Abstract
The D2 dopamine receptor (D2R) is the primary site of the therapeutic action of antipsychotics and is involved in essential brain functions relevant to schizophrenia, such as attention, memory, motivation, and emotion processing. Moreover, the gene coding for D2R (DRD2) has been associated with schizophrenia at a genome-wide level. Recent studies have shown that a polygenic co-expression index (PCI) predicting the brain-specific expression of a network of genes co-expressed with DRD2 was associated with response to antipsychotics, brain function during working memory in patients with schizophrenia, and with the modulation of prefrontal cortex activity after pharmacological stimulation of D2 receptors. We aimed to investigate the relationship between the DRD2 gene network and in vivo striatal dopaminergic function, which is a phenotype robustly associated with psychosis and schizophrenia. To this aim, a sample of 92 healthy subjects underwent 18F-DOPA PET and was genotyped for genetic variations indexing the co-expression of the DRD2-related genetic network in order to calculate the PCI for each subject. The PCI was significantly associated with whole striatal dopamine synthesis capacity (p = 0.038). Exploratory analyses on the striatal subdivisions revealed a numerically larger effect size of the PCI on dopamine function for the associative striatum, although this was not significantly different than effects in other sub-divisions. These results are in line with a possible relationship between the DRD2-related co-expression network and schizophrenia and extend it by identifying a potential mechanism involving the regulation of dopamine synthesis. Future studies are needed to clarify the molecular mechanisms implicated in this relationship.
Subject terms: Psychosis, Schizophrenia, Genetics of the nervous system
Introduction
The D2 dopamine receptor (D2R) is a G protein-coupled receptor coded by the DRD2 gene and is involved in essential brain functions such as learning, memory, locomotion, attention, motivation, sleep, emotion processing, reproductive behaviour1–3. The D2R is also the primary site of the therapeutic action of antipsychotics4–7. Furthermore, one of the schizophrenia-associated loci from Genome-Wide Association Studies (GWAS)8,9 includes the D2R coding gene (DRD2), implicating this gene in the pathophysiology of schizophrenia10.
Genetic variations within DRD2 have been associated with brain-related phenotypes, including working memory, sustained attention, variable attention control, emotion processing, dopamine binding in the striatum, suggesting that genetic mechanisms influence the effects of the D2R on brain function11–17. However, it is unlikely that genetic variations within a single gene explain the entire physiology related to specific brain phenotypes. In this regard, previous investigations have elucidated that genes involved in complex traits do not work in isolation but operate in networks of interacting genes18–22 acting via molecular pathways23–25. Genetic networks can be investigated in detail using methods for the analysis of gene co-expression patterns26,27. This approach is based on the evidence that the expression of different genes is influenced by common regulatory molecules, and that such gene expressions correlate24,28–32. Co-expressed genes are often related in terms of function33,34. A widely used technique to study gene co-expression is the weighted gene co-expression network analysis (WGCNA). WGCNA represents correlated gene expression into a graph that is designed to be scale invariant, hence reflecting the basic property of biological networks that include highly connected central hubs and more peripheral genes. Hierarchical clustering is used in WGCNA to define gene sets, called modules, that are tightly co-expressed. This approach has been used to identify, in post-mortem dorsolateral prefrontal cortex of healthy controls, a network of genes co-expressed with DRD235, including genes associated with schizophrenia identified in the PGC29 and PGC3 GWAS36. A follow-up study has supported with in vitro evidence the link between some of these genes and identified potential co-regulators32. Interestingly, a Polygenic Co-expression Index (PCI) predicting the brain-specific expression of this network of co-expressed genes was associated with response to antipsychotics and prefrontal inefficiency during working memory35, which has been consistently associated with schizophrenia37. Moreover, healthy subjects with higher PCI showed increased activation in the prefrontal cortex and longer reaction times when performing a working memory task35. Interestingly, in a recent network control theory study38 the same PCI has been shown to be related to dynamical brain state transitions during working memory in healthy volunteers. Furthermore, this PCI has been associated with within-subject variation of prefrontal cortex activity following pharmacological stimulation of D2R in a double-blind crossover design39.
While these studies focused on the frontal cortex, DRD2 has its highest expression in the striatum40. Interestingly, molecular imaging studies show evidence that presynaptic striatal dopamine dysfunction plays an important role in abnormal reward processing and anomalies of other aspects of cognitive function41,42. Moreover, elevated striatal dopamine synthesis and release capacity are associated with schizophrenia43–53, psychotic symptoms54 and risk of psychosis55,56.
Whilst the findings discussed above show that the DRD2 gene network is associated with cortical brain function relevant to cognitive phenotypes of schizophrenia57, it remains unknown if and how the genetic underpinnings of cortical dopaminergic function are related to striatal dopaminergic phenotypes associated with psychosis. The exploration of this relationship can be considered as particularly relevant in view of the connections between cortex and striatum58. The aim of the present study is to investigate the relationship between striatal dopamine synthesis capacity and co-expression of the DRD2-related genetic network35. To this aim, we analysed data from a cohort of healthy subjects that underwent 18F-DOPA PET and were genome-wide genotyped; we used the genetic variants indexing the co-expression of the DRD2-related genetic network to compute an individual PCI35. We hypothesised that higher PCI, which has been previously associated with greater prefrontal BOLD response (see also59) and longer reaction times during working memory processing35, would be associated with higher striatal dopamine synthesis capacity—thus outlining a consistent pattern of results resembling the physiological observations in patients with schizophrenia.
Methods
Participants
A total of 92 healthy subjects (demographics in Table 1) underwent 18F-DOPA PET scans60. The study was conducted in accordance with the Declaration of Helsinki and Good Clinical Practice. All participants gave informed written consent. The study was approved by the Administration of Radioactive Substances Advisory Committee (ARSAC), the South London and Maudsley/Institute of Psychiatry NHS Trust, the London Bentham Research Ethics Committee, and the Hammersmith Research Ethics Committee.
Table 1.
Demographic characteristics of the sample.
| Total | |
|---|---|
| N | 92 |
| Age (yr ± SD) | 29.93 ± 8.84 |
| Gender (male/female) | 52/40 |
| PET scanner (scanner 1/scanner 2/scanner 3) | 37/35/20 |
| Kicer (1/min) whole striatum (mean ± SD) | 0.0129 ± 0.0012 |
| Kicer (1/min) associative striatum (mean ± SD) | 0.0128 ± 0.0012 |
| Kicer (1/min) limbic striatum (mean ± SD) | 0.0130 ± 0.0014 |
| Kicer (1/min) sensorimotor striatum (mean ± SD) | 0.0132 ± 0.0015 |
| PCI (mean ± SD) | − 0.0085 ± 0.0986 |
Inclusion criteria were: age range 18–65 years, no history of major medical condition, good physical health. Exclusion criteria were: significant medical disorder or treatment, history of psychiatric illness (assessed using the Structured Clinical Interview for DSM-IV Axis I Disorders) including alcohol or substance abuse or dependence. The dataset has been gathered from our publicly available imaging data archive (https://maudsleybrc.nihr.ac.uk/research/precision-psychiatry/neuroimaging/neuroimaging-database-node/). The PET data have been previously published61–65, but the integration with the PCI has not been published before.
Polygenic co-expression index
DNA was extracted from whole blood samples or cheek swabs using standard procedures66. Genome-wide genotyping was performed at Cardiff University, using HumanCore Exome 1.1 arrays ("Psych-chip", Illumina, San Diego, California, USA).
A PCI was calculated as previously described35. Briefly, a DRD2 co-expression gene set, including 85 genes, was identified with a Weighted Genes Co-expression Network Analysis67 using the post mortem frontal cortex mRNA expression Braincloud database68. A set of 8 SNPs (CHIT1 rs2486064, GPLD1 rs6902039, OSR1 rs851436, POP1 rs9297283, SDK2 rs1294071, DHX33 rs1805453, BTG4 rs1121391, AGR2 rs1037791) associated with the first principal component of gene set co-expression was used to compute the PCI; a weight based on the co-expression profile of the gene set was assigned to each genotype of each SNP (Table S1). Genotyping was conducted for these SNPs. Genotype quality control for these SNPs was performed according to standard parameters69. Briefly, these included an individual missingness rate < 0.98, a SNP call rate > 0.98 and a Hardy–Weinberg equilibrium (HWE) p value > 10−4, as computed by the PLINK v1.9 software70.
Population stratification
The Principal Components Analysis in Related samples (PC-AiR) method71 was used in R (GENESIS R/Bioconductor package72) on the full set of genotypes to generate the top 10 principal components of the sample, which were included as covariates of no interest in all the analyses, in order to correct for population stratification.
PET scanning
18F-DOPA PET scans were performed to measure dopamine synthesis capacity (indexed as the influx rate constant Kicer)73.
Image acquisition
Images were acquired in three-dimensional mode using three different PET scanners: an ECAT HR + 962 PET scanner (CTI/Siemens, Knoxville, Tennessee) and two Siemens Biograph HiRez XVI PET-CT scanners (Siemens Healthcare, Erlangen, Germany). After the administration of approximately 150 MBq of 18F-DOPA, dynamic PET data were acquired over a period of 95 min as previously described61–64,74.
Image processing
The frames were aligned using a mutual information algorithm75. A movement-corrected dynamic image was then used in the analysis. A tracer-specific (18F-DOPA) template76 was normalised together with a striatal probabilistic atlas77 to the individual PET summation images. The influx constant (Kicer) for striatum was calculated using the cerebellum as a reference region78. For the exploratory analyses, the striatum was sub-divided into limbic, associative and sensorimotor parts on the basis of function and the topography of brain projections from limbic, associative and sensorimotor cortical areas to the striatum48,77,78.
Statistical analysis
The effect of the PCI on whole striatal Kicer was tested using a linear model (lm) regression in R79 with age, gender, PET scanner and the first 10 genetic principal components as covariates of no interest in view of their potential effect on dopamine synthesis capacity80,81. To facilitate the interpretation of the results, PCI values were standardised using the scale() function in R before being entered in the model82. Injected dose of radiotracer was not considered, as it is not associated with 18F-DOPA Kicer estimates74. A significance threshold of α < 0.05 was used. Separate exploratory analyses were conducted to test the effect of the PCI on associative striatum, limbic striatum and sensorimotor striatum Kicer. R79 was used for all the statistical analyses. The R package ggplot283 was used to plot the main results. To exclude the presence of outliers, the Rosner's test function ("rosnerTest") of the R package EnvStats84 was used to remove extreme observations.
Results
Demographic (± SD) and Kicer values included are reported in Table 1.
The Rosner's test did not reveal any outliers. PCI was significantly associated with whole striatal dopamine synthesis capacity (t value = 2.106, p = 0.038). Figure 1 illustrates a positive correlation between whole striatum Kicer (y axis) and PCI. PET scanners, included as covariates of no interest, did not show a statistically significant association with dopamine synthesis capacity (t value = 1.603, p = 0.112).
Figure 1.
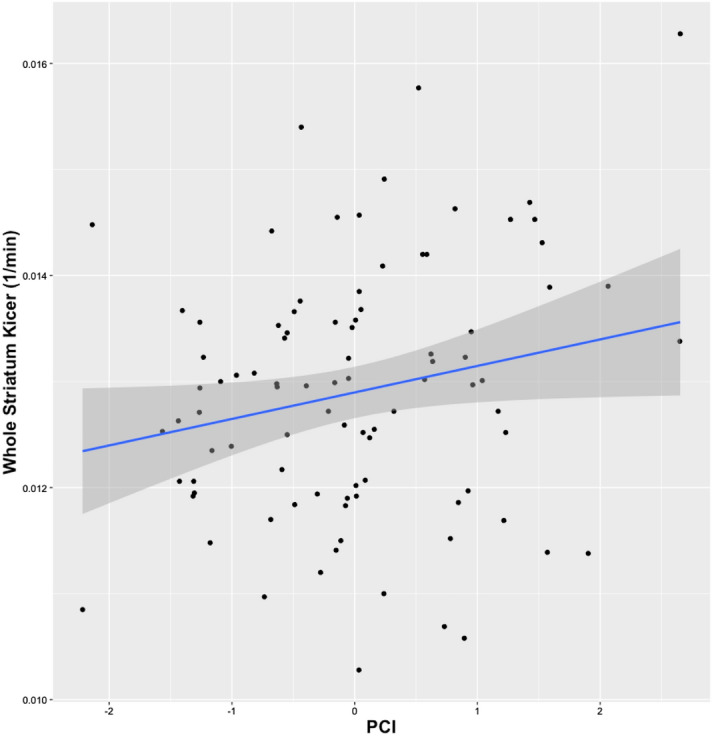
Scatterplot illustrating the correlation between whole striatum Kicer (y axis) and PCI.
The exploratory analyses in the striatal subdivisions revealed an effect of the PCI on dopamine synthesis capacity for the associative striatum only (t value = 2.063, p = 0.042) (Fig. 2), while there was not a significant correlation with limbic striatum (t value = 1.957, p = 0.054) or sensorimotor striatum (t value = 1.841, p = 0.069). The interaction among striatal subdivision, PCI, and Kicer was not significant (p = 0.738).
Figure 2.
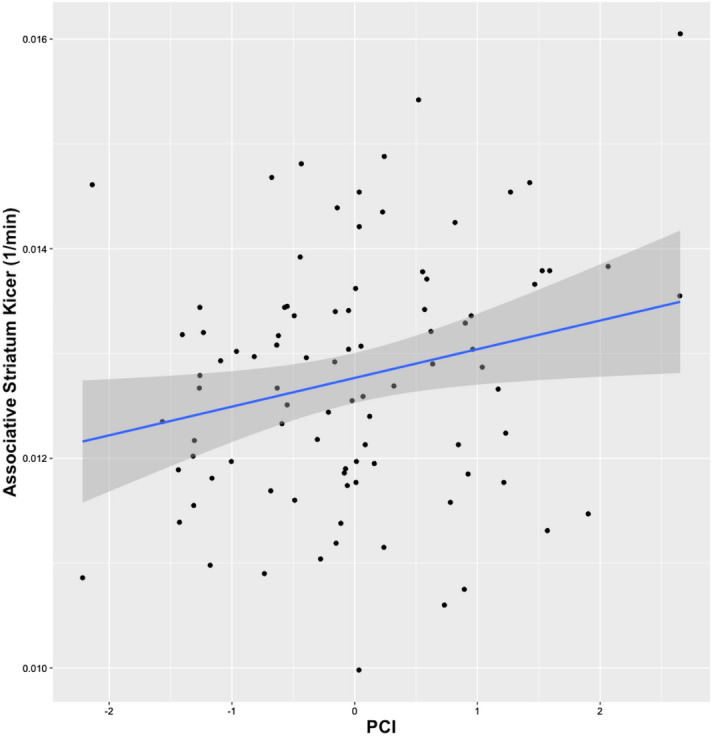
Scatterplot illustrating the correlation between associative striatum Kicer (y axis) and PCI.
Discussion
The present study shows for the first time an in vivo association between striatal dopamine synthesis capacity and a DRD2-related co-expression score in a cohort of healthy subjects. Specifically, high polygenic co-expression index, reflecting greater prefrontal co-expression of a DRD2-related genetic network, was associated with elevated striatal dopamine synthesis capacity. These results suggest that, besides DRD2, several genes and related products may be relevant to the modulation of striatal dopamine function.
Gene co-expression networks have been instrumental in identifying gene sets associated with antipsychotic treatment response30,35, phenotypes associated with schizophrenia25,28,35,85, clinical state and risk for schizophrenia31, and changes in prefrontal function after D2R stimulation39. As reviewed previously, increased dopamine synthesis capacity represents a phenotype associated with schizophrenia47,86,87. Thus, the results of the present study suggest a possible relationship between the DRD2-related co-expression network identified and schizophrenia. Consistently, the exploratory analyses in the different striatal subdivisions suggest that the PCI is associated with dopamine capacity in the associative striatum, which is the striatal region showing greater dopaminergic dysfunction in patients with schizophrenia compared with other striatal subdivisions according to meta-analytic evidence88. Nevertheless, it is important to note that the analyses in the different striatal subdivisions were only exploratory and there was no significant difference between effect sizes across striatal subdivisions. Additionally, the association with the associative striatum Kicer would not survive correction for multiple comparisons. Therefore, these results should be considered as exploratory and require further evaluation in a larger sample.
The hypothesis of a positive relationship between PCI and striatal dopamine synthesis capacity was based on a study35 demonstrating in healthy subjects an association of this index with prefrontal inefficiency during working memory, another phenotype related to schizophrenia. However, it is noteworthy that previous studies have shown both positive89–91 and negative92,93 correlations between striatal dopamine synthesis capacity and working memory. Thus, a study investigating striatal dopamine function, working memory efficiency and PCI in the same sample would be necessary to elucidate the relationship between these factors.
Interestingly, fifteen genes of the DRD2 Co-Expression Network (ACR, ALDH3A1, BTN3A1, CALHM3, CES3, DRD2, EFCAB6, GALNT10, GATAD2A, GLI1, HIST1H1E, HIST1H3G, IL31, RBM6, SLC28A1) are located within schizophrenia-associated loci in the latest Psychiatric Genetic Consortium investigation36. Notably, GATAD2A is among the genes resulting from the PGC3 prioritisation analysis due to its eQTL co-localisation profile36. Accordingly, it is considered a plausible causal gene for schizophrenia94. This gene codes for the protein GATA zinc finger domain containing 2A, a transcriptional repressor95, which is preferentially expressed during foetal brain development96. Its involvement in cell proliferation97 indicates a key role in development94. Furthermore, it has been implicated in schizophrenia through its involvement in the regulation of gene expression98,99. Consistently, it is upregulated in the hippocampus of patients with schizophrenia compared with healthy controls94.
Moreover, it has been recently demonstrated32 that the expression of genes of the DRD2 co-expression module can be regulated by NURR1, a transcription factor regulating genes involved in the dopaminergic system100. As D2R is a potent NURR1 activator101,102, it has been hypothesised that antipsychotics, through the blockade of the D2R, can impact the expression of NURR1, which in turn can regulate the transcription of the genes included in the DRD2 co-expression module32. The results of the present study—indicating a relationship between the DRD2 co-expression network and an established phenotype linked to schizophrenia such as PET-estimated dopamine synthesis capacity—are consistent with the hypothesis of the involvement of the genes of this module in the pathophysiology of schizophrenia and mechanisms underlying the response to antipsychotics.
Notably, the approach used in the present study is data-driven and the genes within the network are not pre-defined; thus, the mechanisms through which the proteins coded by the genes of this network interact with the dopaminergic pathway still need to be clarified. In fact, it needs to be understood how the DRD2 co-expression network influences striatal presynaptic dopamine synthesis capacity. In this context, it should be considered that post-synaptic D2 receptors play a role in the regulation of dopamine synthesis and release through inhibitory feedback loops103,104. It is also possible that the PCI reflects a different expression of the D2 autoreceptors, which regulate dopamine synthesis, although it should be noted that this score was developed analysing the expression of transcripts including exon 6, which is characteristic of the long isoform of D2R more often found post-synaptically105,106. Moreover, in view of the fact that the DRD2 co-expression gene-set indexed by the PCI is enriched for “negative regulation of dopamine secretion (GO:0033602)”35,39, preclinical studies are needed to test the hypothesis that the transcriptomic context of DRD2 influences dopamine presynaptic signalling.
Furthermore, in order to examine the potential involvement of this co-expression network in the regulation of expression and availability of the post-synaptic D2 receptors, it would be helpful to investigate the in vivo relationship between PCI and D2 receptor availability through studies using other PET tracers (e.g. 11C-raclopride).
The present study was conducted on healthy subjects; thus, the results were not influenced by medication or disease status. A key next step is thus to explore the effect of the DRD2-Polygenic Co-expression Index on dopamine function in disorders where involvement of the dopamine system has been demonstrated, such as psychosis, addiction, bipolar disorder54,107,108.
A potential limitation of the study is the use of data from three different PET scanners. However, the scanner was used as a covariate of no interest. Furthermore, we did not find a statistically significant association of PET scanner with Kicer; consistently, our recent investigation on the effect of the scanner in a similar dataset acquired from three different PET tomographs (Siemens Biograph 6 Hi-Rez, Siemens Biograph 6TruePoint, ECAT/EXACT3D) with an injected radioactivity below 200 MBq and acquisition time of 95 min did not reveal significant effects109.
Moreover, it should be considered that the DRD2 co-expression pathway, and therefore the PCI used in this study, was calculated by using post mortem mRNA from the frontal cortex35,68, thus it would be interesting to test if the DRD2 co-expression network remains the same in the striatum and the midbrain, where the dopamine neuron cell bodies are located.
Furthermore, in the present study, we did not examine dopamine function in the frontal cortex, due to lower 18F-DOPA signal reliability in frontal cortical regions when quantified without arterial blood input function76,110. Therefore, a study using PET tracers more suitable for the measurement of the cortical dopamine system59,111 would be helpful in understanding the relationships between PCI, striatal and cortical dopamine systems.
Conclusions
The results from the present study indicate that a polygenic score indexing a DRD2-related co-expression network is associated with striatal dopamine function measured in vivo with 18F-DOPA imaging. Our findings suggest that the same genetic variants associated with prefrontal inefficiency during working memory are also associated with greater estimated dopamine synthesis in the striatum. In view of the hypothesised link between striatal hyperdopaminergia and prefrontal hypodopaminergia59,112, it is tempting to observe that these variants originally found analysing the prefrontal cortex may have more widespread system-level correlates38.
Supplementary Information
Acknowledgements
This research was funded in whole, or in part, by the Wellcome Trust [Grant number 094849/Z/10/Z]. For the purpose of open access, the author has applied a CC BY public copyright licence to any Author Accepted Manuscript version arising from this submission. MV is supported by MIUR, Italian Ministry for Education, under the initiatives “Departments of Excellence” (Law 232/2016), by National Institute for Health Research (NIHR) Maudsley Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King’s College London and by the Wellcome Trust Innovator Award 215747/Z/19/Z. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health. AFP was supported by an Academy of Medical Sciences “Springboard” award (SBF005\1083). JTRW is an investigator on a grant from Takeda Pharmaceuticals Ltd. to Cardiff University, for a project unrelated to the work presented here. KG is supported by the National Institute for Health Research (NIHR) Biomedical Research Centre at South London and Maudsley (SLaM) NHS Foundation Trust and King’s College London. TD is supported by the NIHR. ODH is supported by Medical Research Council-UK (no. MC_U120097115) and Wellcome Trust (no. 094849/Z/10/Z).
Author contributions
E.D., G.P., G.B., A.B., O.D.H. conceptualised the study and wrote the first draft of the manuscript. E.D., T.D., M.V., S.J., M.R., M.A.P.B., S.F.W., I.B., J.T.R.W., A.F.P., O.D.H. performed data collection and curation. E.D., G.P., A.F.P., T.D., M.V., P.T., L.S., K.G. performed data analyses. All the authors provided major contributions to the interpretation of the data, writing, and critical review of the manuscript. All the authors approved the final version of the manuscript.
Data availability
The PET data are available in The NeurOimaging DatabasE (NODE) repository (https://maudsleybrc.nihr.ac.uk/research/precision-psychiatry/neuroimaging/neuroimaging-database-node/) upon request. The data68 used for the WGCNA performed to identify the DRD2 co-expression gene set35 are available in the database of Genotypes and Phenotypes (dbGaP, https://www.ncbi.nlm.nih.gov/gap/, Study Accession: phs000417.v2.p1) and Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/, Study Accession: GSE30272). The weights assigned to each genotype of each SNP are available in Supplementary Table S1.
Competing interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. ODH is a part-time employee of H. Lundbeck A/S and has received investigator-initiated research funding from and/or participated in advisory/speaker meetings organised by Angelini, Autifony, Biogen, Boehringer-Ingelheim, Eli Lilly, Heptares, Global Medical Education, Invicro, Janssen, Lundbeck, Neurocrine, Otsuka, Sunovion, Rand, Recordati, Roche and Viatris/Mylan. Neither ODH nor his family have holdings/a financial stake in any pharmaceutical company. ODH has a patent for the use of dopaminergic imaging. JTRW is an investigator on a grant from Takeda Pharmaceuticals Ltd. to Cardiff University, for a project unrelated to the work presented here. The other authors have nothing to declare.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Alessandro Bertolino, Email: alessandro.bertolino@uniba.it.
Oliver D. Howes, Email: oliver.howes@kcl.ac.uk
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-022-16442-6.
References
- 1.Beaulieu J-M, Gainetdinov RR. The physiology, signaling, and pharmacology of dopamine receptors. Pharmacol. Rev. 2011;63:182–217. doi: 10.1124/pr.110.002642. [DOI] [PubMed] [Google Scholar]
- 2.Mishra, Singh, Shukla Physiological and functional basis of dopamine receptors and their role in neurogenesis: Possible implication for Parkinson’s disease. J. Exp. Neurosci. 2018 doi: 10.1177/1179069518779829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Pezze, Feldon Mesolimbic dopaminergic pathways in fear conditioning. Prog. Neurobiol. 2004;74:301–320. doi: 10.1016/j.pneurobio.2004.09.004. [DOI] [PubMed] [Google Scholar]
- 4.Seeman, Kapur Schizophrenia: More dopamine, more D2 receptors. Proc. Natl. Acad. Sci. 2000;97:7673–7675. doi: 10.1073/pnas.97.14.7673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Seeman P, Chau-Wong M, Tedesco J, Wong K. Brain receptors for antipsychotic drugs and dopamine: Direct binding assays. Proc. Natl. Acad. Sci. 1975;72:4376–4380. doi: 10.1073/pnas.72.11.4376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wang S, et al. Structure of the D2 dopamine receptor bound to the atypical antipsychotic drug risperidone. Nature. 2018;555:269–273. doi: 10.1038/nature25758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kaar SJ, Natesan S, McCutcheon R, Howes OD. Antipsychotics: Mechanisms underlying clinical response and side-effects and novel treatment approaches based on pathophysiology. Neuropharmacology. 2019;172:107704. doi: 10.1016/j.neuropharm.2019.107704. [DOI] [PubMed] [Google Scholar]
- 8.Pardiñas AF, et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 2018;50:381–389. doi: 10.1038/s41588-018-0059-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ripke S, et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–427. doi: 10.1038/nature13595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.McCutcheon RA, Krystal JH, Howes OD. Dopamine and glutamate in schizophrenia: Biology, symptoms and treatment. World Psychiatry. 2020;19:15–33. doi: 10.1002/wps.20693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bertolino A, et al. Genetically determined measures of striatal D2 signaling predict prefrontal activity during working memory performance. PLoS ONE. 2010;5:e9348. doi: 10.1371/journal.pone.0009348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Blasi G, et al. Variation in dopamine D2 and serotonin 5-HT2A receptor genes is associated with working memory processing and response to treatment with antipsychotics. Neuropsychopharmacology. 2015;40:1600–1608. doi: 10.1038/npp.2015.5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Di Giorgio A, et al. DRD2/CHRNA5 interaction on prefrontal biology and physiology during working memory. PLoS ONE. 2014;9:e95997. doi: 10.1371/journal.pone.0095997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gluskin BS, Mickey BJ. Genetic variation and dopamine D2 receptor availability: A systematic review and meta-analysis of human in vivo molecular imaging studies. Transl. Psychiatry. 2016;6:e747. doi: 10.1038/tp.2016.22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Luykx JJ, Broersen JL, de Leeuw M. The DRD2 rs1076560 polymorphism and schizophrenia-related intermediate phenotypes: A systematic review and meta-analysis. Neurosci. Biobehav. Rev. 2017;74:214–224. doi: 10.1016/j.neubiorev.2017.01.006. [DOI] [PubMed] [Google Scholar]
- 16.Sambataro F, et al. DRD2 genotype-based variation of default mode network activity and of its relationship with striatal DAT binding. Schizophr. Bull. 2013;39:206–216. doi: 10.1093/schbul/sbr128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wiers CE, et al. Association of genetic ancestry with striatal dopamine D2/D3 receptor availability. Mol. Psychiatry. 2018;23:1711–1716. doi: 10.1038/mp.2017.208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Boyle EA, Li YI, Pritchard JK. An expanded view of complex traits: From polygenic to omnigenic. Cell. 2017;169:1177–1186. doi: 10.1016/j.cell.2017.05.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chen J, Cao H, Meyer-Lindenberg A, Schwarz E. Male increase in brain gene expression variability is linked to genetic risk for schizophrenia. Transl. Psychiatry. 2018;8:140. doi: 10.1038/s41398-018-0200-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Gandal MJ, et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science. 2018;362:eaat8127. doi: 10.1126/science.aat8127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li M, et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science. 2018;362:eaat7615. doi: 10.1126/science.aat7615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hartl CL, et al. Coexpression network architecture reveals the brain-wide and multiregional basis of disease susceptibility. Nat. Neurosci. 2021;24:1313–1323. doi: 10.1038/s41593-021-00887-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.McGuffin P, Riley B, Plomin R. Toward behavioral genomics. Science. 2001;291:1232–1249. doi: 10.1126/science.1057264. [DOI] [PubMed] [Google Scholar]
- 24.Parikshak NN, Gandal MJ, Geschwind DH. Systems biology and gene networks in neurodevelopmental and neurodegenerative disorders. Nat. Rev. Genet. 2015;16:441–458. doi: 10.1038/nrg3934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pergola G, et al. A miR-137-related biological pathway of risk for Schizophrenia is associated with human brain emotion processing. bioRxiv. 2020 doi: 10.1101/2020.08.03.230227. [DOI] [PubMed] [Google Scholar]
- 26.Eisen MB, Spellman PT, Brown PO, Botstein D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. 1998;95:14863–14868. doi: 10.1073/pnas.95.25.14863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Fromer M, et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 2016;19:1442–1453. doi: 10.1038/nn.4399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Fazio L, et al. Transcriptomic context of DRD1 is associated with prefrontal activity and behavior during working memory. Proc. Natl. Acad. Sci. 2018;115:201717135. doi: 10.1073/pnas.1717135115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gaiteri, Ding, French, Tseng, Sibille Beyond modules and hubs: The potential of gene coexpression networks for investigating molecular mechanisms of complex brain disorders. Genes Brain Behav. 2014;13:13–24. doi: 10.1111/gbb.12106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pergola G, et al. Prefrontal co-expression of schizophrenia risk genes is associated with treatment response in patients. Biol. Psychiatry. 2019;86:45–55. doi: 10.1016/j.biopsych.2019.03.981. [DOI] [PubMed] [Google Scholar]
- 31.Radulescu E, et al. Identification and prioritization of gene sets associated with schizophrenia risk by co-expression network analysis in human brain. Mol. Psychiatry. 2020;25:791–804. doi: 10.1038/s41380-018-0304-1. [DOI] [PubMed] [Google Scholar]
- 32.Torretta S, et al. NURR1 and ERR1 modulate the expression of genes of a DRD2 coexpression network enriched for schizophrenia risk. J. Neurosci. 2020;40:932–941. doi: 10.1523/JNEUROSCI.0786-19.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.van Dam S, Võsa U, van der Graaf A, Franke L, de Magalhães JP. Gene co-expression analysis for functional classification and gene–disease predictions. Brief Bioinform. 2017 doi: 10.1093/bib/bbw139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Obayashi T, et al. COXPRESdb: A database of coexpressed gene networks in mammals. Nucleic Acids Res. 2008;36:D77–D82. doi: 10.1093/nar/gkm840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pergola G, et al. DRD2 co-expression network and a related polygenic index predict imaging, behavioral and clinical phenotypes linked to schizophrenia. Transl. Psychiatry. 2017;7:e1006. doi: 10.1038/tp.2016.253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Trubetskoy V, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022 doi: 10.1038/s41586-022-04434-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Callicott JH, et al. Physiological dysfunction of the dorsolateral prefrontal cortex in schizophrenia revisited. Cereb. Cortex. 2000;10:1078–1092. doi: 10.1093/cercor/10.11.1078. [DOI] [PubMed] [Google Scholar]
- 38.Braun U, et al. Brain network dynamics during working memory are modulated by dopamine and diminished in schizophrenia. Nat. Commun. 2021;12:3478. doi: 10.1038/s41467-021-23694-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Selvaggi P, et al. Genetic variation of a DRD2 co-expression network is associated with changes in prefrontal function after D2 receptors stimulation. Cereb. Cortex. 2019;29:1162–1173. doi: 10.1093/cercor/bhy022. [DOI] [PubMed] [Google Scholar]
- 40.Kessler RM, et al. Identification of extrastriatal dopamine D2 receptors in post mortem human brain with [125I]epidepride. Brain Res. 1993;609:237–243. doi: 10.1016/0006-8993(93)90878-Q. [DOI] [PubMed] [Google Scholar]
- 41.Nour MM, et al. Dopaminergic basis for signaling belief updates, but not surprise, and the link to paranoia. Proc. Natl. Acad. Sci. 2018;115:201809298. doi: 10.1073/pnas.1809298115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.McCutcheon RA, et al. Mesolimbic dopamine function is related to salience network connectivity: An integrative positron emission tomography and magnetic resonance study. Biol. Psychiatry. 2019;85:368–378. doi: 10.1016/j.biopsych.2018.09.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Laruelle M, et al. Single photon emission computerized tomography imaging of amphetamine-induced dopamine release in drug-free schizophrenic subjects. Proc. Natl. Acad. Sci. 1996;93:9235–9240. doi: 10.1073/pnas.93.17.9235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Abi-Dargham A, et al. Increased baseline occupancy of D2 receptors by dopamine in schizophrenia. Proc. Natl. Acad. Sci. 2000;97:8104–8109. doi: 10.1073/pnas.97.14.8104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Howes OD, et al. Midbrain dopamine function in schizophrenia and depression: A post-mortem and positron emission tomographic imaging study. Brain. 2013;136:3242–3251. doi: 10.1093/brain/awt264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Howes OD, et al. The nature of dopamine dysfunction in schizophrenia and what this means for treatment: Meta-analysis of imaging studies. Arch. Gen. Psychiatry. 2012;69:776–786. doi: 10.1001/archgenpsychiatry.2012.169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Jauhar S, et al. Determinants of treatment response in first-episode psychosis: An 18F-DOPA PET study. Mol. Psychiatry. 2019;24:1502–1512. doi: 10.1038/s41380-018-0042-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.McCutcheon R, Beck K, Jauhar S, Howes OD. Defining the locus of dopaminergic dysfunction in schizophrenia: A meta-analysis and test of the mesolimbic hypothesis. Schizophrenia Bull. 2017;44:1301–1311. doi: 10.1093/schbul/sbx180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Kumakura Y, et al. Elevated [18F]fluorodopamine turnover in brain of patients with schizophrenia: An [18F]fluorodopa/positron emission tomography study. J. Neurosci. 2007;27:8080–8087. doi: 10.1523/JNEUROSCI.0805-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Meyer-Lindenberg A, et al. Reduced prefrontal activity predicts exaggerated striatal dopaminergic function in schizophrenia. Nat. Neurosci. 2002;5:267–271. doi: 10.1038/nn804. [DOI] [PubMed] [Google Scholar]
- 51.Mizrahi R, et al. Increased stress-induced dopamine release in psychosis. Biol. Psychiatry. 2012;71:561–567. doi: 10.1016/j.biopsych.2011.10.009. [DOI] [PubMed] [Google Scholar]
- 52.Reith J, et al. Elevated dopa decarboxylase activity in living brain of patients with psychosis. Proc. Natl. Acad. Sci. 1994;91:11651–11654. doi: 10.1073/pnas.91.24.11651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Abi-Dargham A, van de Giessen E, Slifstein M, Kegeles LS, Laruelle M. Baseline and amphetamine-stimulated dopamine activity are related in drug-naïve schizophrenic subjects. Biol. Psychiatry. 2009;65:1091–1093. doi: 10.1016/j.biopsych.2008.12.007. [DOI] [PubMed] [Google Scholar]
- 54.Jauhar S, et al. A test of the transdiagnostic dopamine hypothesis of psychosis using positron emission tomographic imaging in bipolar affective disorder and schizophrenia. JAMA Psychiat. 2017;74:1206. doi: 10.1001/jamapsychiatry.2017.2943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Howes O, et al. Progressive increase in striatal dopamine synthesis capacity as patients develop psychosis: A PET study. Mol. Psychiatry. 2011;16:885–886. doi: 10.1038/mp.2011.20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Rogdaki M, et al. Striatal dopaminergic alterations in individuals with copy number variants at the 22q11.2 genetic locus and their implications for psychosis risk: A [18F]-DOPA PET study. Mol. Psychiatry. 2021 doi: 10.1038/s41380-021-01108-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.McCutcheon RA, Marques TR, Howes OD. Schizophrenia—An overview. JAMA Psychiat. 2020;77:201–210. doi: 10.1001/jamapsychiatry.2019.3360. [DOI] [PubMed] [Google Scholar]
- 58.Haber SN. Corticostriatal circuitry. Dialogues Clin. Neurosci. 2016;18:7–21. doi: 10.31887/DCNS.2016.18.1/shaber. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Slifstein M, et al. Deficits in prefrontal cortical and extrastriatal dopamine release in schizophrenia: A positron emission tomographic functional magnetic resonance imaging study. JAMA Psychiat. 2015;72:316–324. doi: 10.1001/jamapsychiatry.2014.2414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.D’Ambrosio E, et al. The effect of a genetic variant at the schizophrenia associated AS3MT/BORCS7 locus on striatal dopamine function: A PET imaging study. Psychiatry Res. Neuroimaging. 2019;291:34–41. doi: 10.1016/j.pscychresns.2019.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Bloomfield MAP, et al. Dopaminergic function in cannabis users and its relationship to cannabis-induced psychotic symptoms. Biol. Psychiatry. 2014;75:470–478. doi: 10.1016/j.biopsych.2013.05.027. [DOI] [PubMed] [Google Scholar]
- 62.Bloomfield MAP, Morgan CJA, Kapur S, Curran HV, Howes OD. The link between dopamine function and apathy in cannabis users: An [18F]-DOPA PET imaging study. Psychopharmacology. 2014;231:2251–2259. doi: 10.1007/s00213-014-3523-4. [DOI] [PubMed] [Google Scholar]
- 63.Froudist-Walsh S, et al. The effect of perinatal brain injury on dopaminergic function and hippocampal volume in adult life. Elife. 2017;6:e29088. doi: 10.7554/eLife.29088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Jauhar S, et al. Regulation of dopaminergic function: An [18F]-DOPA PET apomorphine challenge study in humans. Transl. Psychiatry. 2017;7:e1027–e1027. doi: 10.1038/tp.2016.270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Dahoun T, et al. The effect of the DISC1 Ser704Cys polymorphism on striatal dopamine synthesis capacity: An [18F]-DOPA PET study. Hum. Mol. Genet. 2018;27:3498–3506. doi: 10.1093/hmg/ddy242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Freeman B, et al. DNA from buccal swabs recruited by mail: Evaluation of storage effects on long-term stability and suitability for multiplex polymerase chain reaction genotyping. Behav. Genet. 2003;33:67–72. doi: 10.1023/A:1021055617738. [DOI] [PubMed] [Google Scholar]
- 67.Zhang B, Horvath S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. 2005 doi: 10.2202/1544-6115.1128. [DOI] [PubMed] [Google Scholar]
- 68.Colantuoni C, et al. Temporal dynamics and genetic control of transcription in the human prefrontal cortex. Nature. 2011;478:519–523. doi: 10.1038/nature10524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Anderson CA, et al. Data quality control in genetic case-control association studies. Nat. Protoc. 2010;5:1564–1573. doi: 10.1038/nprot.2010.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Chang CC, et al. Second-generation PLINK: Rising to the challenge of larger and richer datasets. Gigascience. 2015;4:1–16. doi: 10.1186/s13742-015-0047-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Conomos MP, Miller MB, Thornton TA. Robust inference of population structure for ancestry prediction and correction of stratification in the presence of relatedness. Genet. Epidemiol. 2015;39:276–293. doi: 10.1002/gepi.21896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Gogarten SM, et al. Genetic association testing using the GENESIS R/bioconductor package. Bioinformatics. 2019;35:5346–5348. doi: 10.1093/bioinformatics/btz567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Kumakura Y, Cumming P. PET studies of cerebral levodopa metabolism: A review of clinical findings and modeling approaches. Neuroscientist. 2009;15:635–650. doi: 10.1177/1073858409338217. [DOI] [PubMed] [Google Scholar]
- 74.Veronese M, et al. A potential biomarker for treatment stratification in psychosis: Evaluation of an [18F] FDOPA PET imaging approach. Neuropsychopharmacology. 2020 doi: 10.1038/s41386-020-00866-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Turkheimer FE, Brett M, Visvikis D, Cunningham VJ. Multiresolution analysis of emission tomography images in the wavelet domain. J. Cereb. Blood Flow Metab. 1999;19:1189–1208. doi: 10.1097/00004647-199911000-00003. [DOI] [PubMed] [Google Scholar]
- 76.Egerton A, Demjaha A, McGuire P, Mehta MA, Howes OD. The test–retest reliability of 18F-DOPA PET in assessing striatal and extrastriatal presynaptic dopaminergic function. Neuroimage. 2010;50:524–531. doi: 10.1016/j.neuroimage.2009.12.058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Martinez D, et al. Imaging human mesolimbic dopamine transmission with positron emission tomography. Part II: Amphetamine-induced dopamine release in the functional subdivisions of the striatum. J. Cereb. Blood Flow Metab. 2003;23:285–300. doi: 10.1097/01.WCB.0000048520.34839.1A. [DOI] [PubMed] [Google Scholar]
- 78.Howes O, et al. Elevated striatal dopamine function linked to prodromal signs of schizophrenia. Arch. Gen. Psychiatry. 2009;66:13–20. doi: 10.1001/archgenpsychiatry.2008.514. [DOI] [PubMed] [Google Scholar]
- 79.Team, R. C. R: A Language and Environment for Statistical Computing (2021).
- 80.Kumakura Y, et al. Age-dependent decline of steady state dopamine storage capacity of human brain: An FDOPA PET study. Neurobiol. Aging. 2010;31:447–463. doi: 10.1016/j.neurobiolaging.2008.05.005. [DOI] [PubMed] [Google Scholar]
- 81.Egerton A, et al. Elevated striatal dopamine function in immigrants and their children: A risk mechanism for psychosis. Schizophrenia Bull. 2017;43:293–301. doi: 10.1093/schbul/sbx023.081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Collister JA, Liu X, Clifton L. calculating polygenic risk scores (PRS) in UK Biobank: A practical guide for epidemiologists. Front. Genet. 2022;13:818574. doi: 10.3389/fgene.2022.818574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Wickham H. ggplot2, Elegant Graphics for Data Analysis. Springer; 2016. pp. 241–253. [Google Scholar]
- 84.Millard SP. EnvStats: An R Package for Environmental Statistics. Springer; 2013. [Google Scholar]
- 85.Pergola G, et al. Combined effect of genetic variants in the GluN2B coding gene (GRIN2B) on prefrontal function during working memory performance. Psychol. Med. 2016;46:1135–1150. doi: 10.1017/S0033291715002639. [DOI] [PubMed] [Google Scholar]
- 86.Demjaha A, Murray R, McGuire P, Kapur S, Howes O. Dopamine synthesis capacity in patients with treatment-resistant schizophrenia. AJP. 2012;169:1203–1210. doi: 10.1176/appi.ajp.2012.12010144. [DOI] [PubMed] [Google Scholar]
- 87.Jauhar S, et al. The effects of antipsychotic treatment on presynaptic dopamine synthesis capacity in first-episode psychosis: A positron emission tomography study. Biol. Psychiatry. 2019;85:79–87. doi: 10.1016/j.biopsych.2018.07.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.McCutcheon R, et al. Antipsychotic plasma levels in the assessment of poor treatment response in schizophrenia. Acta Psychiatr. Scand. 2018;137:39–46. doi: 10.1111/acps.12825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Cools R, Gibbs SE, Miyakawa A, Jagust W, D’Esposito M. Working memory capacity predicts dopamine synthesis capacity in the human striatum. J. Neurosci. 2008;28:1208–1212. doi: 10.1523/JNEUROSCI.4475-07.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Landau SM, Lal R, O’Neil JP, Baker S, Jagust WJ. Striatal dopamine and working memory. Cereb. Cortex. 2009;19:445–454. doi: 10.1093/cercor/bhn095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Vernaleken I, et al. ‘Prefrontal’ cognitive performance of healthy subjects positively correlates with cerebral FDOPA influx: An exploratory [18F]-fluoro-L-DOPA-PET investigation. Hum. Brain Mapp. 2007;28:931–939. doi: 10.1002/hbm.20325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Braskie MN, et al. Correlations of striatal dopamine synthesis with default network deactivations during working memory in younger adults. Hum. Brain Mapp. 2011;32:947–961. doi: 10.1002/hbm.21081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Braskie MN, et al. Relationship of striatal dopamine synthesis capacity to age and cognition. J. Neurosci. 2008;28:14320–14328. doi: 10.1523/JNEUROSCI.3729-08.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Ma C, Gu C, Huo Y, Li X, Luo X-J. The integrated landscape of causal genes and pathways in schizophrenia. Transl. Psychiatry. 2018;8:67. doi: 10.1038/s41398-018-0114-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Whitton L, et al. Cognitive analysis of schizophrenia risk genes that function as epigenetic regulators of gene expression. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2016;171:1170–1179. doi: 10.1002/ajmg.b.32503. [DOI] [PubMed] [Google Scholar]
- 96.Li J, et al. Identification of human neuronal protein complexes reveals biochemical activities and convergent mechanisms of action in autism spectrum disorders. Cell Syst. 2015;1:361–374. doi: 10.1016/j.cels.2015.11.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Wang Z, et al. Knockdown of GATAD2A suppresses cell proliferation in thyroid cancer in vitro. Oncol. Rep. 2017;37:2147–2152. doi: 10.3892/or.2017.5436. [DOI] [PubMed] [Google Scholar]
- 98.Fullard JF, et al. Open chromatin profiling of human postmortem brain infers functional roles for non-coding schizophrenia loci. Hum. Mol. Genet. 2017;26:ddx103. doi: 10.1093/hmg/ddx103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Bipolar Disorder and Schizophrenia Working Group of the Psychiatric Genomics Consortium et al. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell. 2018;173:1705–1715. doi: 10.1016/j.cell.2018.05.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Wallén Å, et al. Orphan nuclear receptor Nurr1 is essential for ret expression in midbrain dopamine neurons and in the brain stem. Mol. Cell. Neurosci. 2001;18:649–663. doi: 10.1006/mcne.2001.1057. [DOI] [PubMed] [Google Scholar]
- 101.Tseng KY, et al. Selective increase of Nurr1 mRNA expression in mesencephalic dopaminergic neurons of D2 dopamine receptor-deficient mice. Mol. Brain Res. 2000;80:1–6. doi: 10.1016/S0169-328X(00)00107-8. [DOI] [PubMed] [Google Scholar]
- 102.Kim S, et al. The Dopamine D<sub>2</sub> receptor regulates the development of dopaminergic neurons via extracellular signal-regulated kinase and Nurr1 activation. J. Neurosci. 2006;26:4567. doi: 10.1523/JNEUROSCI.5236-05.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Anzalone A, et al. Dual control of dopamine synthesis and release by presynaptic and postsynaptic dopamine D2 receptors. J. Neurosci. 2012;32:9023–9034. doi: 10.1523/JNEUROSCI.0918-12.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Paladini CA, Robinson S, Morikawa H, Williams JT, Palmiter RD. Dopamine controls the firing pattern of dopamine neurons via a network feedback mechanism. Proc. Natl. Acad. Sci. 2003;100:2866–2871. doi: 10.1073/pnas.0138018100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Kehr W, Carlsson A, Lindqvist M, Magnusson T, Atack C. Evidence for a receptor-mediated feedback control of striatal tyrosine hydroxylase activity. J. Pharm. Pharmacol. 1972;24:744–747. doi: 10.1111/j.2042-7158.1972.tb09104.x. [DOI] [PubMed] [Google Scholar]
- 106.Wolf ME, Roth RH. Autoreceptor regulation of dopamine synthesis. Ann. N.Y. Acad. Sci. 1990;604:323–343. doi: 10.1111/j.1749-6632.1990.tb32003.x. [DOI] [PubMed] [Google Scholar]
- 107.Bello EP, et al. Cocaine supersensitivity and enhanced motivation for reward in mice lacking dopamine D2 autoreceptors. Nat. Neurosci. 2011;14:1033–1038. doi: 10.1038/nn.2862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Marinelli M, Cooper DC, Baker LK, White FJ. Impulse activity of midbrain dopamine neurons modulates drug-seeking behavior. Psychopharmacology. 2003;168:84–98. doi: 10.1007/s00213-003-1491-1. [DOI] [PubMed] [Google Scholar]
- 109.Nordio G, et al. Digital data repository and automatic analysis framework for FDOPA PET neuroimaging. Biorxiv. 2022 doi: 10.1101/2022.04.14.488129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Cropley VL, et al. Pre- and post-synaptic dopamine imaging and its relation with frontostriatal cognitive function in Parkinson disease: PET studies with [11C]NNC 112 and [18F]FDOPA. Psychiatry Res. Neuroimaging. 2008;163:171–182. doi: 10.1016/j.pscychresns.2007.11.003. [DOI] [PubMed] [Google Scholar]
- 111.Kilbourn MR. 11C- and 18F-radiotracers for in vivo imaging of the dopamine system: Past: past, present and future. Biomedicines. 2021;9:108. doi: 10.3390/biomedicines9020108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Frankle WG, Himes M, Mason NS, Mathis CA, Narendran R. Prefrontal and striatal dopamine release are inversely correlated in schizophrenia. Biol. Psychiatry. 2022 doi: 10.1016/j.biopsych.2022.05.009. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The PET data are available in The NeurOimaging DatabasE (NODE) repository (https://maudsleybrc.nihr.ac.uk/research/precision-psychiatry/neuroimaging/neuroimaging-database-node/) upon request. The data68 used for the WGCNA performed to identify the DRD2 co-expression gene set35 are available in the database of Genotypes and Phenotypes (dbGaP, https://www.ncbi.nlm.nih.gov/gap/, Study Accession: phs000417.v2.p1) and Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/, Study Accession: GSE30272). The weights assigned to each genotype of each SNP are available in Supplementary Table S1.


