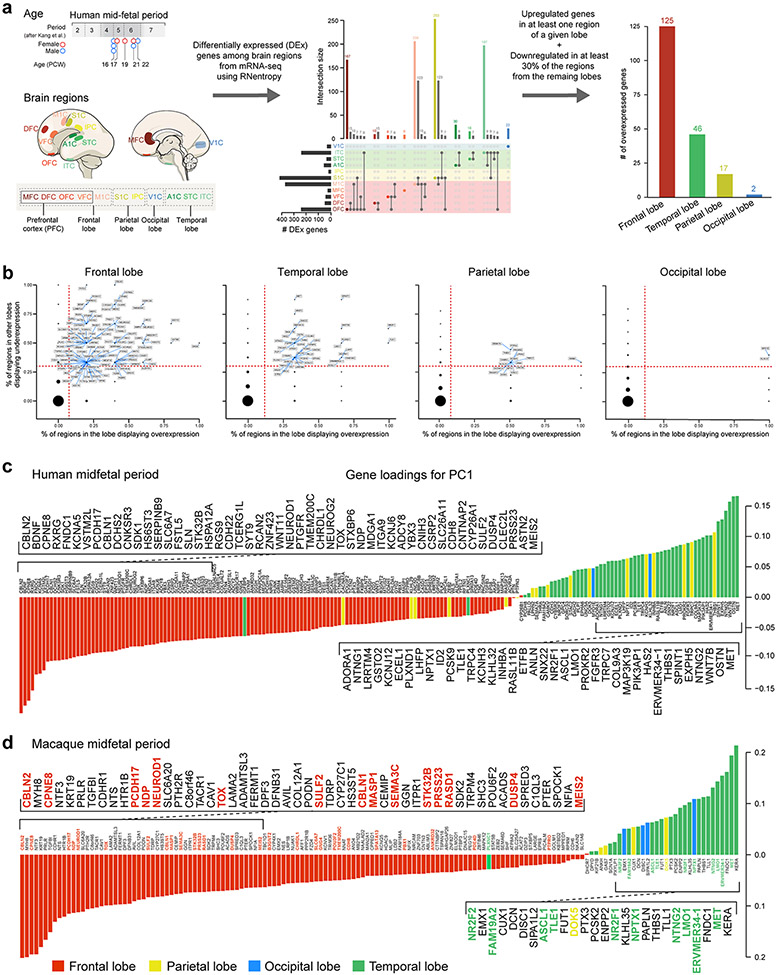
Extended Data Fig. 1 ∣. Workflow for analysis of midfetal RNA-seq data and genes upregulated in individual lobes.
a, Human developmental neocortical RNA-seq dataset and workflow for analysis to identify genes upregulated in each cortical lobe. b, Genes upregulated in the frontal, temporal, parietal and occipital lobes in comparison to the other lobes. We identified 190 protein-coding genes that are specifically upregulated, using stringent criteria, in at least one area within a lobe in comparison with areas from other lobes: 125 in the frontal lobe, 46 in the temporal lobe, 17 in the parietal lobe, and 2 in the occipital lobe. The X-axis represents proportion of putative areas in the frontal lobe in which the gene is significantly upregulated according to RNentropy. The Y-axis represents proportion of areas in the other lobes in which the gene is significantly downregulated according to RNentropy. Upregulated genes, the ones delimited by dashed red lines, are labeled. c,d, Gene loadings of PC1 from PCA of protein-coding genes that are specifically enriched in one of the four lobes of the midfetal human (c) and macaque (d) cortex. Colors represent the cortical lobe where the gene was found to be specifically upregulated. For reproducibility information, see Methods.