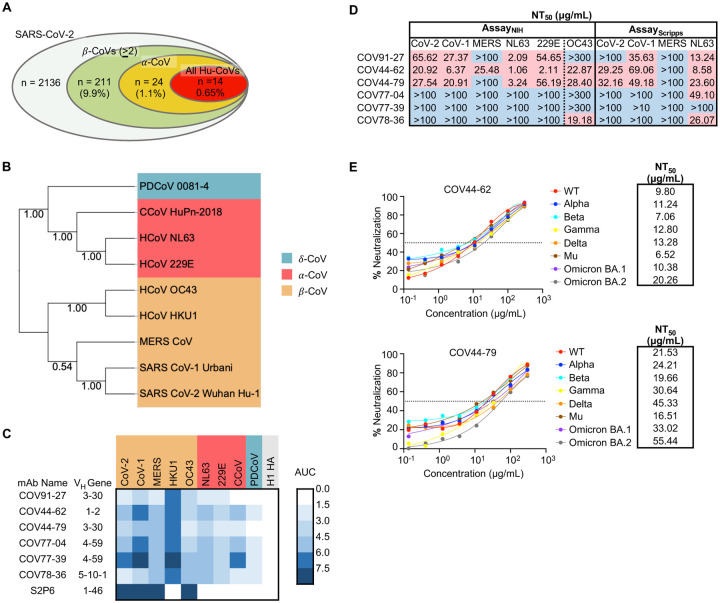
Figure 1. Broadly neutralizing antibodies target coronaviruses associated with human disease.
(A) Analysis of the frequency of MBCs expressing broadly reactive antibodies from n = 19 donors. Values in parentheses represent the percentage of SARS-CoV-2 reactive supernatants that also bind the specified subsets of non-SARS coronavirus spikes. A total of 10,356 MBC culture supernatants (50–100 B cells/well) was screened.
(B) Phylogenetic relationships across the coronavirus spike proteins targeted by the broadly reactive mAbs were inferred by the Neighbor-Joining method in MEGA11 using full-length amino-acid sequences of CoV spike proteins. Bootstrap values from 500 samplings are shown on the branches.
(C) Heat map representing the binding of broadly reactive mAbs to spike proteins from coronaviruses across the alpha, beta and deltacoronavirus genera. H1 hemagglutinin was included as a negative control for mAb binding experiments and area under the curve (AUC) values for each antigen are shown after subtraction with values for the negative control antigen CD4.
(D) Values represent antibody titer at 50% neutralization (NT50) against SARS-CoV-2 Wuhan Hu-1, SARS-CoV-1, MERS-CoV, HCoV-NL63 and HCoV-229E envelope-pseudotyped lentivirus, as well as authentic HCoV-OC43. NT50 values were calculated using the dose-response-inhibition model with 5-parameter Hill slope equation in GraphPad Prism 9.
(E) Neutralization of SARS-CoV-2 variants of concern (pseudovirus) by COV44-62 and COV44-79.