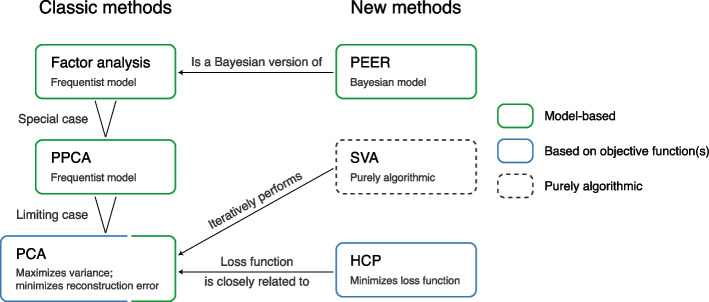
Fig. 6.
PCA, SVA, PEER, and HCP are closely related statistical methods despite their apparent dissimilarities. In particular, the methodology behind SVA, PEER, and HCP can all be traced back to PCA. PCA [34–38] is traditionally derived by optimizing some objective functions (either maximum variance or minimum reconstruction error; Additional file 1: Section S5.1), but more recently, it is shown that PCA can be derived as a limiting case of probabilistic principal component analysis (PPCA) [45], which in turn is a special case of factor analysis [35, 46]. PEER [24, 32] is based on a Bayesian probabilistic model and can be considered a Bayesian version of factor analysis. SVA [25, 26] is purely algorithmic and is not defined based on a probabilistic model or objective function. The steps of the SVA algorithm are complicated [44], but in a nutshell, SVA iterates between two steps: (1) reweight the features of the molecular phenotype matrix, and (2) perform PCA on the resulting matrix (with centering but without scaling) [26]. Lastly, HCP [33] is defined by minimizing a loss function that is very similar to the minimum-reconstruction-error loss function of PCA (Additional file 1: Section S5.2)