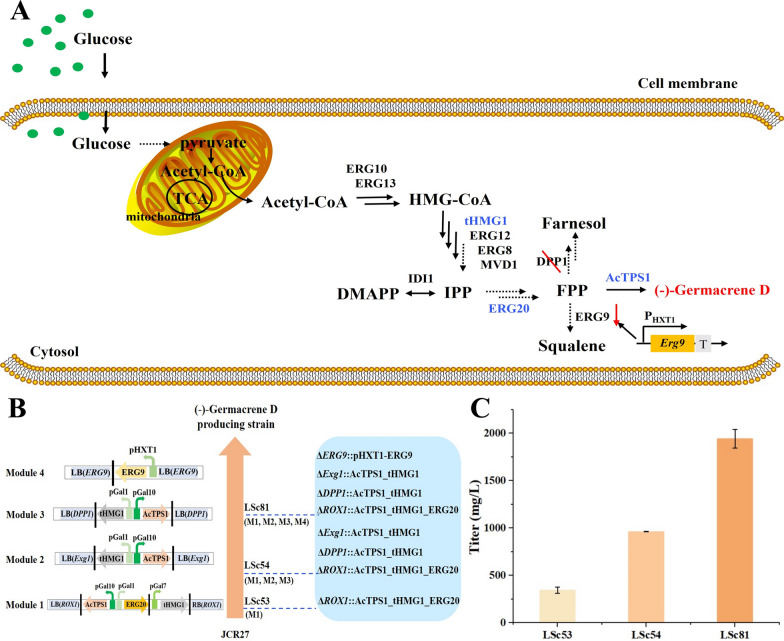
Fig. 6.
Titer of (–)-germacrene D in engineered yeast strains in shake-flask fermentation. A Biosynthesis pathways for (–)-germacrene D in engineered yeast. ERG10 acetyl-CoA C-acetyltransferase, ERG13 3-hydroxy-3-methylglutaryl coenzyme A synthase, tHMG1 truncated 3-hydroxy-3-methylglutaryl-coenzyme A reductase 1, ERG8 phosphomevalonate kinase, ERG12 mevalonate kinase, MVD1 mevalonate diphosphate decarboxylase 1, IDI1 isopentenyl-diphosphate delta isomerase 1, ERG20 farnesyl pyrophosphate synthetase, AcTPS1 (–)-germacrene D synthase. Overexpressed enzymes are shown in blue. Red slash, gene knockout; Red downward arrow, downregulated gene expression. B The engineered yeast strains integrated with gene modules. These expression modules were integrated into the chromosomal sites of ROX1 (module 1, chromosome XIV), Exg1 (module 2, chromosome XII), DPP1 (module 3, chromosome IV), and ERG9 (module 4, chromosome VIII). The engineered strain LSc53 contained expression module 1 (abbreviated as ‘M1’). The engineered strain LSc54 contained the expression module 1 (‘M1’), module 2 (‘M2’) and module 3 (‘M3’). The engineered strain LSc81 contained the expression module 1 (‘M1’), module 2 (‘M2’), module 3 (‘M3’) and module 4 (‘M4’). (C) The production of different engineered stains. Germacrene D levels were determined after 72 h fermentation in YPDH medium as described in “Methods”. Error bars represent standard deviations from three independent experiments