Figure 2. Synthetic DNA oligos spiked into amp-seq reactions designed to flag contamination and sample swaps.
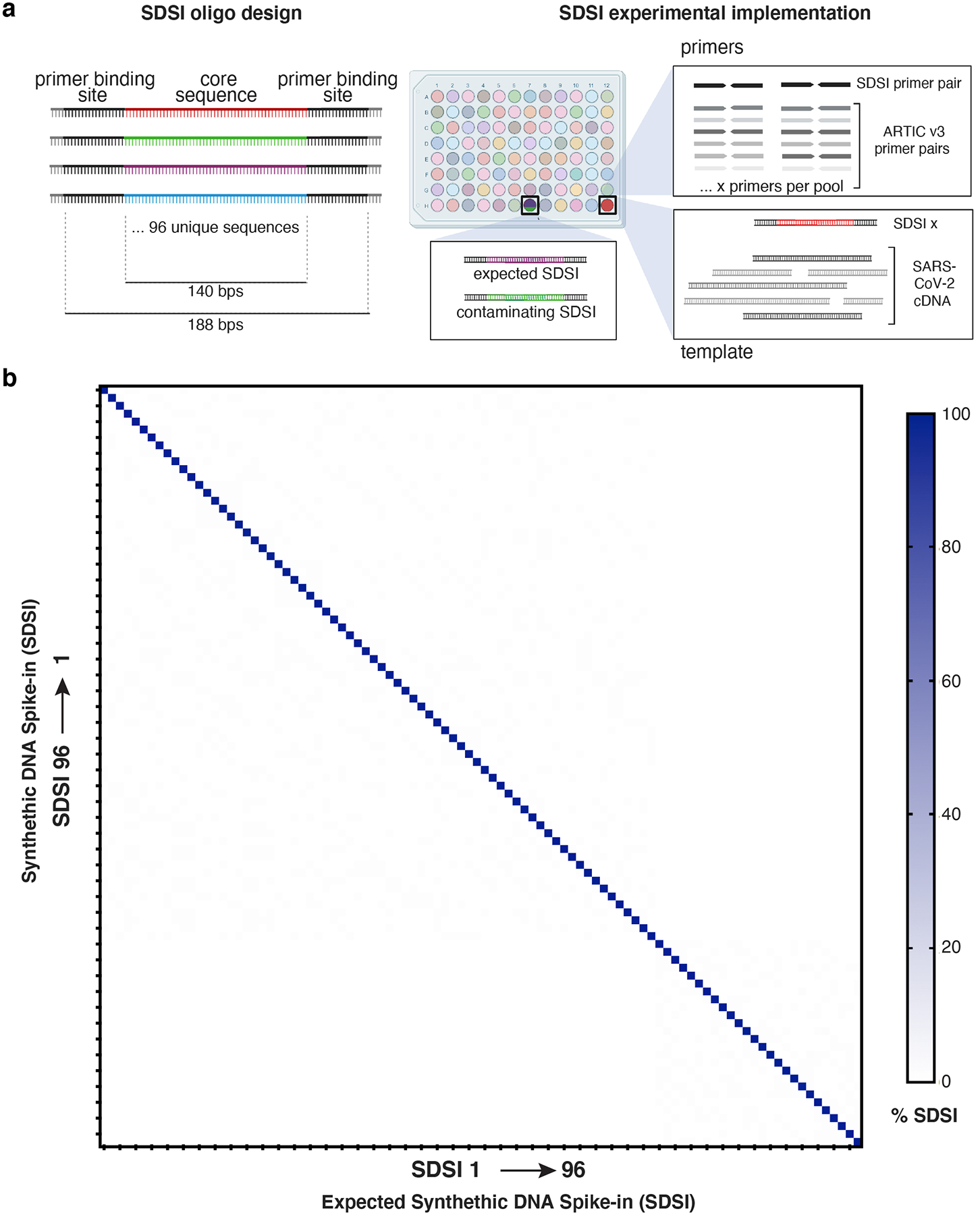
a, Schematic of SDSI design. Each oligo contains 140 bp of unique sequence flanked by common primer binding sites. Primers designed to amplify all SDSIs are added to ARTIC primer pools, and a unique SDSI is added to each clinical sample. Identification of multiple SDSIs in the same sample indicates contamination. b, Percent of SDSI reads mapping for each of the 96 SDSIs (horizontal axis) were quantified for each of the 96 SDSIs (vertical axis). Any off-diagonal signal would indicate non-specific identification of SDSIs.
