Abstract
We present a de novo mitogenome assembly obtained from specimens sampled in the so-called summer population (SP) of Thaumetopoea pityocampa (Denis and Schiffermüller, 1775) in Portugal. Contrary to the typical larval development occurring in winter in this species, the larvae of this unique population develop during summer. The sequencing data used were obtained from genomic libraries originally generated to assemble the nuclear genome of T. pityocampa[1]. We also provide a complete annotation and a phylogenetic representation which positions the Portuguese summer population of T. pityocampa and an Italian typical individual of the same species among the Notodontidae family and more distant Noctuoidea species. This data represents a valuable new resource for an expanding and urticating insect pest.
Keywords: Annotation, Assembly, Lepidoptera, Mitogenome, Notodontidae, Phylogeny, Thaumetopoeinae
Specifications Table
| Subject | Omics: Genomics |
| Specific subject area | Lepidoptera, Notodontidae, Mitogenomics |
| Type of data |
|
| How the data were acquired | Two whole-genome Illumina shotgun libraries were sequenced on an Illumina HiSeq2000 platform using the paired-end protocol (2 × 100 base pairs (bp)). |
| Data format | Analysed: de novo assembled and annotated mitogenome |
| Description of data collection | MITObim v1.9 assembled genomic libraries generated from male individuals. Genomic regions were annotated with MITOS v2 and represented as circular mitogenome map with OGDRAW. Multiple sequence alignments were generated with MAFFT v7.471 and subsequently concatenated to build phylogenetic trees with PHYML v3.0 and MrBayes v3.2.7a. Trees were visualized with FigTree v1.4.4. |
| Data source location | Institution: INRAE City/Town/Region: Mata Nacional de Leiria Country: Portugal Latitude and longitude (and GPS coordinates, if possible) for collected samples/data: 39°47’N, 8°58’W |
| Data accessibility | Repository name: NCBI BioProject Data identification number: PRJNA344465 Direct URL to data: http://www.ncbi.nlm.nih.gov/bioproject/344465 Repository name: NCBI BioSample Data identification number: SAMN24475646 Direct URL to data: https://www.ncbi.nlm.nih.gov/biosample/24475646 Repository name: NCBI Genbank Data identification number: OM478592 Direct URL to data: https://www.ncbi.nlm.nih.gov/nuccore/OM478592 |
Value of the Data
-
•
This data represents the mitogenome of a phenologically shifted specimen of Thaumetopoea pityocampa collected in Portugal.
-
•
The present data can be used for phylogenetic studies of Lepidoptera especially for getting deeper insights in diversification dynamics of Notodontidae species.
-
•
The resource will be helpful to study the evolution of the Thaumetopoea genus and the T. pityocampa / T. wilkinsoni species complex.
1. Data Description
The T. pityocampa Summer Population (“TpitSP”) mitogenome presented here had an A + T content of 77.8% and a size of 15,729 bp. All 38 genetic regions reported for arthropod mitogenomes [2], were identified (Supplementary GFF annotation file): 13 protein-coding genes, 22 tRNA and 2 rRNA (12S rRNA and 16S rRNA) genes, and one AT-rich control region. Genes were ordered identically to the Italian T. pityocampa [3] and other Notodontidae [4] mitogenomes. Start and stop codons of the protein-coding genes corresponded also to those observed in other Noctuoidea. In total, 22 intergenic spacer regions (see Table 1: positive intergenic nucleotide values) with an average size of 20 bp and a total length of 437 bp were identified. These regions ranged from 1 to 96 bp. The longest intergenic spacer was located between the ND4 and ND4L genes. Furthermore, three gene pairs showed overlaps (see Table 1: negative intergenic nucleotide values) of 1 to 8 bp. All annotation details are given in Table 1. In Fig. 1 the TpitSP mitogenome genetic regions are depicted as a circular map. Table 2 further shows type base compositions, AT and GC skews for each annotation type. The typical clover-leaf secondary structures can be observed for all tRNA except trnS1 in which the dihydrouridine (DHU) arm was replaced by an unstable loop (Supplementary Fig. 1). Fig. 2 shows characteristics of the AT-rich region. At last, Fig. 3 and Supplementary Fig. 2 show phylogenies built on the mitogenomes of SP, 30 other Noctuoidea and two Geometridae species representing the outgroup (Table 3).
Table 1.
Gene features of the SP T. pityocampa mitogenome.
| Gene (anticodon) | Start | 0 | Strand | Size | Intergenic nucleotides | Start/stop codon |
|---|---|---|---|---|---|---|
| trnM(cat) | 1 | 68 | + | 68 | 0 | |
| trnI(gat) | 69 | 135 | + | 67 | 2 | |
| trnQ(ttg) | 138 | 206 | - | 69 | 63 | |
| ND2 | 270 | 1,283 | + | 1,014 | 2 | ATT/TAA |
| trnW(tca) | 1,286 | 1,351 | + | 66 | -8 | |
| trnC(gca) | 1,344 | 1,408 | - | 65 | 10 | |
| trnY(gta) | 1,419 | 1,484 | - | 66 | 4 | |
| COX1 | 1,489 | 3,021 | + | 1,533 | 2 | CGA/TAA |
| trnL2(taa) | 3,024 | 3,093 | + | 70 | 2 | |
| COX2 | 3,096 | 3,774 | + | 679 | 0 | ATG/ATT |
| trnK(ctt) | 3,775 | 3,845 | + | 71 | 10 | |
| trnD(gtc) | 3,856 | 3,931 | + | 76 | 0 | |
| ATP8 | 3,932 | 4,102 | + | 171 | -7 | ATT/TAA |
| ATP6 | 4,096 | 4,776 | + | 681 | 28 | ATG/TAA |
| COX3 | 4,805 | 5,593 | + | 789 | 7 | ATG/TAA |
| trnG(tcc) | 5,601 | 5,675 | + | 75 | 0 | |
| ND3 | 5,676 | 6,029 | + | 354 | 64 | ATT/TAA |
| trnA(tgc) | 6,094 | 6,162 | + | 69 | 1 | |
| trnR(tcg) | 6,164 | 6,230 | + | 67 | 13 | |
| trnN(gtt) | 6,244 | 6,312 | + | 69 | 24 | |
| trnS1(gct) | 6,337 | 6,402 | + | 66 | 62 | |
| trnE(ttc) | 6,465 | 6,534 | + | 70 | 6 | |
| trnF(gaa) | 6,541 | 6,612 | - | 72 | 3 | |
| ND5 | 6,616 | 8,364 | - | 1,749 | 0 | ATT/TAA |
| trnH(gtg) | 8,365 | 8,430 | - | 66 | 0 | |
| ND4 | 8,431 | 9,769 | - | 1,339 | 96 | ATG/TTT |
| ND4L | 9,866 | 10,156 | - | 291 | 5 | ATG/TAA |
| trnT(tgt) | 10,162 | 10,225 | + | 64 | 0 | |
| trnP(tgg) | 10,226 | 10,290 | - | 65 | 8 | |
| ND6 | 10,299 | 10,829 | + | 531 | -1 | ATG/TAA |
| CYTB | 10,829 | 11,983 | + | 1,155 | 3 | ATG/TAA |
| trnS2(tga) | 11,987 | 12,053 | + | 67 | 22 | |
| ND1 | 12,076 | 13,011 | - | 936 | 0 | GTG/TAA |
| trnL1(tag) | 13,012 | 13,082 | - | 71 | 0 | |
| 16S rRNA | 13,083 | 14,463 | - | 1,381 | 0 | |
| trnV(tac) | 14,464 | 14,531 | - | 68 | 0 | |
| 12S rRNA | 14,532 | 15,309 | - | 778 | 0 | |
| AT-rich region | 15,310 | 15,729 | + | 420 | 0 |
The gene names, the coordinates, the orientation, the size, the gene overlaps/intergenic spacers (column 6) and the start/stop codons of the SP mitogenome are listed.
Fig. 1.
Map of the T. pityocampa SP mitogenome generated with OGDRAW. Annotated genes are highlighted.
Table 2.
Base composition and skewness indices for each gene region of the SP T. pityocampa mitogenome.
| Type | Size [bp] | A% | G% | T% | C% | A+T% | ATskewness | GCskewness |
|---|---|---|---|---|---|---|---|---|
| Entire mitogenome | 15,729 | 40.6 | 7.6 | 37.3 | 14.6 | 77.8 | 0.042 | -0.318 |
| Protein-coding genes | 11,222 | 32.6 | 12.1 | 43.0 | 12.4 | 75.5 | -0.138 | -0.009 |
| tRNA | 1,507 | 41.4 | 11.3 | 39.3 | 7.9 | 80.8 | 0.025 | 0.179 |
| rRNA | 2,159 | 39.2 | 13.1 | 43.1 | 4.6 | 82.3 | -0.048 | 0.482 |
| AT-rich region | 420 | 43.6 | 2.9 | 50.0 | 3.6 | 93.6 | -0.069 | -0.111 |
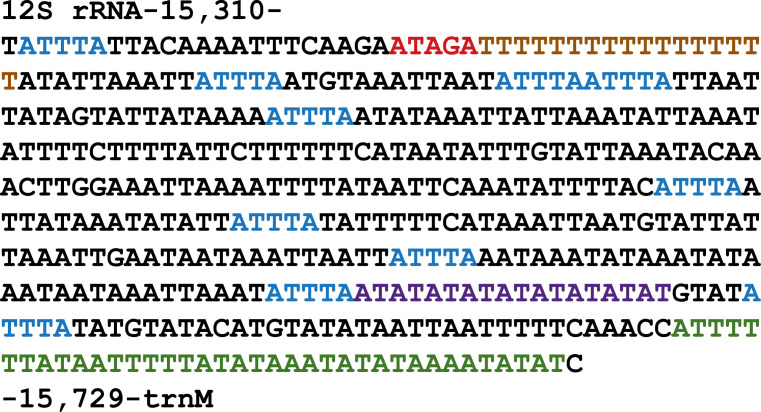
Fig. 2.
Characteristics of the AT-rich region of the T. pityocampa SP mitogenome.
The AT-rich region consists of an ATAGA (red) motif followed by a 17 bp poly-T stretch (brown). Furthermore, the control region contains ten ATTTA motifs (blue) and an (AT)9 microsatellite stretch (purple). The region ends with a polyT/polyA region (green).
Fig. 3.
Phylogenetic tree built on concatenated alignments of the nucleotide sequences of the 13 protein-coding and both rRNA genes applying a Maximum Likelihood approach with PHYML. For both T. pityocampa populations the countries are given.
Table 3.
Lepidoptera mitogenomes used to build the phylogenetic trees. For both T. pityocampa populations the countries are given.
| Super-family | Family | Subfamily | Species | Genbank no. |
|---|---|---|---|---|
| Geometroidea | Geometridae | Ennominae | Biston panterinaria | NC_020004 |
| Geometroidea | Geometridae | Ennominae | Phthonandria atrilineata | NC_010522 |
| Noctuoidea | Erebidae | Aganainae | Asota plana lacteata | KJ173908 |
| Noctuoidea | Erebidae | Arctiinae | Amata formosae | NC_021416 |
| Noctuoidea | Erebidae | Arctiinae | Callimorpha dominula | NC_027094 |
| Noctuoidea | Erebidae | Arctiinae | Lemyra melli | NC_026692 |
| Noctuoidea | Erebidae | Arctiinae | Vamuna virilis | NC_026844 |
| Noctuoidea | Erebidae | Erebinae | Grammodes geometrica | NC_039810 |
| Noctuoidea | Erebidae | Herminiinae | Hydrillodes lentalis | NC_041106 |
| Noctuoidea | Erebidae | Hypeninae | Paragabara curvicornuta | NC_034638 |
| Noctuoidea | Erebidae | Lymantriinae | Euproctis pseudoconspersa | NC_027145 |
| Noctuoidea | Erebidae | Lymantriinae | Gynaephora aureata | NC_029162 |
| Noctuoidea | Erebidae | Lymantriinae | Lachana alpherakii | NC_029172 |
| Noctuoidea | Erebidae | Lymantriinae | Lymantria dispar | NC_012893 |
| Noctuoidea | Euteliidae | Euteliinae | Eutelia adulatricoides | NC_026840 |
| Noctuoidea | Noctuidae | Heliothinae | Helicoverpa armigera | NC_014668 |
| Noctuoidea | Noctuidae | Noctuinae | Agrotis ipsilon | NC_022185 |
| Noctuoidea | Noctuidae | Noctuinae | Mythimna separata | NC_023118 |
| Noctuoidea | Noctuidae | Noctuinae | Sesamia inferens | NC_015835 |
| Noctuoidea | Noctuidae | Noctuinae | Spodoptera frugiperda | NC_027836 |
| Noctuoidea | Noctuidae | Noctuinae | Striacosta albicosta | NC_025774 |
| Noctuoidea | Noctuidae | Plusiinae | Ctenoplusia agnata | NC_021410 |
| Noctuoidea | Noctuidae | Plusiinae | Diachrysia nadeja | NC_053855 |
| Noctuoidea | Noctuidae | Plusiinae | Trichoplusia ni | NC_045936 |
| Noctuoidea | Nolidae | Chloephorinae | Gabala argentata | NC_026842 |
| Noctuoidea | Nolidae | Chloephorinae | Sinna extrema | NC_045120 |
| Noctuoidea | Nolidae | Risobinae | Risoba prominens | NC_026841 |
| Noctuoidea | Notodontidae | Phalerinae | Phalera flavescens | NC_016067 |
| Noctuoidea | Notodontidae | Pygaerinae | Clostera anachoreta | NC_034740 |
| Noctuoidea | Notodontidae | Pygaerinae | Clostera anastomosis | NC_041140 |
| Noctuoidea | Notodontidae | Thaumetopoeinae | Ochrogaster lunifer | NC_011128 |
| Noctuoidea | Notodontidae | Thaumetopoeinae | Thaumetopoea pityocampa (Italy) | NC_053256 |
| Noctuoidea | Notodontidae | Thaumetopoeinae | Thaumetopoea pityocampa (Portugal) | OM478592 |
2. Experimental Design, Materials and Methods
2.1. DNA sequence libraries
To generate the present SP mitogenome assembly we used two shotgun whole-genome paired-end sequence libraries originally built to produce the reference nuclear genome of T. pityocampa SP [1]. Each of these libraries was built from one SP male which was sampled between 2010 and 2012 from the Mata Nacional de Leiria in Portugal (39°47’N, 8°58’W). The libraries had insert sizes of 300 bp and 600 bp, respectively, and were sequenced on an Illumina HiSeq2000 platform at the Edinburgh Genomics sequencing facility in Scotland, UK.
The raw whole-genome shot-gun sequence reads were trimmed and cleaned with Trimmomatic v0.22 [5] (specific parameters were: window size of 20 bp, minimum PHRED quality of 20, minimum length of 70 bp), and subsequently assembled by Eurofins MWG Operon using their own bioinformatics pipeline (CONVEY, http://www.conveycomputer.com), implementing multiple Velvet assemblies.
2.2. Mitogenome assembly and feature analyses
The SP mitogenome was generated in two subsequent steps. First, as described in [1] the nuclear genome assembly contained one scaffold of 15,717 bp which was identified as partial mitochondrion based on blastn searches against the NCBI nucleotide database. In a second step, MITObim v1.9 [6] was applied to extend and refine this mitogenome backbone using the cleaned (--clean) Illumina paired-end (--pair) reads with a k-mer size of 41 bp (--kbait 41), a maximum of 5 mismatches per mapped read (--mismatch 5) and a minimum average contig coverage of 500 reads per base (--min_cov 500). Furthermore, the PPM mitogenome was globally aligned with MAFFT v7.471 [7] (executable mafft-ginsi, using a maximum of 1,000 refinement iterations (--maxiterate 1000)) to five other Notontidae mitogenomes of the following species: Thaumetopoea pityocampa (Italian population, Genbank no. NC_053256), Ochrogaster lunifer (Genbank no. NC_011128), Clostera anastomosis (Genbank no. NC_041140), Clostera anachoreta (Genbank no. NC_034740), Phalera flavescens (Genbank no. NC_016067). This allowed to identify overlapping repeat regions at both sequence termini which were then identified as assembly artefacts and thus manually removed (linear vs circular structure). This resulted in a mitogenome size of 15,729 bp. Annotation of the genetic regions was conducted by the MITOS web server v2 [8], using as reference set ‘RefSeq 89 Metazoa‘ and the genetic code ‘5‘ for invertebrates. Furthermore, the MITOS gene annotations were manually curated by aligning the predicted genes via web service of NCBI blastn [9] (with default parameters, version of January 2022) to gene annotations of other Notodontidae mitogenomes (same as described above). The web tool OGDRAW v1.3.1 [10] was run to create the circular map of the SP T. pityocampa mitogenome with its annotations. We applied the wordcount program of the EMBOSS toolkit v6.6.0.0 [11] to calculate the base content of each genetic region type. The genomic compositional asymmetry of the SP mitogenome of each specific annotation type region was described by the AT/GC skewness indices which were calculated as follows with each base count being represented by the corresponding base letter: AT skew = [A-T] / [A + T] and GC skew = [G-C] / [G + C].
2.3. Phylogenetic analyses
For phylogenetic trees were built on 33 Lepidoptera mitogenomes – including 30 Noctuoidea species (Table 3). Two representatives of the Geometroidea were used as outgroup to root the trees following the conclusions of Wahlberg et al. [12]. We obtained the nucleotidic sequences of the 13 PCGs and 2 rRNA genes from the NCBI ‘nucleotide’ database (https://www.ncbi.nlm.nih.gov/nuccore, July 2021). The superfamily Noctuoidea was chosen for taxon sampling, encompassing five species belonging to the Notodontidae (including T. pityocampa). Regarding T. pityocampa, we included the mitogenomes of the Portuguese TpitSP (provided here) and an Italian typical population [3].
The nucleotide sequences of all species were grouped into 15 gene-specific sequence sets, corresponding to 13 PCG and 2 rRNA genes. Global sequence alignment on each gene set using the Needleman-Wunsch algorithm was conducted with the program MAFFT v7.471 (executable mafft-ginsi for PCGs and mafft-qinsi for rRNA sequences), applying a maximum of 1,000 refinement iterations (--maxiterate 1000). Finally, in-house Perl and Python and scripts were run to concatenate for each species all gene-specific alignments following the same gene order.
A local installation of MrBayes v3.2.7a [13] and the PHYML web server v3.0 [14] were run to calculate phylogenetic tree structures. For each gene a distinct partition was configured in the MrBayes analysis with the following parameters each: GTR substitution model (nst = 6) with gamma-distributed rate variation (rates = invgamma) across sites and a proportion of invariable sites (GTR + I + #). As substitution model the standard nucleotide model (nucmodel = 4by4) was chosen with priors being left to default settings. We configured MrBayes to use 100,000 samples (ngen = 1,000,000, samplefreq = 10) in order to generate phylogenetic trees. Additional settings were: printfreq = 100, diagnfreq = 1,000, nchains = 4, savebrlens = yes, starttree = random, startparams = reset, outgroup bipan (corresponding to Biston panterinaria), sump burnin = 25,000, sumt burnin = 25,000. Regarding the PHYML analysis, the Smart Model Selection [15] parameter was chosen with the Akaike Information Criterion and the tree searching method BIONJ being set. Furthermore, 1,000 bootstraps were applied. Finally, the program FigTree v1.4.4 [16] was run to draw the phylogenetic tree.
CRediT Author Statement
Bernhard Gschloessl: Conceptualization, Methodology, Formal analysis, Validation, Writing – review & editing; Carole Kerdelhué: Project administration, Funding acquisition, Writing – review & editing.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
The authors would like to thank Gael J Kergoat for his advice in phylogenetic analyses. Grants were provided by the French National Agency for Research (ANR-10-JCJC-1705-01, GENOPHENO) and by the Institut National de la Recherche Agronomique (INRA AIP BioRessources 2012, GAPP). This work has been realized with the support of MESO@LR-Platform at the University of Montpellier.
Footnotes
Supplementary material associated with this article can be found in the online version at doi:10.1016/j.dib.2022.108376.
Appendix. Supplementary materials
Supplementary Fig. 1. MITOS-predicted secondary tRNA structures of the SP Thaumetopoea pityocampa mitogenome.
Supplementary Fig. 2. Phylogenetic tree constructed with a Bayesian inference approach using MrBayes on concatenated alignments of the nucleotide sequences of the 13 protein-coding and both rRNA genes. For both T. pityocampa populations the countries are given.
Data Availability
BioSample of the TpitSP mitogenome (Original data) (NCBI BioSample).
BioProject comprising the TpitSP mitogenome (Original data) (NCBI BioProject).
Genbank entry of the TpitSP mitogenome (Original data) (NCBI Genbank).
References
- 1.Gschloessl B., Dorkeld F., Berges H., Beydon G., Bouchez O., Branco M., Bretaudeau A., Burban C., Dubois E., Gauthier P., Lhuillier E., Nichols J., Nidelet S., Rocha S., Sauné L., Streiff R., Gautier M., Kerdelhué C. Draft genome and reference transcriptomic resources for the urticating pine defoliator Thaumetopoea pityocampa (Lepidoptera: Notodontidae) Mol. Ecol. Resour. 2018;18:602–619. doi: 10.1111/1755-0998.12756. [DOI] [PubMed] [Google Scholar]
- 2.Boore J.L. Animal mitochondrial genomes. Nucleic Acids Res. 1999;27:1767–1780. doi: 10.1093/nar/27.8.1767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wu K., Yang J., Ni Y., Liu Q. Identification and analysis of the complete mitochondrial genome of Thaumetopoea pityocampa (Lepidoptera: Notodontidae) Mitochondrial DNA Part B. 2019;4:3654–3656. doi: 10.1080/23802359.2019.1678422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Zhu X.-Y., Xin Z.-Z., Liu Y., Wang Y., Huang Y., Yang Z.-H., Chu X.-H., Zhang D.-Z., Zhang H.-B., Zhou C.-L., Wang J.-L., Tang B.-P., Liu Q.-N. The complete mitochondrial genome of Clostera anastomosis (Lepidoptera: Notodontidae) and implication for the phylogenetic relationships of Noctuoidea species. Int. J. Biol. Macromol. 2018;118:1574–1583. doi: 10.1016/j.ijbiomac.2018.06.188. [DOI] [PubMed] [Google Scholar]
- 5.Bolger A.M., Lohse M., Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–2120. doi: 10.1093/bioinformatics/btu170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hahn C., Bachmann L., Chevreux B. Reconstructing mitochondrial genomes directly from genomic next-generation sequencing reads–a baiting and iterative mapping approach. Nucleic Acids Res. 2013;41:e129. doi: 10.1093/nar/gkt371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Katoh K., Standley D.M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 2013;30:772–780. doi: 10.1093/molbev/mst010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bernt M., Donath A., Jühling F., Externbrink F., Florentz C., Fritzsch G., Pütz J., Middendorf M., Stadler P.F. MITOS: improved de novo metazoan mitochondrial genome annotation. Mol. Phylogenet. Evol. 2013;69:313–319. doi: 10.1016/j.ympev.2012.08.023. [DOI] [PubMed] [Google Scholar]
- 9.Johnson M., Zaretskaya I., Raytselis Y., Merezhuk Y., McGinnis S., Madden T.L. NCBI BLAST: a better web interface. Nucleic Acids Research. 2008;36:W5–W9. doi: 10.1093/nar/gkn201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Greiner S., Lehwark P., Bock R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 2019;47:W59–W64. doi: 10.1093/nar/gkz238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Rice P., Longden I., Bleasby A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet. 2000;16:276–277. doi: 10.1016/s0168-9525(00)02024-2. [DOI] [PubMed] [Google Scholar]
- 12.Wahlberg N., Wheat C.W., Peña C. Timing and patterns in the taxonomic diversification of Lepidoptera (butterflies and moths) PLoS ONE. 2013;8:e80875. doi: 10.1371/journal.pone.0080875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ronquist F., Teslenko M., van der Mark P., Ayres D.L., Darling A., Höhna S., Larget B., Liu L., Suchard M.A., Huelsenbeck J.P. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012;61:539–542. doi: 10.1093/sysbio/sys029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Guindon S., Lethiec F., Duroux P., Gascuel O. PHYML Online–a web server for fast maximum likelihood-based phylogenetic inference. Nucleic Acids Res. 2005;33:W557–W559. doi: 10.1093/nar/gki352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lefort V., Longueville J.-E., Gascuel O. SMS: Smart Model Selection in PhyML. Mol. Biol. Evol. 2017;34:2422–2424. doi: 10.1093/molbev/msx149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.A. Rambaut, A.J. Drummond, FigTree version 1.4. 0, 2012. http://tree.bio.ed.ac.uk/software/figtree/.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Fig. 1. MITOS-predicted secondary tRNA structures of the SP Thaumetopoea pityocampa mitogenome.
Supplementary Fig. 2. Phylogenetic tree constructed with a Bayesian inference approach using MrBayes on concatenated alignments of the nucleotide sequences of the 13 protein-coding and both rRNA genes. For both T. pityocampa populations the countries are given.
Data Availability Statement
BioSample of the TpitSP mitogenome (Original data) (NCBI BioSample).
BioProject comprising the TpitSP mitogenome (Original data) (NCBI BioProject).
Genbank entry of the TpitSP mitogenome (Original data) (NCBI Genbank).