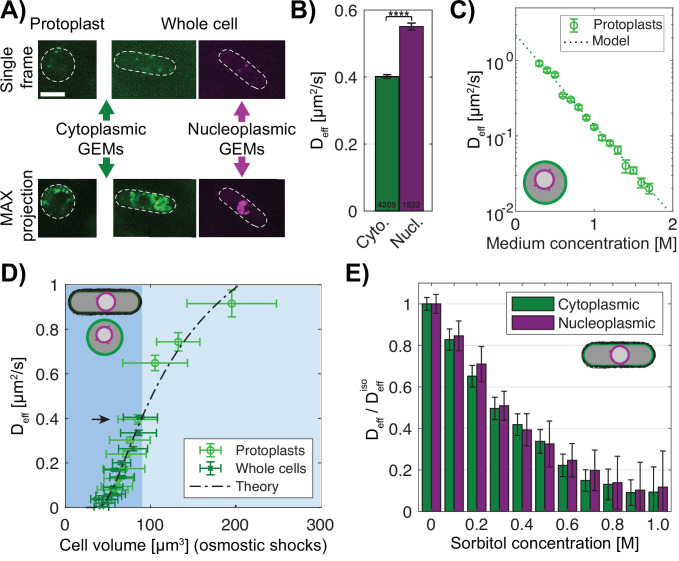
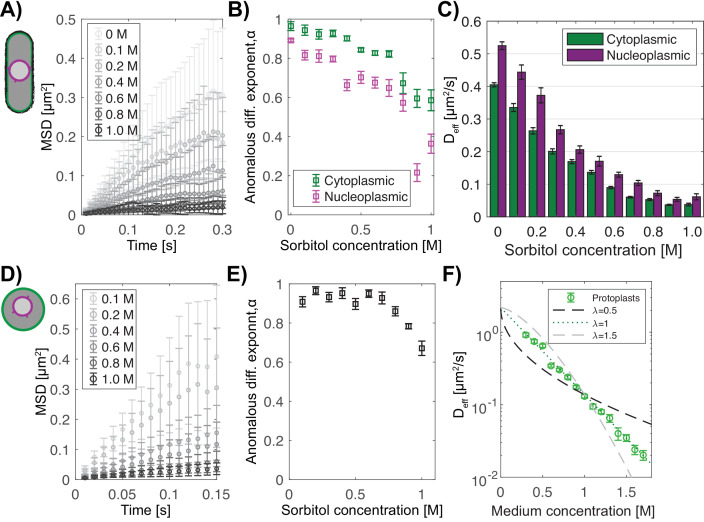
Figure 4. Macromolecular crowding is affected similarly in the nucleus and cytoplasm under osmotic shocks.
(A) Images of protoplasts (left) and whole cells (right) expressing cytoplasmic 40 nm GEMs and nucleoplasmic 40 nm GEMs. Top, single time point image; bottom, maximum projection of 100 frames. Dashed lines, the cell boundary. Scale bar = 5 µm. (B) GEMs effective diffusion coefficient (mean ± SEM) is slower in the cytoplasm (green) than in the nucleoplasm (purple) in whole cells in YE medium. Numbers indicate the number of tracks, p-value <0.0001 Mann-Whitney U test. (C) Effective diffusion coefficient of cytoplasmic GEMs (mean ± SEM) in protoplasts shifted to various sorbitol concentrations in the medium. Dashed lines, predictions of Phillies’ model for diffusion with a power law λ=1. NGEMs tracked = 4058, from at least two biological replicates per condition. (D) Effective diffusion coefficient of cytoplasmic GEMs (mean ± SEM) plotted against cell volume under hypotonic and hypertonic shock (light blue and blue background respectively). Volumes represent mean distribution of an asynchronous culture ± STD. Dashed line, fit of Phillies’ model for self-diffusing trackers in a polymer solution. Black arrow indicates Deff for a population of cells in YE and protoplasts in isotonic condition. Protoplasts: NGEMs = 3355, NVolume = 2216 cells, whole cells: NGEMs = 9849, NVolume = 981 cells. (E) Effects of hyperosmotic shifts on the relative effective diffusion coefficients (mean ± SEM) of cytoplasmic and nuclear GEMs; no statistically significant difference was detected (F-test, p-value = 0.90). Cytoplasm NGEMs = 9365,, nuclear NGEMs = 3732, from at least two biological replicates per condition. See also Figure 4—figure supplement 1.