Fig. 3.
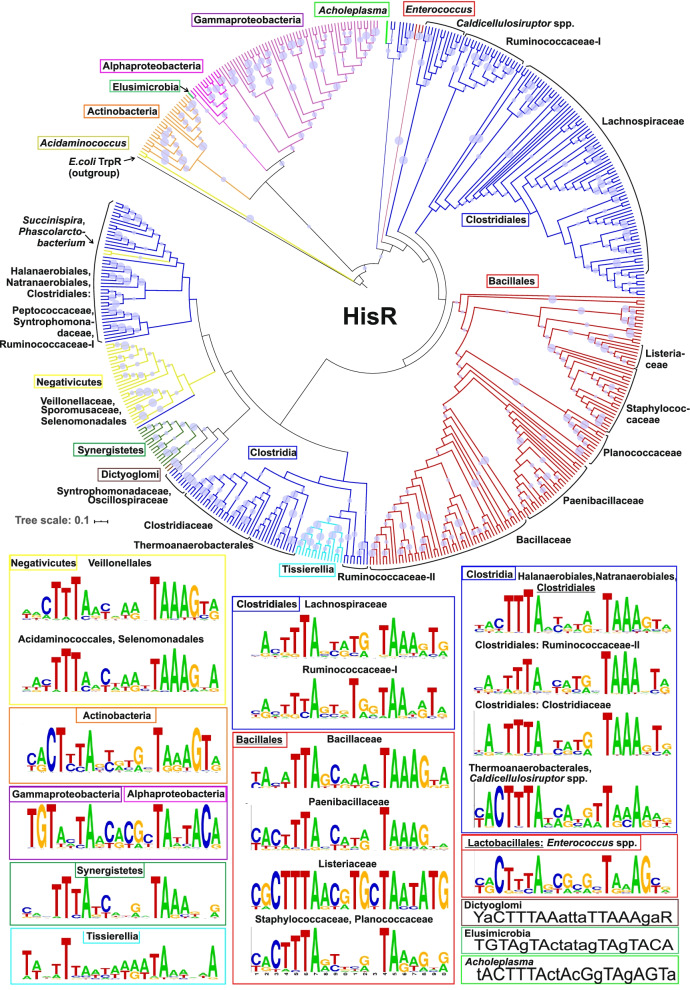
DNA binding motifs and phylogeny of HisR regulators in various taxonomic groups of Bacteria. Sequence logos representing the consensus HisR binding site motifs were constructed using all candidate sites in the respective bacterial lineage. Maximum likelihood phylogenetic tree for HisR proteins in studied genomes of Firmicutes (395 genomes), Proteobacteria (62 genomes), Actinobacteria (22 genomes), Synergistetes (9 genomes), Dictyoglomi (2 genomes), Elusimicrobia and Tenericutes. Branches representing HisR proteins from each of the above phyla are highlighted by colors matching their corresponding HisR motif box outline colors. The size of circles in internal nodes represents bootstrap values in percentages (out of 100). TrpR repressor from E. coli was used as an outgroup. Detailed tree with protein IDs and genome names is presented in the rectangular form in Additional file 1