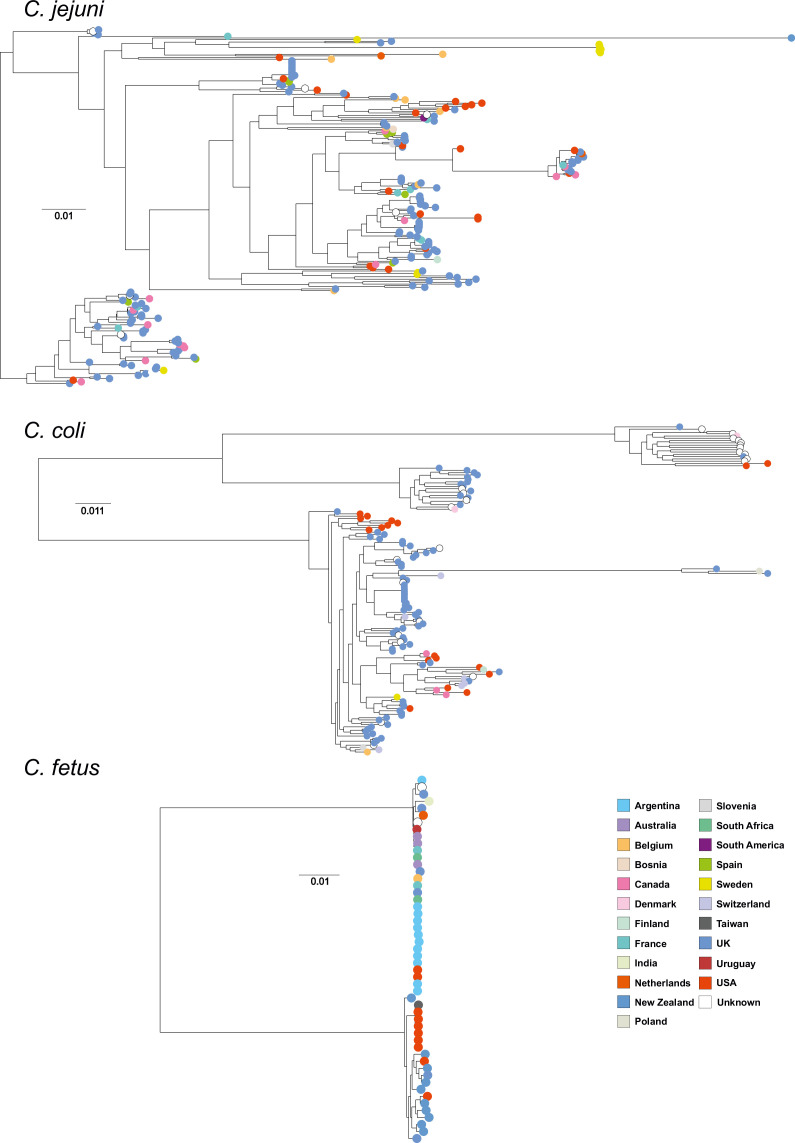
Figure 2. Population structure and host ecology in the genus Campylobacter.
(a) Phylogenetic tree of 631 Campylobacter isolates from 30 species reconstructed using a gene-by-gene concatenated alignment of 820 core genes (shared by >95% of isolates) and an approximation of the maximum-likelihood (ML) algorithm implemented in RAxML. The species name is indicated adjacent to the associated sequence cluster. The scale bar indicates the estimated number of substitutions per site. (b) Isolation source of Campylobacter species with n ≥ 3 isolates.