Figure 8.
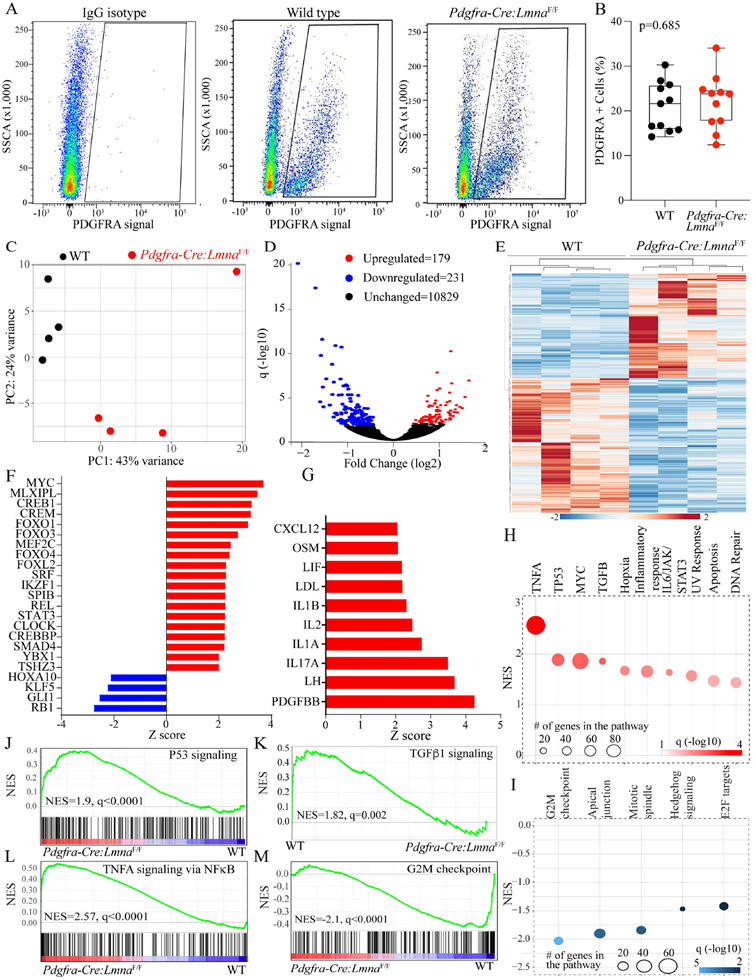
Differentially expressed genes, dysregulated transcriptional regulators, and biological pathways in the LMNA-deficient cardiac fibroblasts. (A) Fluorescence-activated cell sorting (FACS) panels showing the gating for IgG isotype and isolation of the cells expressing PDGFRA in the WT and Pdgfra-Cre:LmnaF/F mice. (B) Quantitative data showing the percentage of the non-myocyte cells isolated based on the signaling from an antibody against PDGFRA in the 6-week-old WT and Pdgfra-Cre:LmnaF/F mice. (C) Principle component analysis (PCA) of the cardiac fibroblast transcripts shows distinct separation between the WT and Pdgfra-Cre:LmnaF/F genotypes. (D) Volcano plot showing the down-regulated (blue), upregulated (red), and unchanged (black) genes in the Pdgfra-Cre:LmnaF/F as compared to the WT fibroblasts. (E) Heat map of the differentially expressed genes (DEGs) between the WT and Pdgfra-Cre:LmnaF/F fibroblasts. (F) The bar chart represents the predicted transcriptional regulators of the DEGs in Pdgfra-Cre:LmnaF/F fibroblasts. Red indicated predicted activation and blue predicted suppression. (G) The bar chart represents predicting growth and mitotic factors based on the DEGs in the Pdgfra-Cre:LmnaF/F fibroblasts. (H) Biological pathways predicted to be activated in the Pdgfra-Cre:LmnaF/F fibroblasts. (I) Biological pathways predicted to be suppressed in the Pdgfra-Cre:LmnaF/F fibroblasts. (J-M) Plots representing gene set enrichment analysis (GSEA) predicting activation of the TP53, TNFA/NFκB, and TGFβ1 and suppression of the cell cycle G2M pathways.