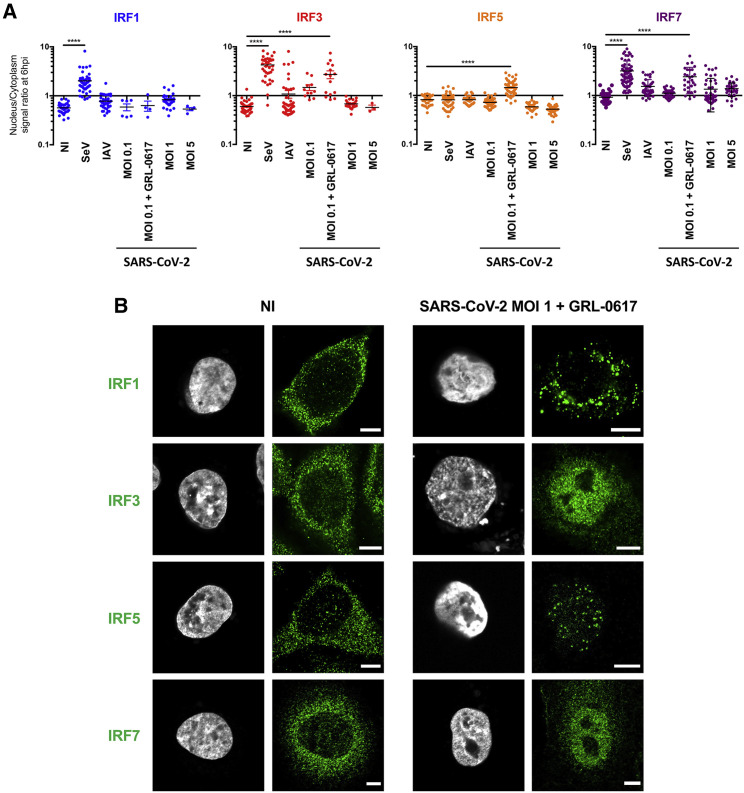
Figure 3.
SARS-CoV-2 and other RNA viruses activate IRFs differentially
(A) Localization of endogenous transcription factors in resting and stimulated A549-ACE2 cells. Cells were infected with SeV, influenza A virus (H1N1 WSN) (IAV) at MOI 1, or SARS-CoV-2 at the indicated MOIs, for 6 h or left uninfected (NI). Alternatively, cells were infected in the presence of the PLP inhibitor GRL-0617 (50 μM). Cells fixed and labeled with anti-IRF1, IRF3, IRF5, or IRF7 were acquired with Airyscan mode. Fluorescence intensities in the nuclear and cytoplasmic regions of interest (ROIs) were measured using FiJi. Each point represents the nucleus/cytoplasm ratios of mean gray values for a single cell. The graph shows the median and interquartile range for ∼30 cells per condition from 3 independent experiments. ∗∗∗∗p < 0.0001, as determined by one-way ANOVA with Bonferroni post hoc test.
(B) Representative images from (A). Scale bar: 5 μm.