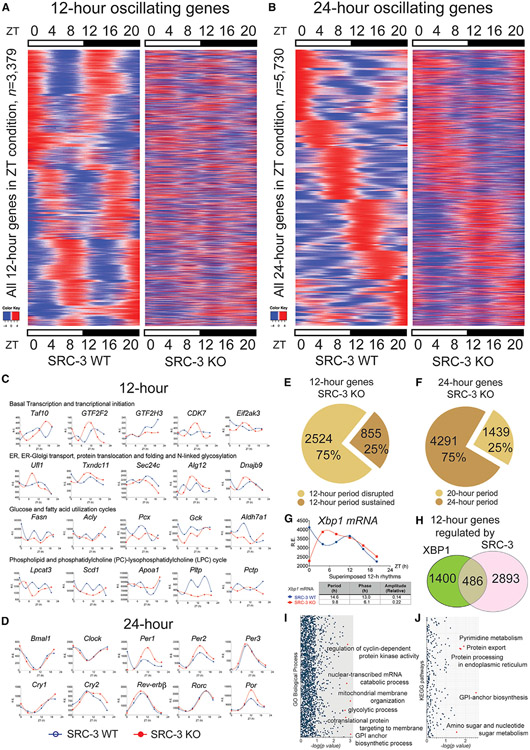
Figure 2. SRC-3 regulates the hepatic 12-h transcriptome. Rhythmic transcripts detected by RNA-seq and the RAIN method were plotted as heatmaps for 12- and 24-h cycling genes.
(A) Heatmaps of the 12-h-oscillating transcripts for the SRC-3 WT and KO mice (n = 3,379).
(B) Heatmaps of the 24-h-oscillating transcripts (n = 5,730).
(C) Representative 12-h gene oscillations plotted against zeitgeber time (ZT) from 4-h-resolution transcriptome profiling of the SRC-3 WT (blue lines) and KO (red lines) mice.
(D) Representative 24-h gene oscillations of core components of the circadian clock plotted against ZT time from 4-h resolution transcriptome profiling of the SRC-3 WT and KO mice. Relative expression (R.E.) is shown as DESeq2 normalized transcript counts.
(E) The proportion of the 12-h gene oscillations that had abolished (n = 2,524, 75%) or sustained (n = 855, 25%) 12-h periods in the SRC-3 KO mice.
(F) The proportion of the 24-h gene oscillations that had a sustained 20-h period (n = 1,439, 25%) or sustained 24-h periods (n = 4,291, 75%) in the SRC-3 KO mice. All the 24-h gene oscillations remain between 20 and 24-h periods in the SRC-3 KO mice.
(G) Xbp1 gene oscillations plotted against ZT from 4-h-resolution transcriptome profiling of the SRC-3 WT (blue lines) and KO (red lines) mice.
(H) Venn diagram of the 12-h transcriptome (n = 1,886) regulated by XBP1 and the 12-h transcriptome (n = 3,379) regulated by SRC-3. Intersection: concordant 12-h-oscillating genes regulated by both XBP1 and SRC-3 (n = 486).
(I and J) Gene enrichment analysis of the 486 concordant 12-h-oscillating genes regulated by both XBP1 and SRC-3 in GO biological processes (I) and KEGG pathways (J). Normalized p values are shown. Red dots indicate the top significant biological processes and KEGG pathways that are regulated by both XBP1 and SRC-3.