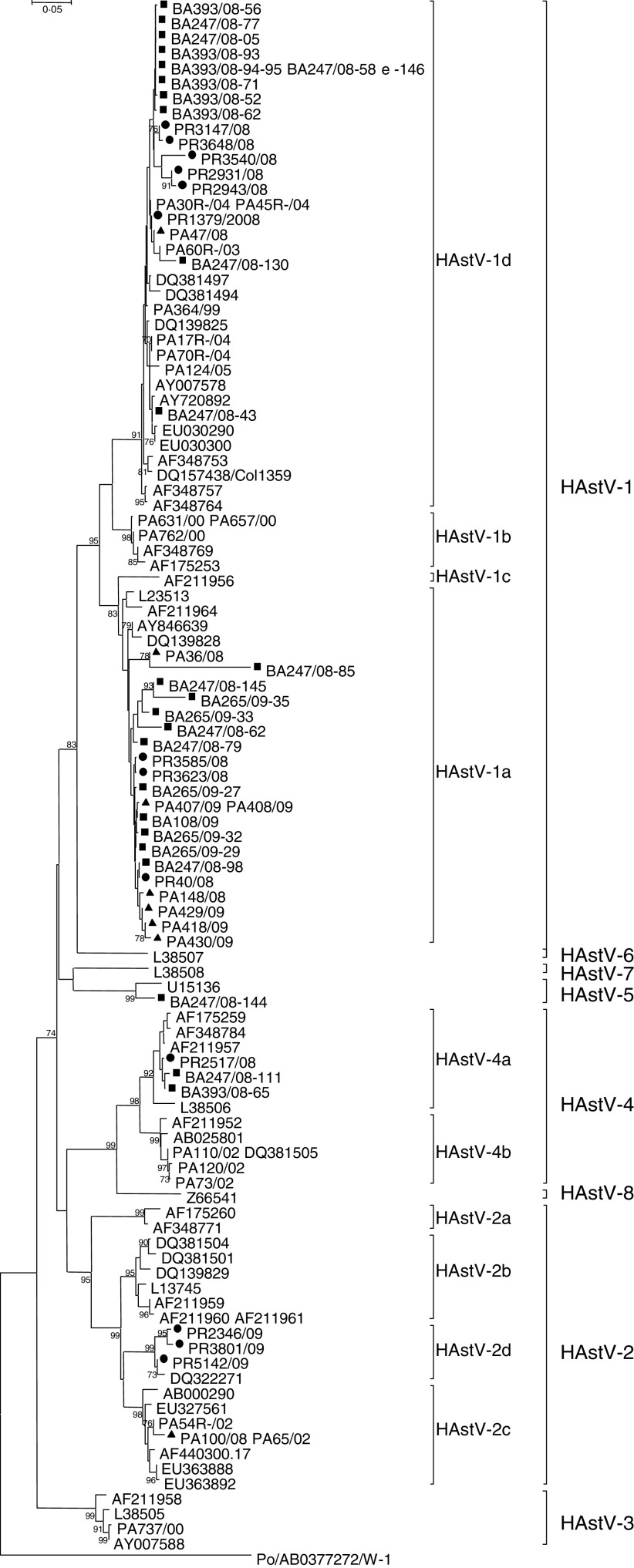
Fig. 1.
Phylogenetical analysis of partial nucleotide sequences (348 bp) of HAstV ORF2 capsid region of 49 strains collected in Bari, Palermo, and Parma from 2008 to 2009. The Kimura two-parameter model of substitution and the neighbour-joining method were used to construct the phylogenetic tree. Bootstrap values above 70%, estimated with 1000 pseudoreplicate datasets, are indicated at each node. Italian strains of this study are indicated as follows: ▪, HAstV from Bari; ▴, HAstV from Palermo; •, HAstV from Parma.