FIG 2.
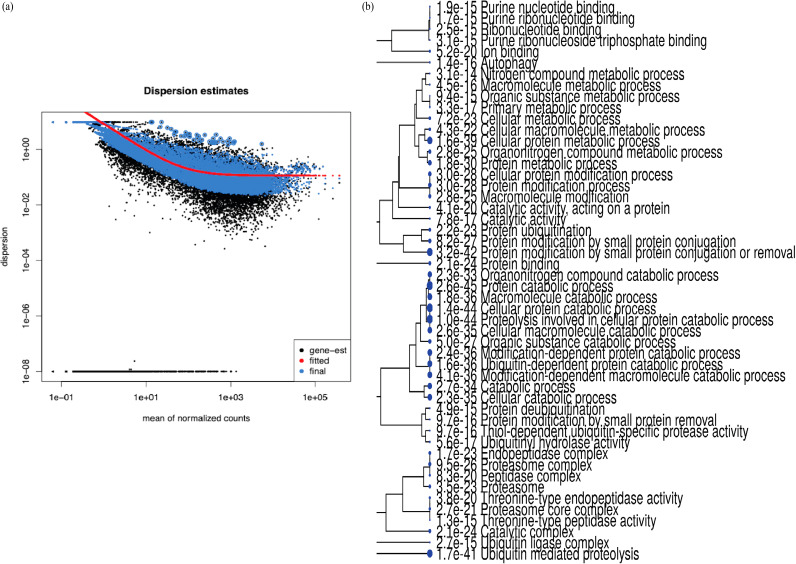
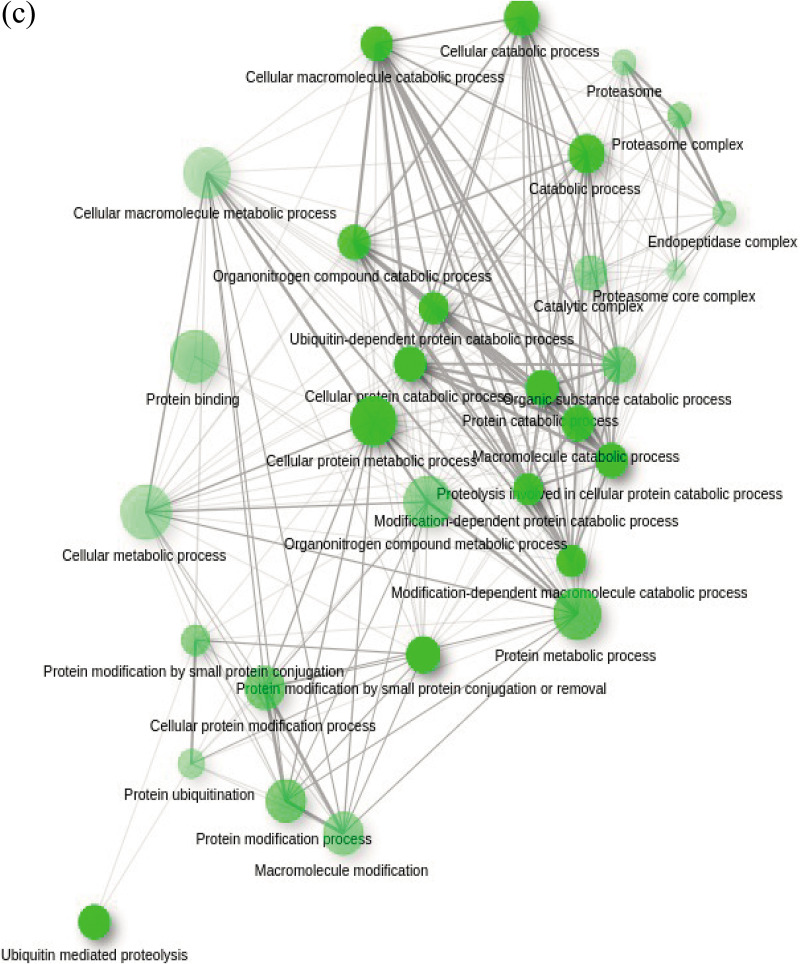
Identification of putative A. aegypti host factors upon CHIKV infection. (a) Representation of differentially expressed genes between two libraries. Black dots represent overall differentially expressed genes while blue dots represent significantly expressed genes (Fold change ± 1.5 and P value < 0.05). (b) Pathway analysis of all 8,608 transcripts found common between lab-generated transcriptomics libraries and DRSC database. Transcripts from pathways such as ubiquitin-related pathways, proteolysis pathways, protein catabolic processes, protein modification, and cellular protein metabolic processes were significantly affected upon CHIKV infection of Aag2 cell line. The sizes of the blue dots represent significance, with bigger dots having a lower P value. (c) Functional analysis of identified orthologous 346 genes found from significant selected pathways. Darker nodes are more significantly enriched gene sets. Bigger nodes represent larger gene sets. Thicker edges represent more overlapped genes.