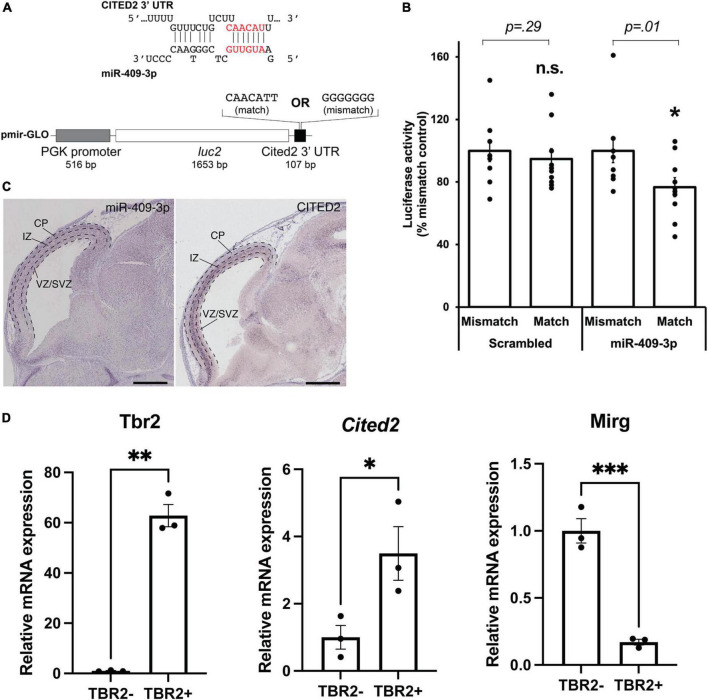
FIGURE 1.
miR-409-3p represses the transcriptional regulator Cited2. (A) Sequence alignment demonstrating the predicted miR-409-3p target site in the Cited2 3′ UTR. Seed sequence base-pairing is in red. Luciferase reporter constructs were generated with the Cited2 3′ UTR fused to luc2, containing the wild-type predicted miR-409-3p target sequence (match). Mismatch reporter vectors were identical except that the target sequence was replaced by a sequence that would not be recognized by miR-409-3p (mismatch). (B) miR-409-3p oligonucleotides repress a Cited2 3′UTR luciferase reporter gene bearing wild-type, but not mismatch, miR-409-3p target sequences. Scrambled miRNA does not repress the Cited2 reporter. Error bars represent SEM. *p < 0.05 compared to mismatch control. (C) In the e14.5 dorsal telencephalon, miR-409-3p is enriched in the VZ and CP, and not in the upper SVZ and IZ, with minimal overlap with Cited2. Dashed lines indicate boundaries of the ventricular zone/sub-ventricular zone (VZ/SVZ), intermediate zone (IZ), and cortical plate (CP). Scale bars = 1 mm. Raw images from the Eurexpress database (Diez-Roux et al., 2011). (D) Cited2 expression is enriched in IPCs at e15.5 by qPCR, while Mirg, which encodes miR-409-3p, is enriched in the TBR2- cells. Error bars represent SEM. Each dot depicts an independent biological replicate. *p < 0.05; **p < 0.01 (Two-tailed t-test for Cited2 and Mirg; Welch’s correction was performed for Tbr2 due to unequal variance).