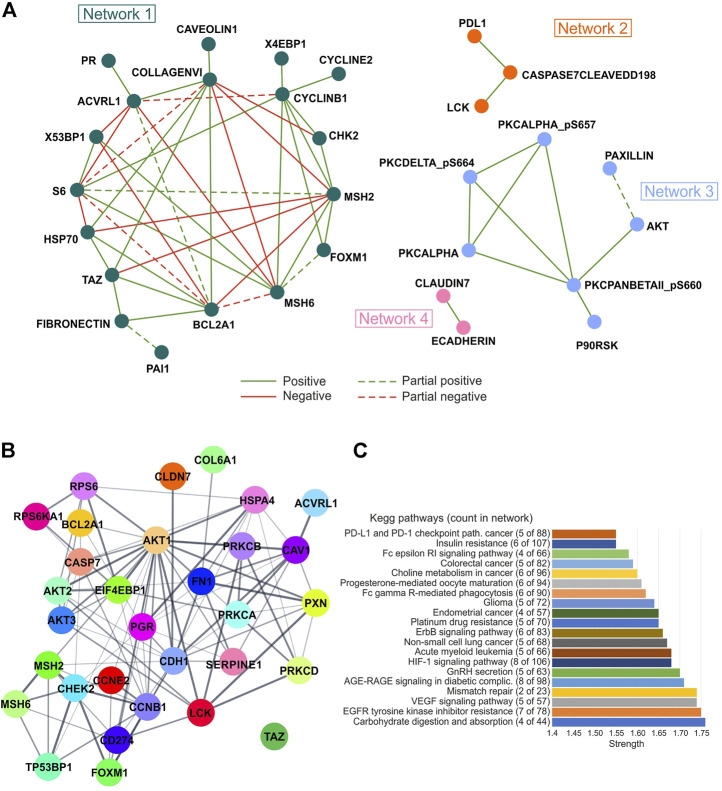
FIGURE 6.
Correlation and pathway analysis of protein levels analyzed in each tumor type. (A) Protein–protein interaction showing correlation (Pearson’s r ≥ 0.3 or ≤ −0.3; p ≤ 0.05) in at least 20 tumor types are represented. The concordance among Pearson’s value means was defined as a degree of agreement among the correlation sign. Partial concordance: dotted interaction lines. Total concordance: solid interaction lines. The network was obtained from Cytoscape 3.8.2 (https://cytoscape.org/). (B) STRING network analysis (https://string-db.org/) of selected proteins significantly correlated with each other (Pearson’s r ≥ 0.3 or ≤ −0.3; p ≤ 0.05). Edge thickness indicates the strength of data support. (C) Gene ontology analysis according to STRING algorithms. Strength of functional enrichments and count of proteins in each KEGG pathway are indicated.