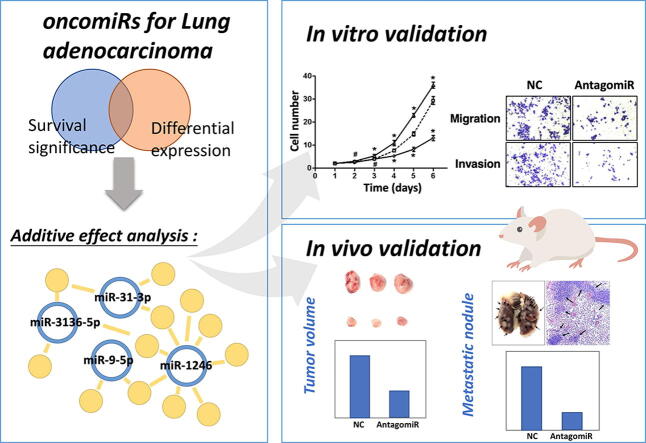
Graphical abstract
Keywords: oncomiR, antagomiR, Lung adenocarcinoma, miRNA treatment
Abstract
Lung cancer is a major cause of cancer-associated deaths worldwide, and lung adenocarcinoma (LUAD) is the most common lung cancer subtype. Micro RNAs (miRNAs) regulate the pattern of gene expression in multiple cancer types and have been explored as potential drug development targets. To develop an oncomiR-based panel, we identified miRNA candidates that show differential expression patterns and are relevant to the worse 5-year overall survival outcomes in LUAD patient samples. We further evaluated various combinations of miRNA candidates for association with 5-year overall survival and identified a four-miRNA panel: miR-9-5p, miR-1246, miR-31-3p, and miR-3136-5p. The combination of these four miRNAs outperformed any single miRNA for predicting 5-year overall survival (hazard ratio [HR]: 3.47, log-rank p-value = 0.000271). Experiments were performed on lung cancer cell lines and animal models to validate the effects of these miRNAs. The results showed that singly transfected antagomiRs largely inhibited cell growth, migration, and invasion, and the combination of all four antagomiRs considerably reduced cell numbers, which is twice as effective as any single miRNA-targeted transfected. The in vivo studies revealed that antagomiR-mediated knockdown of all four miRNAs significantly reduced tumor growth and metastatic ability of lung cancer cells compared to the negative control group. The success of these in vivo and in vitro experiments suggested that these four identified oncomiRs may have therapeutic potential.
1. Introduction
Lung cancer is a major cause of cancer-associated deaths in a number of countries, and lung adenocarcinoma (LUAD) is the leading lung cancer subtype [1]. Multiple environmental and risk factors have been associated with the incidence and mortality of lung cancer. According to the update of GLOBOCAN (The Global Cancer Observatory) in 2020, lung cancer remained the leading cause of cancer death, resulting in more than 18 % of global cancer-related deaths [2]. Currently available treatments include various therapeutic targets, such as epithelial growth factor receptor (EGFR) and anaplastic lymphoma kinase (ALK), or immune therapy [3]; however, therapeutic outcomes remain far from ideal.
Micro RNAs (miRNAs) are small, endogenous RNAs that regulate the pattern of gene expression in multiple cancer types and in various tumor cell growth levels, including migration, invasion, and metastasis [4]. With these known crucial roles, miRNAs have been explored as potential tools for cancer diagnosis and prognosis [5]. MiRNAs can be characterized as either tumor suppressor miRNAs or oncogenic miRNAs (oncomiRs), depending on their functions [6]. Developed miRNA mimics and miRNA inhibitors, also known as antagomiRs, have achieved encouraging results in preclinical and clinical trials [7]. For example, antagomiRs against miR-155 have been designed as anti-miRNA-based treatments for cutaneous T cell lymphoma (CTCL) in Phase I clinical trials [8]. In addition to miR-155, many miRNA-related therapies have entered clinical trials [9]. Although multiple tumor suppressor miRNAs have been explored for therapeutic development, treatments that target the dysregulation of oncomiRs have been found they also play significant roles in cancer studies. For example, miR-21 had been identified to play a pro-tumorigenic role in non-small cell lung cancer (NSCLC) [7]. Another study also indicated that the amplification of the miR-21 locus could serve as a prognostic marker, based on analyses of LUAD data in The Cancer Genome Atlas (TCGA) [10]. One study examined a miRNA-based diagnostic assay and determined that evaluating the expression levels of eight miRNAs could effectively distinguish between lung cancer subtypes [11]. In the field of lung cancer research, the miRNAs let-7, miR-34, miR-150, and miR-200c have been well-studied for therapeutic effects [12]. Although a considerable amount of research has been devoted to the development of miRNA-based therapeutic strategies, a limited number of studies have progressed to preclinical or clinical applications. The determination of the best miRNA candidates to target specific diseases and cancer types during the therapeutic development of miRNAs has been a major challenge [7].
In this study, an integrative bioinformatics method was applied to identify multiple oncomiRs candidates, and experiments were performed in cell lines and animal models to validate the effects by using antagomiRs of the miRNA candidates in inhibiting the tumor growth.
2. Materials and methods
2.1. Data collection and preparation
The processed datasets, RNA sequencing (RNA-seq) profiles and smRNA sequencing (smRNA-seq) profiles, were collected from two databases: DriverDB database (greater than9500 cancer-related samples) [13], [14], [15] and YM500 (8105 smRNA samples) [16], [17], [18] database. Briefly, in DriverDB, RNA-seq data were retrieved from the TCGA data portal (https://portal.gdc.cancer.gov/) (such as primary tumor, normal and metastatic tissues) and annotated [14]. In YM500, smRNA-seq data were downloaded from CGHub (https://cghub.ucsc.edu/), processed by the miR-seq pipeline of YM500 [16], [17], [18], and annotated with miRBase database R21 [19] and DASHR database v1.0 [20]. A total of 20,495 genes and 2588 miRNAs were curated in this study. Because not every patient in the TCGA-LUAD cohort conducted both RNA-seq and miRNA-seq, RNA-seq data from 59 normal and 515 LUAD tumor samples and miRNA-seq data from 46 normal and 513 LUAD tumor samples were included in the following analyses. For RNA-seq, we used FPKM-UQ, whereas the miRNAseq was normalized by RPM.
2.2. Identification of differentially expressed miRNAs
To define significant differentially expressed miRNAs between primary tumor samples and adjacent normal samples, we applied an R package, DEseq (Version 1.28.0) [21]. The criteria used to filter candidates included adjusted p-values < 0.05 and log2 fold-change values greater than 3 for miRNAs, which efficiently identified oncogenic expression between tumor samples and normal samples. To eliminate candidates with excessively low expression levels, we further filtered out candidates with normalized mean counts (baseMean) < 1 for miRNAs.
2.3. Survival analysis of miRNA
To define clinically relevant candidates, a R package, Survival (Version 2.41–3) [22], was applied to calculate the Cox proportional hazards model between two predefined groups, which provides survival estimates for time-to-event datasets. LUAD cancer patients were stratified according to median miRNA expression levels. The number of patients included in the clinical TCGA dataset for survival analysis was 503. A total of 114 deaths occurred over a five-year period. Overall, 2588 miRNAs and 20,495 genes were included in the analysis. miRNA with survival significance was defined as those with log-rank p-values < 0.05 and hazard ratio (HR) values greater than 1 for miRNAs. Five-year overall survival was used for the following analyses.
2.4. Additive effect on survival for miRNAs
To further reduce the number of miRNA candidates, an analytic model was established to evaluate the additive effects on survival for co-expressed miRNAs that derived from the above analysis. First, a survival estimate was computed for all 31 combinations of miRNA candidates in no particular order. For individual miRNA, we used a median cut-off to split patients into “high” and “low” groups, respectively. Consequently, patients may be assigned to the “high” group by one miRNA and the “low” group by another. In the combination of miRNAs, the patient must be identified to be “high” by all miRNAs and then be assigned into the “all high” group and, on the flip side, into the “all low” group and vice versa. Significant combinations were defined as those with log-rank p-values < 0.05 and HR values greater than 1.
2.5. Identification of miRNA-target genes
We used four criteria to identify miRNA-target candidates: (1) Genes with differentially expressed between normal and tumor. DEseq (Version 1.28.0) [21] was used to explore the differential expressed miRNA-target genes between tumor samples and normal samples which were filtered by adjusted p-values < 0.05 and log2 fold-change values <− 1. Low expression candidates were also filtered out with < 10 normalized mean counts (baseMean) for genes. (2) Survival significant genes. These candidates were further analyzed with the five-year survival by Cox proportional hazards model. Log-rank p-values < 0.05 and HR < 1 were defined as survival significance genes. (3) Genes with expression correlation to the miRNAs. To confirm the relationships between miRNAs and target genes, we performed the following analytical steps. Firstly, we calculated correlations between gene and miRNA expression levels, using Pearson’s [18], Spearman’s [23], and Kendall’s coefficients [18]. This method comprises both linear and non-linear correlation, increasing the sensitivity of detecting potential target genes of miRNA candidates. Those genes and miRNAs for which any one of the three correlation coefficients was <− 0.3 were considered. (4) Twelve additional bioinformatic prediction tools, which predicted the interaction between miRNA and corresponding target genes, were applied to investigate the interactions between miRNAs and their target genes, as detailed in a previous publication [18]. Targeting gene with only negative correlation with a miRNA but without a bioinformatic tool supporting the interaction was regarded as a “weak linkage”. Instead, if an interaction was supported by more than one or more than 6 of 12 bioinformatic tools, it was regarded as “medium” or “strong”, respectively.
2.6. Functional enrichment analysis
Functional enrichment analysis of the target genes was performed as described in our previous publications [13], [14]. Reactome pathway annotations were used with adjusted p-values < 0.05. The top 10 significant Reactome, Gene Ontology Biological Process (GO-BP) and Kyoto Encyclopedia of Genes and Genomes (KEGG) terms were extracted from the functional annotation outcomes. Moreover, a gene set overrepresentation analysis was performed using another R package, GSOAP [24], which investigates intersecting gene sets across multiple functions.
2.7. Cell culture and transfection
All cell lines were maintained in a medium (Supplementary Table 1) containing 10 % fetal bovine serum (FBS) and 1 % penicillin/streptomycin (PS; Gibco, Thermo Fisher Scientific, Waltham, MA, USA) at 37 °C with 5 % CO2. The functional assays were performed to validate the biological role of these oncomiRs using three lung cancer cell lines, including A549, Hop62, and a brain metastatic LUAD cell line Bm7 [25] cultured in Dulbecco’s modified Eagle medium (DMEM)/F12 medium. The A549, Bm7, and Hop62 cells were transfected with antagomiRs targeting miR-9-5p, miR-31-3p, miR-1246, and miR-3136-5p or the anti-miR miRNA Inhibitor Negative Control (NC; Ambion, Thermo Fisher Scientific) at a final concentration of 20 nM using Lipofectamine 2000 (Invitrogen) for 48 h, according to the manufacturer’s instructions. The anti-miR miRNA Inhibitor Negative Control is a random sequence anti-miR molecule with no identifiable effects on known miRNA functions in human cell lines and tissues.
2.8. Quantitative real-time PCR
Total RNA was extracted using TRIzol reagent (Invitrogen, Thermo Fisher Scientific) according to the manufacturer’s instructions, and then reverse-transcribed into complementary DNA using the MultiScribe Reverse Transcriptase system and specific primers designed for miR-9-5p (#000583), miR-31-3p (#002113), miR-1246 (#462575), or miR-3136-5p (#243337; Applied Biosystems, Thermo Fisher Scientific). The miRNA expression levels were determined with the ABI StepOne Plus system through quantitative real-time PCR using TaqMan Universal PCR Master Mix and TaqMan MicroRNA Assays (Applied Biosystems, Thermo Fisher Scientific). The relative miRNA levels were normalized against the expression of RNU48 small nuclear RNA, which is a stably expressed and well-validated gene used as a normalizer [26].
Fold change = 2^ - [(CT target - CT RNU48) test sample - (CT target - CT RNU48) control sample].
For miRNA target gene expression analysis, the cDNA was amplified with primers (Supplementary Table 2). The relative mRNA levels were normalized to the expression of 18S rRNA.
Fold change = 2^ - [(CT target - CT 18S rRNA) test sample - (CT target - CT 18S rRNA) control sample].
2.9. Western blot analysis
As previously described [27], the tumor cell lysates were prepared and analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis using antibodies against AK1, CACNA2D2, ESYT3, NFIX, SELENBP1, SNX30 (Novus Biologicals, Centennial, CO, USA), and β-actin (Genetex, Alton Pkwy Irvine, CA, USA).
2.10. Cell growth and viability assays
For the measurement of cell growth, transfected cells (3 × 105) were seeded into 6-well plates and analyzed by the trypan blue exclusion method at the indicated times, as described previously [27]. For the assessment of cell viability, transfected cells (1 × 104) were seeded into 24-well plates. After incubation for 24, 48, or 72 h, 50 µL of 3-(4, 5-dimethyl-2-thiazolyl)-2, 5-diphenyl-2H-tetrazolium bromide (MTT; Sigma-Aldrich) solution was added to the medium for an additional 3 h. The purple formazan precipitates were dissolved in 250 µL dimethyl sulfoxide, and absorbance was determined using a microplate enzyme-linked immunosorbent assay (ELISA) reader (TECAN Infinite 200) [27].
2.11. Soft agar colony formation assay
As described previously, for the evaluation of anchorage-independent cell growth, transfected cells were plated at 5,000 cells/well on soft agar [27], [28]. After 14 days, the cells were stained with 0.005 % crystal violet, and colonies larger than 100 µm in diameter were counted in 10 random fields under a light microscope (40 × magnification times).
2.12. Migration and invasion assays
Migration and invasion assays (3 × 104 cells for the migration assay and 5 × 104 cells for the invasion assay) were performed in 24-well plates. Cells were seeded onto Millicell tissue culture plate well inserts with 8-μm pores (Millipore, Bedford, MA) for 12 h and BD BioCoat Matrigel Invasion Chambers (Becton Dickson, Mountain View, CA) for 20 h, respectively, as described previously [27], [29]. The migrating or invading cells were fixed with methanol for 10 min and stained with crystal violet for 1 h. The total number of stained cells was counted under a light microscope.
2.13. Xenograft tumorigenicity assay in nude mice
All animal experiments were performed with the approval of the Institutional Animal Care and Use Committee (IACUC) of China Medical University. Five-week-old male mice (BALB/c nu/nu; National Science Council Animal Center, Taipei, Taiwan) were subcutaneously injected with control (NC) or four antagomiRs-transfected Bm7 cells (3 × 106 cells) into both flanks near hind limbs (n = 8 per group). The mice were sacrificed on day 30 after implantation, and the tumor volumes were measured. The expression levels of miR-9-5p, miR-31-3p, miR-1246, and miR-3136-5p in the xenografted tumors were determined by miRNA quantitative real-time PCR, as described previously [27], [30].
2.14. In vivo tail vein metastasis assay
As described previously [27], [30], controls (NC) or four antagomiRs-transfected Bm7 cells (1 × 106) were intravenously injected into the tail veins of six-week-old male non-obese diabetic severe-combined immunodeficiency mice (NOD-SCID mice; n = 6 per group). After 15 weeks, the mice were sacrificed, and metastatic lung nodules were counted by gross and microscopic examination. The expression levels of miR-9-5p, miR-31-3p, miR-1246, and miR-3136-5p in excised lung tumor samples were detected by miRNA quantitative real-time PCR.
2.15. Immunohistochemical staining
Immunohistochemical staining was performed as described previously [31]. Briefly, tissue sections were incubated with primary antibodies against Ki-67 (ab15580, abcam, Cambridge, MA, United Kingdom) at 1:2000 dilution. The stained cells were considered as Ki-67-positive.
3. Statistical analysis for in vivo and in vitro experiments
Unless otherwise noted, each sample was assayed in triplicate. Statistical analyses were performed using the Mann Whitney U test for a simple comparison between two groups. A p-value<0.05 was considered significant.
4. Results
4.1. Identification of oncomiR candidates by integrative bioinformatics approaches
To identify survival-relevant miRNA candidates that might play oncogenic roles in LUAD, we incorporated several approaches, as shown in Fig. 1A. Firstly, a differential expression analysis between tumor and normal samples was performed to identify 37 abnormally expressed miRNAs (Supplementary Table 3). Among those miRNAs, we identified five miRNAs (Supplementary Table 4) associated with clinical outcomes related to 5-year overall survival. Finally, we conducted an additive effect analysis to explore the possibility of reducing candidate number while keeping or scarifying a tolerable statistical power as a tradeoff. As shown in Fig. 1B, the highest 5-year overall survival HR of a 4-gene combination signature was comparable to the HR of the 5-gene combination signature (HR4-gene = 3.47 vs HR5-gene = 3.60). As a consequence, we choose the combination of miR-1246, miR-9-5p, miR-31-3p, and miR-3136-5p as the final signature with the highest HR among all examined combinations of 4 miRNAs. Fig. 1C demonstrates that all four miRNAs in this combination were upregulated in tumor samples compared with normal samples, and Supplementary Fig. 1A shows a significant difference in survival when the population was stratified according to the expression levels of these four miRNAs. The combined evaluation of all four miRNAs in this panel (Supplementary Table 5) showed a prognostic signature for 5-year overall survival, as shown in Fig. 1D (log-rank p-value < 0.001) and Supplementary Fig. 1B. The biological functions of these four oncomiRs were further investigated to determine whether they were oncogenic relevant.
Fig. 1.
Integrative bioinformatics analysis for the identification of oncomiRs in LUAD. (A) A bioinformatics approach, integrating differential expression analysis (DEseq), survival analysis, and additive effects analysis, was used to identify four oncomiRs candidates. OC miR: onco-miRNA; LUAD: lung adenocarcinoma. (B) The box plot reveals the results of additive effect on survival analysis for five candidates oncomiRs from the previous correlation analysis (* indicates the adjusted p-value of DESeq). A total of 31 combinations were examined. Significant combinations were defined as those with log-rank p-values < 0.05 and HR values greater than 1. The red spot shows the highest hazard ratio within all 4-miR combinations. (C) The boxplots show the expression levels of the miRNA candidates in tumor and normal samples (* indicates the adjusted p-value of DESeq). (D) The KM (Kaplan-Meier) plot shows the 5-year overall survival analysis for the four-miRNA panel: all.high (N = 62; 12.55 %) and all.low (N = 68, 13.77 %). The KM plot of all stratification groups was shown in Supplementary Fig. 1B. The 5-year progression-free survival analysis result was shown in Supplementary Fig. 1C. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
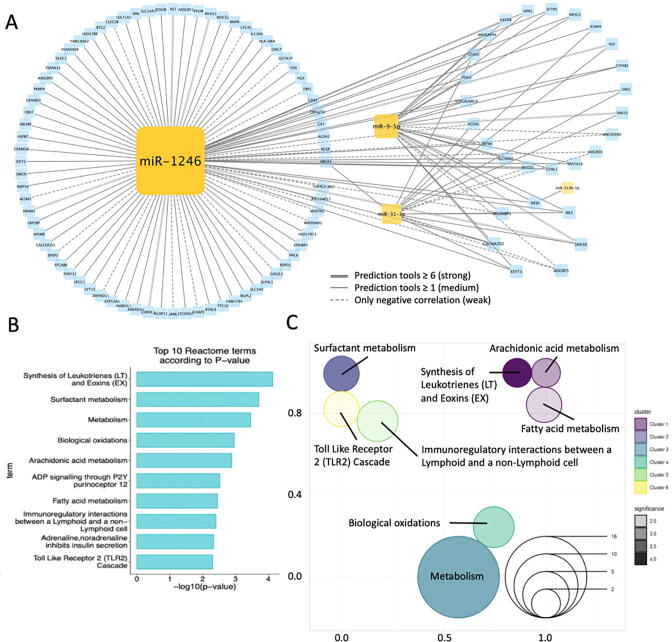
To further investigate the functions associated with these miRNA candidates, we identified their target genes and functions in a systematic approach (see Identification of miRNA-target genes in Materials and Methods for more details). In short, the target genes for each miRNA were defined by the negative correlation between the expression of miRNAs and target genes. The strength of the interaction was defined according to the number of positive results predicted by target prediction tools, which were used in our previous publications [18]. Fig. 2A shows the network constructed by the four miRNA candidates and their target genes, displaying several number of potential targets regulated by these four miRNA candidates. Functional enrichment analysis was performed to analyze target genes, and the top 10 Reactome terms are presented in Fig. 2B and Supplementary Fig. 2, revealing multiple terms focused on metabolism-related functions. In addition, the gene set representation displayed several main relevant functions: metabolism-related functions, immunoregulatory-related functions and others (Fig. 2C).
Fig. 2.
OncomiR-gene interaction network and annotation. (A) The interactions between miRNA candidates and their target genes were defined by prediction tools or negative correlation (see Supplementary Table 6). Strong: Negative correlation and more than 6 tools predict the interaction. Medium: Negative correlation and more than 1 tool predict the interaction. Weak: Only negative correlation but none of the tools predicted the interaction. (B) The top Reactome terms were identified in the functional enrichment analysis of the miRNAs’ target genes. (C) A representation of gene sets analysis indicates functions that focus on three main types with shorter distances. Different groups were shown as different colors and the corresponding significance (-log10 p-value of enrichment analysis) was shown as the shades of color, respectively. Two functions with insufficient genes (see Supplementary Table 7) were not shown in the figure.
4.2. In vitro assays for the validation of the four oncomiR candidates
We then examined the endogenous expression of four oncomiRs in different cell lines and found that LUAD cell lines had higher levels of four oncomiRs compared to human non-tumorigenic lung epithelial cell line BEAS-2B (Supplementary Fig. 3). To verify the effect of oncomiR candidates, we performed multiple functional assays to confirm the efficacy of the four miRNAs in three lung cancer cell lines (A549, Bm7, and Hop62). Supplementary Fig. 4 shows changes in miRNA expression levels over time following antagomiR transfection, including 2-fold decreases in miRNA expression when the respective antagomiR was transfected, either singly or in combination. Simultaneous inhibition of miR-1246, miR-9-5p, miR-31-3p, and miR-3136-5p significantly decreased cell growth compared to each antagomiR-transfected cell group and negative control group (Fig. 3A). Notably, the results of the MTT assay revealed that inhibition of miR-1246, miR-9-5p, miR-31-3p, and miR-3136-5p, suppressed the growth of LUAD cells (Supplementary Figure 5). The colony-forming assay for each antagomiR-transfected cell group showed 2-fold decreases in colony numbers compared with the negative control group. The combined antagomiR-transfected cells showed a more than 4-fold decrease, which was twice as effective as any single miRNA-targeted transfected (Fig. 3B). The migration and invasion assays demonstrated similar results, with 2-fold decreases observed for each individual antagomiR and a greater than 4-fold decrease in response to the combination of all four (Fig. 3C-E). These assays not only validate the four miRNA candidates but also confirmed the efficacy of targeting all four miRNAs together.
Fig. 3.
Inhibition of oncomiR candidates suppresses tumor progression in LUAD cells. (A–E) Cells were transfected with 20 nM antagomiRs for the following assays. The term “4 miRs” refers to the use of all 4 antagomiRs, with each antagomiR at a final concentration of 5 nM. anti-miR miRNA Inhibitor Negative Control was used as a negative control (NC). The transfected cells were examined using the trypan blue exclusion method. The results indicated that inhibition of miRNAs reduced the cell growth of lung cancer cells (A). The inhibition of miRNAs decreased A549, Bm7, and Hop62 colony numbers in soft agar (B). The inhibition of miRNAs suppressed the migration and invasion abilities of A549 (C), Bm7 (D), and Hop62 cells (E). The mean of three independent experiments performed in triplicate is depicted. The error bars depict the standard deviation of each mean. Mann Whitney U test, * p < 0.05 compared with cells transfected with NC. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
4.3. Animal validation of the four oncomiR candidates
To verify the role of these survival-related oncomiRs in vivo, the xenograft tumorigenicity and tail vein metastasis assays were performed using Bm7 cells, which had metastatic ability in an animal model. Negative Control Bm7 lung cancer cells and cells transfected with antagomiRs targeting all four miRNAs were injected into nude mice (n = 8 per group), and we examined the size of the resulting tumors, as shown in Fig. 4A. Surprisingly, the tumor volume and weight that developed from the injection of cells transfected with all four miRNA-targeting antagomiRs was 4-fold smaller than that in the negative control group (Fig. 4A and B). AntagomiR-mediated knockdown of all four miRNAs significantly reduced Ki-67 levels in the xenografts compared to the NC group (Fig. 4C and D). Fig. 4E indicates that the expression levels of these miRNAs were approximately-one-fourth of those in the negative control group. Accordingly, the mRNA and protein levels of oncomiR targets in tumor xenografts were significantly elevated by antagomiRs (Supplementary Figure 6A and B). The number of metastatic nodules in the lungs of the experimental animals was assessed, and the number of metastatic nodules in Bm7 cell-injected mice transfected with all four antagomiRs was almost one-fourth that in the negative control group (n = 6 per group; Fig. 4F and G). The levels of Ki-67 were decreased in the lung nodules in 4 antagomiRs groups compared to the NC group (Fig. 4H and I). The expression levels of all four miRNAs were approximately-one-fifth of those in the negative control group, as shown in Fig. 4J. The observed reductions in tumor volume and metastatic nodule numbers in the mouse model confirmed the efficacy of targeting all four miRNAs in combination.
Fig. 4.
Inhibition of oncomiR candidates represses tumor growth and lung metastasis in vivo. (A-E) Bm7 cells transfected with 4 antagomiRs or anti-miR miRNA Inhibitor Negative Control (NC) were subcutaneously injected into nude mice to perform xenograft assays. The graph depicted the experiment plan (A, upper). The inhibition of 4 miRNAs (miR-9-5p, miR-31-3p, miR-1246, and miR-3136-5p) significantly reduced tumor volumes (A, lower) and weights (B). The inhibition of 4 miRNAs suppressed Ki-67 levels in xenografts (C and D). The expression levels of the 4 targeted miRNAs in the xenografts were determined using quantitative real-time PCR (E); n = 8 per group. (F-J) Bm7 cells transfected with 4 antagomiRs or NC were intravenously injected into the tail veins of NOD-SCID mice to evaluate the in vivo metastatic activity. The lung metastasis capacity was diminished by the inhibition of all 4 miRNAs. The graph depicted the experiment plan (F, upper). Representative micrographs of NOD-SCID mouse lungs are shown (F, lower left). The lung sections were examined by hematoxylin and eosin staining (F, lower right). The metastatic nodules in the lungs were counted by gross and microscopic examination (G). The inhibition of 4 miRNAs suppressed Ki-67 levels in lung nodules (H and I). The expression levels of 4 miRNAs in the metastatic nodules were determined using quantitative real-time PCR (J); n = 6 per group. The mean of three independent experiments performed in triplicate is depicted. The error bars depict the standard deviation of each mean. Mann Whitney U test, * p < 0.05 compared with NC.
5. Discussion
In this study, we applied a systematic approach that integrated multiple bioinformatics methods to identify four miRNA candidates as oncomiRs with oncogenic roles (miR-9-5p, miR-31-3p, miR-1246, and miR-3136-5p), and conducted multiple functional assays and mouse model experiments to validate the efficacy of inhibiting all four oncomiRs together by applying antagomiRs for all four miRNAs. Our work was a proof of concept that the miRNA candidates could be identified by a bioinformatic approach and then validated in the mouse model. The prognostic power of combining four oncomiRs outperformed any one of them individually in a retrospective cohort. The results of the animal model also imply that the introduction of antagomiRs of the four oncomiRs to the target RNAs may result in the suppression of cancer cell proliferation or metastasis [32]. This suggested that the four oncomiRs may be potential targets of miRNA-based treatments for lung adenocarcinoma.
Multiple studies have supported an association between miRNAs and amelioration of lung cancer with several factors [33]. To develop miRNAs as diagnostic biomarkers, several studies have identified the dysregulation of miRNAs in various lung cancer subtypes. For example, the use of five circulating miRNAs can be used as a molecular signature for the early detection of NSCLC [34]. The finding may support the identification of miRNA candidates has the potential of providing clinically significant diagnostic information. Several miRNA-based therapies have progressed to the clinical trial phase, such as antagomiR-155 for CTCL and mycosis fungoides or the miR-16 mimic for mesothelioma and NSCLC, which is currently being examined in multicenter Phase I trials [7]. A previous study indicated that the high expression of miR-21-5p is associated with resistance to platinum-based chemotherapy, and these patients require anti-miR-21 treatment to enhance phosphatase and tensin homolog (PTEN) expression [33]. This research suggested that miRNA regulation could be significant for future combinatorial therapeutic options. This study aimed to identify dysregulated and survival-relevant miRNA candidates that might serve a therapeutic role in improving clinical outcomes.
In this study, we identified dysregulated and survival-relevant miRNA candidates that might serve as biomarkers and therapeutic targets for LUAD. We used the brain metastatic LUAD cell line Bm7 for in vivo experiments. To create a mouse model of brain metastasis, the mice were administered an intracardiac injection of cancer cells for whole-body distribution rather than retention in the lungs [25]. However, Bm7 cells can induce lung metastasis through tail vein injection [35]. In this study, we investigated the effect of antagomiRs on metastatic ability rather than on brain metastasis. Thus, we chose safer and simpler routes of intravenous injection instead of intracardiac injection to represent lung metastasis. Indeed, antagomiRs significantly reduce the metastatic ability of lung cancer cells in mice. Studying the metastatic potential of Bm7 cells in the brain could strengthen our conclusions. In contrast, primary lung cancer cell lines could be used to evaluate tumor growth. We examined the expression of miRNAs in xenografts after 30 days of transplantation and found low levels of miRNAs in the 4-antagomiRs group compared to those in the NC control. Zhou et al. demonstrated the long-term effects of antagomiR-mediated miRNA inhibition after transfection [36]. In addition, previous studies have suggested that antagomiR lasts long and is stable in vivo [37], [38]. Thus, antagomiR-mediated regulation of miR expression is a potential strategy for cancer treatment. Of note, to mimic the clinical conditions of cancer treatment, intratumoral delivery or systemic administration with antagomiRs in tumor-bearing mice could be performed.
Four oncomiRs were identified by our bioinformatics approach, and several studies have confirmed the role of some candidates in lung cancer. Li et al. demonstrated that miR-9-5p leads to tumor cell growth and metastasis in NSCLC [39]. MiR-31-3p has been found to play an oncogenic role and is upregulated in lung cancer and other cancers in multiple studies [40], [41], [42]. In addition, miR-1246 has been identified as a prognostic biomarker with oncogenic function in NSCLC [43], [44]. Although miR-3136-5p has been found associated with colorectal cancer [45], and serous ovarian carcinoma [46], currently there still no study has identified an oncogenic role for miR-3136-5p in lung cancer. As a consequence, this is the first study to identify an oncogenic role for this miRNA in lung cancer, but the mechanism of miR-3136-5p is still needed to be studied in the future. Furthermore, we are also the first study that demonstrates the combination of the four candidates has a synergic effect in the aspect of patients’ 5-year overall survival. Therefore, our study not only supports existing miRNA candidates with significant roles in lung cancer but was also able to identify a new candidate for the development of miRNA-based therapeutics. Moreover, to visualize miRNA expression in tumor tissues using in situ hybridization-based assays provides the precise localization and expression pattern of miRNA within a histologic section, which may serve as a useful biomarker.
The four oncomiRs target multiple genes were identified through our integrative bioinformatics analysis. In the network, many genes appear to be regulated by more than one miRNA, and miR-1246 appears to target the majority of the identified genes. When the functions of those genes were analyzed, these genes were predominantly associated with cell metabolism. In addition to the target genes of miR-1246, miR-31-3p was previously identified to suppress succinate dehydrogenase complex subunit A (SDHA) expression which plays a crucial role in mitochondrial metabolism in an induced pluripotent stem cell model [47]. MiR-9-5p was shown to regulate Glutamic-Oxaloacetic Transaminase 1 (GOT1) and inhibit pancreatic cancer cell proliferation and invasion [48]. On the other hand, there were several studies had mentioned that miRNA-targeted genes were metabolism relevant. SELENBP1 was found to involve energy metabolism in prostate cancer [49]. AK1 catalyses the reversible reaction, ATP + AMP ↔ 2ADP, and played a crucial role in energy homeostasis involved in the synthesis, equilibration, and regulation of adenine nucleotide [50]. CACNA2D2 was a subunit of the Ca2+-channel complex and was found to be associated with disruption of mitochondria membrane integrity in NSCLC cells [51]. These studies together with our functional assay results may suggest that the four miRNA candidates could be associated with the regulation of cancer cell metabolism. Remarkably, four oncomiRs-associated genes were also enriched in immunoregulatory-related functions. SFTPD, surfactant proteins D, is crucial for maintenance of lung homeostasis as well as orchestrates the innate and adaptive immune system [52]. JAML-knockout mice showed the increased tumor growth, impaired γδ tumor-infiltrating lymphocytes response, and increased CD8 T cell dysfunction. Treatment with JAML agonist may serve as a novel strategy for immunotherapy via facilitating CD8 and γδ T cell anti-tumor immunity [53]. CD300LG has been reported downregulated in lung cancer tissues which might link to immune escape [54]. ICAM4 is critical for immune synapse formation between natural killer cells and hepatocellular carcinoma cells to promote natural killer cell-mediated cancer immunotherapeutic activity [55]. These findings highlight the potential role of targeting four oncomiRs in the LUAD immunotherapy. Our in vivo study revealed that knockdown of all four oncomiRs increased the expression levels of the six target genes in tumors (Supplementary Figure 6). Interestingly, five out of six genes, AK1, CACNA2D2, NFIX, SELENBP1, and SNX30, have been reported to be associated with a favorable prognosis of LUAD [56], [57], [58], [59], [60]. These findings may partly explain the additive effect of multiple miRNAs on survival.
In this study, we applied an additive effect analysis to identify miRNA candidates with the largest changes in HR values. These results indicated that the combination of the four identified oncomiRs resulted in the most significant changes in the survival analysis. This finding was verified in the experiments performed on cancer cell lines and the xenografted mouse model, which demonstrated more effective outcomes when all four miRNAs were targeted in combination. As results are shown in this study, the four clinical-relevant oncomiRs are valuable candidates for the development of miRNA-based treatment. However, we did not investigate the mechanism behind the four combined miRNAs which needed more experiments to validate the mechanism in detail. Still, more experiments are needed to prove the performance of this panel as the diagnostic biomarker or the effect of respective antagomiRs as the correspondent treatment method. Such as another cohort study to decipher how this panel is to be integrated into current LUAD treatment steps [38]. In addition, before entering a first-in-human study, although there are several technologies like locked nucleic acid (LNA) [61], [62], lipid nanoparticle (LNP) [63], and liposome [64], yet there are still various challenges of applying our miRNA candidates including the drug formation and nucleotide deliver efficiency. We believed that with more real-world evidence, this approach may reduce multiple diagnostic and treatment steps and improve patient care in the future [65].
Author Contributions
W-C.C. and K-W.H. conceived and designed the data analysis and the experiments. K-W.H., S-H.L., L-J.L., P-H. P., and J-L.C. performed the experiments., C-H.L., Y-L.L., and C-Y.L. analyzed the data. F-H.C., Y-P.S., H-H.W., and S-C.W. contributed reagents, materials, and/or analysis tools. C-H.L., W-C.C., K-W.H., M-Z.W., C-Y.C., and S-H.L. wrote the paper. All authors reviewed the manuscript.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
This research was funded by Ministry of Science and Technology [MOST 108-2622-E-039-005-CC2; MOST 109-2622-E-039-004-CC2; MOST 109-2314-B-182-078-MY3; MOST 109-2327-B-039-002; MOST 111-2628-B-182-008; MOST 109-2628-E-039- 001-MY3; MOST 109-2628-B-039-006; MOST 110-2628-B-039-007; MOST 111-2628-B-039-007-MY3]; China Medical University (CMU111-MF-72); China Medical University Hospital [DMR-109-055;DMR-109-223; DMR-110-072; DMR-111-249]; The “Drug Development Center, China Medical University” from The Featured Areas Research Center Program within the framework of the Higher Education Sprout Project (Ministry of Education, Taiwan).
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.csbj.2022.08.042.
Contributor Information
Wei-Chung Cheng, Email: wccheng@mail.cmu.edu.tw.
Kai-Wen Hsu, Email: kwhsu@mail.cmu.edu.tw.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
References
- 1.Barta J.A., Powell C.A., Wisnivesky J.P. Global Epidemiology of Lung Cancer. Ann Glob. Health. 2019;85(1) doi: 10.5334/aogh.2419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Sung H., et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a cancer journal for clinicians. 2021;71(3):209–249. doi: 10.3322/caac.21660. [DOI] [PubMed] [Google Scholar]
- 3.Chen J., et al. Genomic landscape of lung adenocarcinoma in East Asians. Nat Genet. 2020;52(2):177–186. doi: 10.1038/s41588-019-0569-6. [DOI] [PubMed] [Google Scholar]
- 4.Grzywa T.M., Klicka K., Włodarski P.K. Regulators at Every Step-How microRNAs Drive Tumor Cell Invasiveness and Metastasis. Cancers (Basel) 2020;12(12) doi: 10.3390/cancers12123709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Lee Y.S., Dutta A. MicroRNAs in cancer. Annu Rev Pathol. 2009;4:199–227. doi: 10.1146/annurev.pathol.4.110807.092222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wu M., et al. MiRNA-based Therapeutics for Lung Cancer. Curr Pharm Des. 2018;23(39):5989–5996. doi: 10.2174/1381612823666170714151715. [DOI] [PubMed] [Google Scholar]
- 7.Rupaimoole R., Slack F.J. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017;16(3):203–222. doi: 10.1038/nrd.2016.246. [DOI] [PubMed] [Google Scholar]
- 8.Chakraborty C., et al. Therapeutic advances of miRNAs: A preclinical and clinical update. J Adv Res. 2021;28:127–138. doi: 10.1016/j.jare.2020.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hanna J., Hossain G.S., Kocerha J. The potential for microRNA therapeutics and clinical research. Front Genet. 2019;10:478. doi: 10.3389/fgene.2019.00478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Campbell J.D., et al. Distinct patterns of somatic genome alterations in lung adenocarcinomas and squamous cell carcinomas. Nat Genet. 2016;48(6):607–616. doi: 10.1038/ng.3564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gilad S., et al. Classification of the four main types of lung cancer using a microRNA-based diagnostic assay. J Mol Diagn. 2012;14(5):510–517. doi: 10.1016/j.jmoldx.2012.03.004. [DOI] [PubMed] [Google Scholar]
- 12.Wu K.L., et al. The Roles of MicroRNA in Lung Cancer. Int J Mol Sci. 2019;20(7) doi: 10.3390/ijms20071611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Cheng W.C., et al. DriverDB: an exome sequencing database for cancer driver gene identification. Nucleic Acids Res. 2014;42(Database issue):D1048–D1054. doi: 10.1093/nar/gkt1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chung I.F., et al. DriverDBv2: a database for human cancer driver gene research. Nucleic Acids Res. 2016;44(D1):D975–D979. doi: 10.1093/nar/gkv1314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Liu S.H., et al. DriverDBv3: a multi-omics database for cancer driver gene research. Nucleic Acids Res. 2019 doi: 10.1093/nar/gkz964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cheng W.C., et al. YM500: a small RNA sequencing (smRNA-seq) database for microRNA research. Nucleic Acids Res. 2013;41(Database issue):D285–D294. doi: 10.1093/nar/gks1238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cheng W.C., et al. YM500v2: a small RNA sequencing (smRNA-seq) database for human cancer miRNome research. Nucleic Acids Res. 2015;43(Database issue):D862–D867. doi: 10.1093/nar/gku1156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chung I.F., et al. YM500v3: a database for small RNA sequencing in human cancer research. Nucleic Acids Res. 2017;45(D1):D925–D931. doi: 10.1093/nar/gkw1084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kozomara A., Griffiths-Jones S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014;42(D1):D68–D73. doi: 10.1093/nar/gkt1181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Leung, Y.Y., et al., DASHR: database of small human noncoding RNAs. Nucleic acids research, 2016. 44(D1): p. D216-D222. [DOI] [PMC free article] [PubMed]
- 21.Anders S., Huber W. Differential expression analysis for sequence count data. Genome Biol. 2010;11(10):R106. doi: 10.1186/gb-2010-11-10-r106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Therneau T.M., Lumley T. Package ‘survival’. R Top Doc. 2015;128:112. [Google Scholar]
- 23.Meng X., et al. CancerNet: a database for decoding multilevel molecular interactions across diverse cancer types. Oncogenesis. 2015;4(12):e177–e. doi: 10.1038/oncsis.2015.40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Tokar T., Pastrello C., Jurisica I. GSOAP: A tool for visualisation of gene set over-representation analysis. Bioinformatics. 2020 doi: 10.1093/bioinformatics/btaa001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lin C.Y., et al. ADAM9 promotes lung cancer metastases to brain by a plasminogen activator-based pathway. Cancer Res. 2014;74(18):5229–5243. doi: 10.1158/0008-5472.CAN-13-2995. [DOI] [PubMed] [Google Scholar]
- 26.Schwarzenbach H., et al. Data Normalization Strategies for MicroRNA Quantification. Clin Chem. 2015;61(11):1333–1342. doi: 10.1373/clinchem.2015.239459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hsu K.W., et al. Downregulation of tumor suppressor MBP-1 by microRNA-363 in gastric carcinogenesis. Carcinogenesis. 2014;35(1):208–217. doi: 10.1093/carcin/bgt285. [DOI] [PubMed] [Google Scholar]
- 28.Hsu K.W., et al. The activated Notch1 receptor cooperates with alpha-enolase and MBP-1 in modulating c-myc activity. Mol Cell Biol. 2008;28(15):4829–4842. doi: 10.1128/MCB.00175-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Hsu K.W., et al. MBP-1 suppresses growth and metastasis of gastric cancer cells through COX-2. Mol Biol Cell. 2009;20(24):5127–5137. doi: 10.1091/mbc.E09-05-0386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hsu K.W., et al. Activation of the Notch1/STAT3/Twist signaling axis promotes gastric cancer progression. Carcinogenesis. 2012;33(8):1459–1467. doi: 10.1093/carcin/bgs165. [DOI] [PubMed] [Google Scholar]
- 31.Tsai L.H., et al. Targeting interleukin-17 receptor B enhances gemcitabine sensitivity through downregulation of mucins in pancreatic cancer. Sci Rep. 2020;10(1):17817. doi: 10.1038/s41598-020-73659-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Inoue J., Inazawa J. Cancer-associated miRNAs and their therapeutic potential. J Hum Genet. 2021:1–9. doi: 10.1038/s10038-021-00938-6. [DOI] [PubMed] [Google Scholar]
- 33.Iqbal M.A., et al. MicroRNA in lung cancer: role, mechanisms, pathways and therapeutic relevance. Mol Aspects Med. 2019;70:3–20. doi: 10.1016/j.mam.2018.07.003. [DOI] [PubMed] [Google Scholar]
- 34.Geng Q., et al. Five microRNAs in plasma as novel biomarkers for screening of early-stage non-small cell lung cancer. Respir Res. 2014;15:149. doi: 10.1186/s12931-014-0149-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liu S.H., et al. Systematic identification of clinically relevant miRNAs for potential miRNA-based therapy in lung adenocarcinoma. Mol Ther Nucleic Acids. 2021;25:1–10. doi: 10.1016/j.omtn.2021.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhou W., et al. MicroRNA-20b promotes cell growth of breast cancer cells partly via targeting phosphatase and tensin homologue (PTEN) Cell Biosci. 2014;4(1):62. doi: 10.1186/2045-3701-4-62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Krutzfeldt J., et al. Silencing of microRNAs in vivo with 'antagomirs'. Nature. 2005;438(7068):685–689. doi: 10.1038/nature04303. [DOI] [PubMed] [Google Scholar]
- 38.Cerro-Herreros E., et al. Therapeutic Potential of AntagomiR-23b for Treating Myotonic Dystrophy. Mol Ther Nucleic Acids. 2020;21:837–849. doi: 10.1016/j.omtn.2020.07.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Li G., et al. MiR-9-5p promotes cell growth and metastasis in non-small cell lung cancer through the repression of TGFBR2. Biomed Pharmacother. 2017;96:1170–1178. doi: 10.1016/j.biopha.2017.11.105. [DOI] [PubMed] [Google Scholar]
- 40.Davenport M.L., et al. miR-31 displays subtype specificity in lung cancer. Cancer Res. 2021 doi: 10.1158/0008-5472.CAN-20-2769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Lv C., et al. MiR-31 promotes mammary stem cell expansion and breast tumorigenesis by suppressing Wnt signaling antagonists. Nat Commun. 2017;8(1):1036. doi: 10.1038/s41467-017-01059-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Zhang J., et al. Integrative analysis of mRNA and miRNA expression profiles reveals seven potential diagnostic biomarkers for nonsmall cell lung cancer. Oncol Rep. 2020;43(1):99–112. doi: 10.3892/or.2019.7407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Fan L., et al. Aberrant miR-1246 expression promotes radioresistance in non-small cell lung cancer: a potential prognostic biomarker and radiotherapy sensitization target. Am J Cancer Res. 2020;10(1):314–335. [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 44.Huang D., Qu D. Early diagnostic and prognostic value of serum exosomal miR-1246 in non-small cell lung cancer. Int J Clin Exp Pathol. 2020;13(7):1601–1607. [PMC free article] [PubMed] [Google Scholar]
- 45.Wang L., et al. miRNA Expression Profile in the N2 Phenotype Neutrophils of Colorectal Cancer and Screen of Putative Key miRNAs. Cancer Management and Research. 2020;12:5491. doi: 10.2147/CMAR.S251427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Chen S.F., et al. Identification of core aberrantly expressed microRNAs in serous ovarian carcinoma. Oncotarget. 2018;9(29):20451. doi: 10.18632/oncotarget.24942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lee M.R., et al. MiR-31/SDHA axis regulates reprogramming efficiency through mitochondrial metabolism. Stem Cell Rep. 2016;7(1):1–10. doi: 10.1016/j.stemcr.2016.05.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wang J., et al. miR-9-5p inhibits pancreatic cancer cell proliferation, invasion and glutamine metabolism by targeting GOT1. Biochem Biophys Res Commun. 2019;509(1):241–248. doi: 10.1016/j.bbrc.2018.12.114. [DOI] [PubMed] [Google Scholar]
- 49.Elhodaky M., et al. Selenium-binding protein 1 alters energy metabolism in prostate cancer cells. Prostate. 2020;80(12):962–976. doi: 10.1002/pros.24028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Dzeja P., Chung S., Terzic A. Integration of adenylate kinase and glycolytic and clycogenolytic circuits in cellular energetics. Wiley–VCH; Weinheim, Germany: 2007. [Google Scholar]
- 51.Carboni G.L., et al. CACNA2D2-mediated apoptosis in NSCLC cells is associated with alterations of the intracellular calcium signaling and disruption of mitochondria membrane integrity. Oncogene. 2003;22(4):615–626. doi: 10.1038/sj.onc.1206134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Watson A., Madsen J., Clark H.W. SP-A and SP-D: Dual Functioning Immune Molecules With Antiviral and Immunomodulatory Properties. Front Immunol. 2020;11 doi: 10.3389/fimmu.2020.622598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.McGraw J.M., et al. JAML promotes CD8 and gammadelta T cell antitumor immunity and is a novel target for cancer immunotherapy. J Exp Med. 2021;218(10) doi: 10.1084/jem.20202644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zhai S., et al. Expression Depression of CD300LG-gamma in Human Pulmonary Carcinoma. Monoclon Antib Immunodiagn Immunother. 2016;35(2):94–99. doi: 10.1089/mab.2015.0043. [DOI] [PubMed] [Google Scholar]
- 55.Kim M., et al. Novel natural killer cell-mediated cancer immunotherapeutic activity of anisomycin against hepatocellular carcinoma cells. Sci Rep. 2018;8(1):10668. doi: 10.1038/s41598-018-29048-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Jan Y.H., et al. A co-expressed gene status of adenylate kinase 1/4 reveals prognostic gene signature associated with prognosis and sensitivity to EGFR targeted therapy in lung adenocarcinoma. Sci Rep. 2019;9(1):12329. doi: 10.1038/s41598-019-48243-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Xu J.Y., et al. Integrative Proteomic Characterization of Human Lung Adenocarcinoma. Cell. 2020;182(1):245–261 e17. doi: 10.1016/j.cell.2020.05.043. [DOI] [PubMed] [Google Scholar]
- 58.Ge J., et al. NFIX downregulation independently predicts poor prognosis in lung adenocarcinoma, but not in squamous cell carcinoma. Future Oncol. 2018;14(30):3135–3144. doi: 10.2217/fon-2018-0164. [DOI] [PubMed] [Google Scholar]
- 59.Caswell D.R., et al. Tumor Suppressor Activity of Selenbp1, a Direct Nkx2-1 Target, in Lung Adenocarcinoma. Mol Cancer Res. 2018;16(11):1737–1749. doi: 10.1158/1541-7786.MCR-18-0392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Shi X., et al. Integrative pan cancer analysis reveals the importance of CFTR in lung adenocarcinoma prognosis. Genomics. 2022;114(2) doi: 10.1016/j.ygeno.2022.110279. [DOI] [PubMed] [Google Scholar]
- 61.Chabot S., et al. LNA-based oligonucleotide electrotransfer for miRNA inhibition. Molecular Therapy. 2012;20(8):1590–1598. doi: 10.1038/mt.2012.95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Mook O.R., et al. Evaluation of locked nucleic acid–modified small interfering RNA in vitro and in vivo. Mol Cancer Ther. 2007;6(3):833–843. doi: 10.1158/1535-7163.MCT-06-0195. [DOI] [PubMed] [Google Scholar]
- 63.Hou X., et al. Lipid nanoparticles for mRNA delivery. Nat Rev Mater. 2021;6(12):1078–1094. doi: 10.1038/s41578-021-00358-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Sercombe L., et al. Advances and Challenges of Liposome Assisted Drug Delivery. Front Pharmacol. 2015;6 doi: 10.3389/fphar.2015.00286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Palekar-Shanbhag P., et al. Theranostics for cancer therapy. Curr Drug Deliv. 2013;10(3):357–362. doi: 10.2174/1567201811310030013. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.