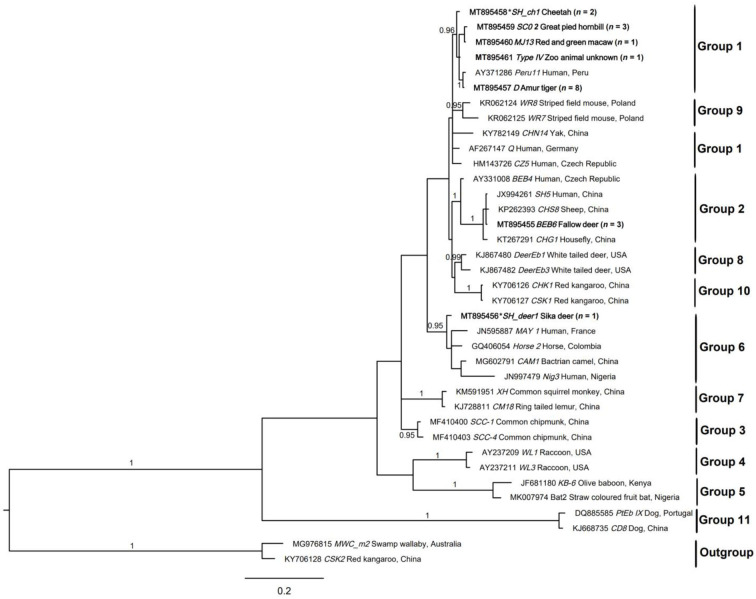
Figure 2.
Relationships among the genotypes of Enterocytozoon bieneusi recorded in the wildlife in this study inferred from the phylogenetic analysis of sequence data for the internal transcribed spacer (ITS) of nuclear ribosomal DNA by Bayesian inference (BI). Statistically significant posterior probabilities (pps) are indicated on branches. Individual GenBank accession numbers precede genotype designation (in italics) followed by sample and locality descriptions. The Enterocytozoon bieneusi genotypes identified and characterized from fecal DNA samples in the present study are indicated in bold type. Clades were assigned group names based on the classification system established by Karim et al. (2015) and Li et al. (2019a). The scale bar represents the number of substitutions per site. The E. bieneusi genotypes PtEbIX (DQ885585) and CD8 (KJ668735) from dogs were used as outgroups. All the groups were strongly supported (pp = 0.96–1). pp < 0.95 were not shown.