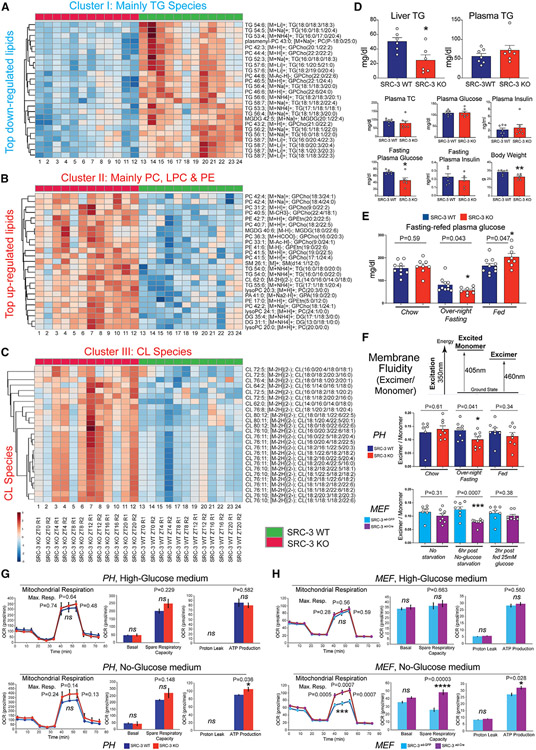
Figure 4. Coactivation of 12-h clock coordinates hepatic lipid metabolic processes.
(A) Heatmap of the cluster I 12-h TG lipid species. The indicated TG lipid species were identified as the top downregulated cluster in the SRC-3 KO mouse livers.
(B) Heatmap of the cluster II 12-h TG lipid species. The indicated PC, LPC, and PE lipid species were identified as the significantly upregulated cluster in the SRC-3 KO mouse livers.
(C) Heatmap of the cluster III 12-h CL lipid species. The indicated CL lipid species were identified as the significant upregulated cluster in the SRC-3 KO mouse livers.
(D) Liver triglycerides (TG), plasma TG, plasma total cholesterol (TC), plasma insulin, plasma glucose, fasting plasma glucose and insulin, and body weight in the indicated mouse strains (n = 6).
(E) Plasma glucose at chow, overnight fasting (16 h) and refed (3 h) in the indicated mouse strains (n = 8).
(F) Membrane fluidity of primary hepatocyte (PHs, n = 6–8 technical replicates from 1 pool of PHs combined from three mouse livers) and MEFs) (n = 6-8 replicates from the MEFs that were independently transfected with Ad-GFP or Ad-Cre for SRC-3 knockout) measured using a fluorescent lipophilic probe, pyrenedecanoic acid (PDA). The absolute excimer: monomer ratio of emission at 460 nm to emission at 405 nm is shown in the indicated SRC-3WT and KO PHs and SRC-3ad–GFP and SRC-3Ade–Cre MEFs.
(G and H) Seahorse metabolic profiling of the indicated SRC-3WT vs. KO PHs (G; n = 4 technical replicates from 1 pool of PHs combined from three mouse livers) and SRC-3ad–GFP vs. SRC-3Ade–Cre MEFs (H; n = 11–12 replicates from the MEFs that were independently transfected with Ad-GFP or Ad-Cre for SRC-3 knockout) under high-glucose medium (top) or no-glucose medium conditions (bottom). Cell Mito Stress test profiles for fundamental mitochondrial functions are shown. OCR, oxygen consumption rate. Unpaired Student’s t test was performed with p value indicated. Data are graphed as mean ± SEM. *p < 0.05, **p < 0.005, ***p < 0.001, ****p < 0.0001.