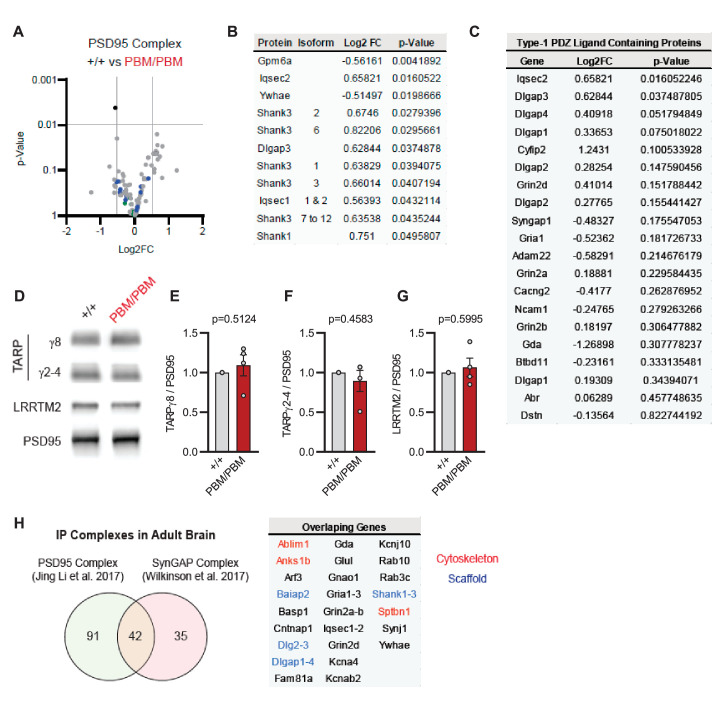
Figure 7. Characterization of native PSD95 complexes from Syngap1PBM animals.
(A) Volcano plot demonstrating the label-free quantitative mass-spectrometry profile of the logarithmic difference in protein levels in the immunoprecipitated PSD95 complexes derived from DIV21 +/+ and PBM/PBM cultures in inactivated state. Only Gpm6a (shown in black) was significantly altered beyond p > 0.001 cutoff. Blue dots represent proteins with type 1 PDZ-ligands. Green dots represent DLG family proteins. p Values were calculated via t-test for each protein. Samples were derived from individual cultures (4 per genotype) which are immunoprecipitated separately. Log2FC was calculated as ratio of PBM/PBM over +/+. (B) List of proteins that are differentially expressed beyond p > 0.05 cutoff. Note that Iqseq2 and Dlgap3 are PDZ-binding proteins. (C) Mass-spectrometry profile of type-1 PDZ binding motif containing proteins in immunoprecipitated PSD95 complex in +/+ vs PBM/PBM inactivated cultures. (D) Western blots showing relative expression of TARPs and Lrrtm2 in PSD fractions from adult hippocampi in +/+ vs PBM/PBM. (E–G) Quantifications of (D). (E) TARPg8 t(6)=0.6961, P = 0.5124. (F) TARPg2-4 t(6)=0.7924, p = 0.4583 (G) Lrrtm2 t(6)=0.5542, p = 0.5995. Each sample represents hippocampi pooled from 2 mice. (H) Comparison of PSD95 and SynGAP IP complexes as reported by Li et al., 2017 and Wilkinson et al., 2017. Note that PSD95 and SynGAP complexes share diverse range of components involving cytoskeletal and scaffolding proteins.