FIGURE 1.
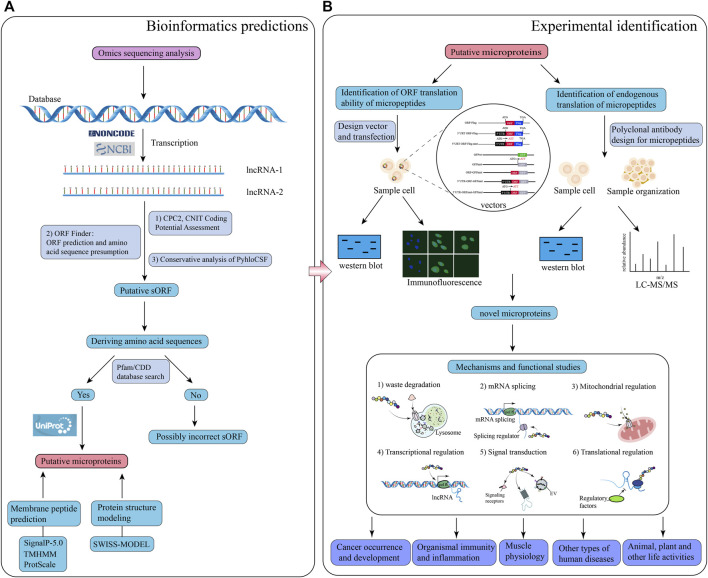
Schematic illustration of the workflow for bioinformatics prediction and experimental analysis of lncRNA-encoded micropeptides. (A) Bioinformatics prediction: firstly, construct a database of putative lncRNA-encoding micropeptides by applying the results of omics sequencing, and search the putative lncRNA sequences with coding potential through NCBI or NONCODE database; secondly, use calculation tools, and databases such as CPC2, CNIT, ORF Finder, PyhloCSF, etc. to evaluate the coding potential of the putative lncRNA, and deduce the corresponding sORF, and amino acid sequence; thirdly, the deduced amino acid sequences were put into the Pfam and CDD databases to look for them, and if they matched, the search for the putative micropeptide information was continued through the UniProt database; finally, the characteristics and structure of the putative micropeptide were predicted and modeled through calculation tools and databases such as SignalP-5.0, TMHMM, ProtScale and SWISS-MODEL; (B) Laboratory identification: design a series of special vectors to be transfected into specific cells, and apply western blot and immunofluorescence experiments to identify micropeptides; meanwhile, polyclonal antibodies to this micropeptide were designed, and detected by western blot and LC-MS/MS experiments on sample cells and tissues. Based on the results of both experimental procedures, the putative micropeptide was identified as a novel micropeptide, and then the function and mechanism of the micropeptide were investigated.