Fig. 2.
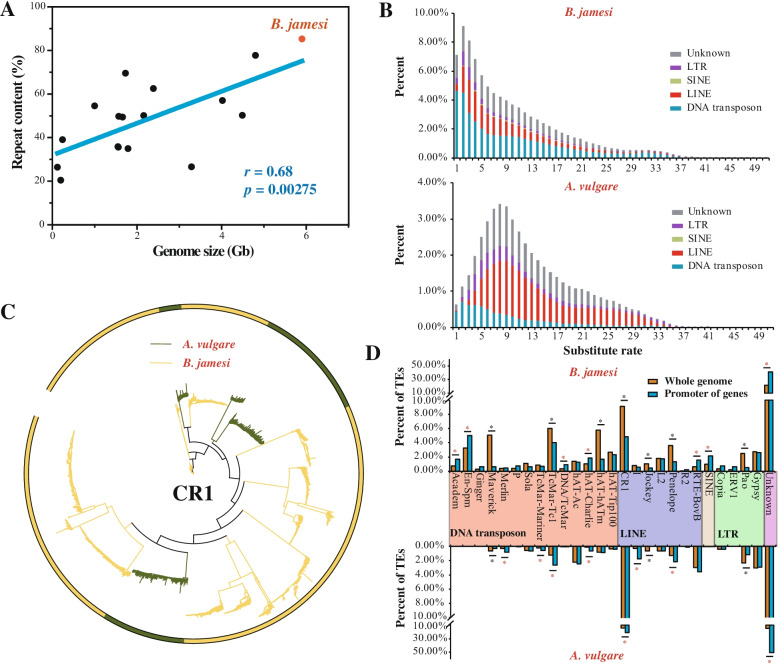
The evolution of transposable elements (TEs) and genome size. A The relationship between the genome size and repeat content. The repeat contents and genome sizes of the sequenced crustacean genomes were summarized in the Additional file 1: Table S2. The TE content and the genome size was positively correlated with the Pearson correlation r = 0.68 and p-value = 0.00275. B Kimura distance-based copy divergence analyses of TEs in the two isopod genomes, B. jamesi and A. vulgare. The graphs represent genome coverage for each TE superfamily in the different genomes analyzed. Clustering was performed according to their Kimura distances (K-value from 0 to 50). C Phylogenetic tree of the CR1 LINEs from B. jamesi (yellow) and A. vulgare (dark gray). D Enrichment analyses of TE families within gene promoters. The closest TE was calculated for each gene, and the content of the closest TEs were calculated and compared with that of the whole genome