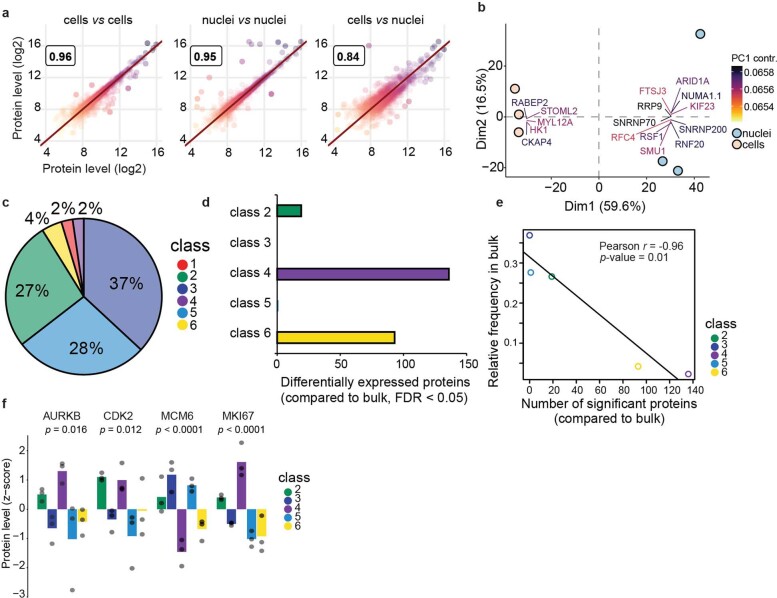
Extended Data Fig. 2. PCA and loadings of cell culture classes at sub-cellular level and number of significantly changed proteins vs. class abundance.
a, Quantitative proteomic results of whole cell and nuclei replicates, and comparison between whole cells and nuclei. b, Principal component analysis (PCA) of whole cell (n = 3) and nuclei proteomes (n = 3). Proteins with the strongest contribution to PC1 are highlighted. c, Relative proportions of the six nuclei classes. d, Number of differentially expressed proteins (two-sided t-test, n = 3 biological replicates) compared to unclassified nuclei (bulk). Proteins with an FDR less than 0.05 were considered significant. e, Correlation between number of significantly regulated proteins per nuclei class vs relative class proportion. A linear model was fitted to the data showing an inverse correlation with Pearson r = -0.96 (p-value = 0.01). f, Relative protein levels (z-score) of known cell cycle markers across the five nuclei classes. All bar graphs represent mean of data (n = 3 biological replicates) and error bars are s.d. ANOVA p-values are shown.