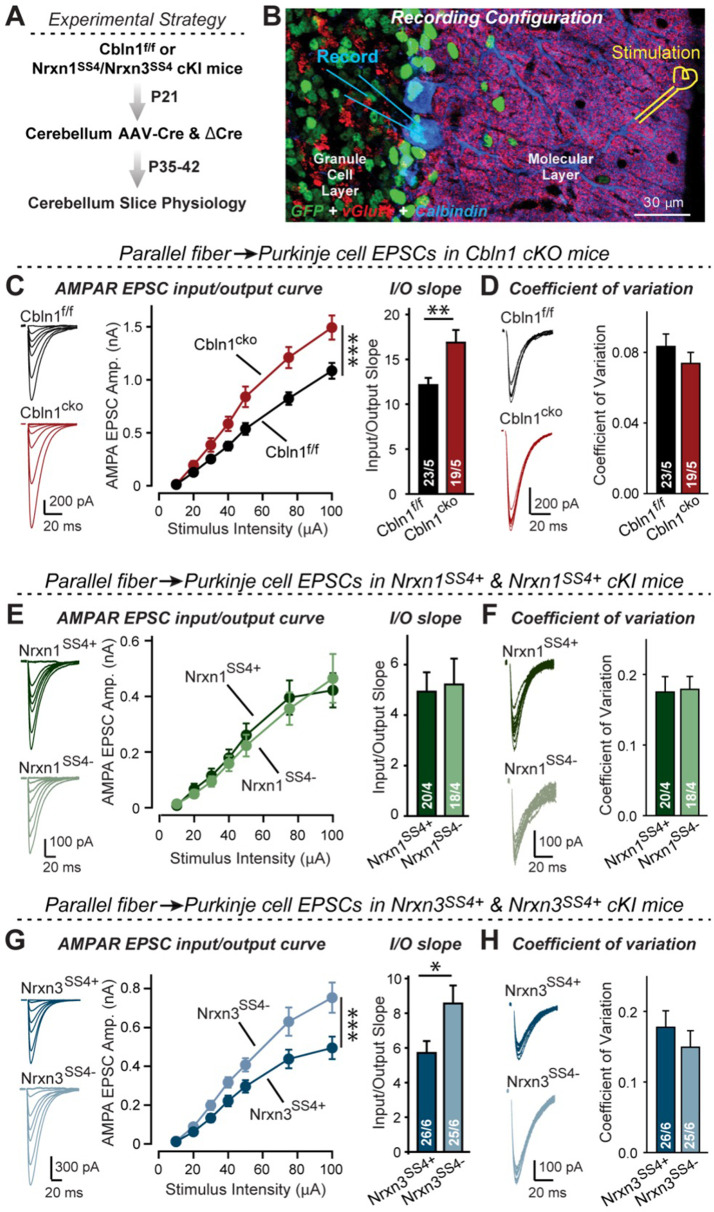
Figure 8. Nrxn3SS4+-Cbln1 signaling controls AMPAR-EPSCs in the cerebellum, but in this brain region Nrxn1SS4+-Cbln1 signaling has no effect.
(A) Experimental workflow for analyzing the effect of the Cbln1 cKO or of the conditional Nrxn1SS4+ or Nrxn3SS4+ knockin on parallel-fiber synaptic transmission in the cerebellum. Note that the expression of ΔCre in Nrxn1SS4+ or Nrxn3SS4+ knockin mice retains the constitutive expression of their SS4 +splice variants, whereas the expression of Cre converts SS4 +into a constitutive SS4- splice variant. (B) Image of a cerebellar cortex section (lobes 4–5) from Cbln1 cKO mouse in which these lobes were infected at P21 by stereotactic injections of AAVs expressing ΔCre-eGFP (Cbln1f/f) or Cre-eGFP (Cbln1cKO). Sections were analyzed at P35 by slice physiology; the positions of the recording electrode in the patched Purkinje cells and of the stimulation electrode in the granule cell layer are indicated. (C) The Cbln1 deletion in cerebellum significantly increases the amplitude of AMPAR-EPSCs at parallel-fiber synapses (left, sample traces of evoked AMPAR-EPSCs; middle, summary plot of AMPAR-EPSCs input-output curves; right, summary graph of the slope of AMPAR-EPSC input/output curves). (D) The Cbln1 deletion in cerebellum has no major effect on the coefficient of variation at parallel-fiber synapses, suggesting that it does not greatly change the release probability (left, sample traces of evoked AMPAR-EPSCs with 50 μA stimulus intensity; right, summary graph of the coefficient of variation of AMPAR-EPSCs). (E & F) Same as (C & D) but recorded from Nrxn1SS4+ knockin mice in which ΔCre retains a constitutive expression of Nrxn1-SS4+splice variants, whereas Cre converts the Nrxn1-SS4+variants into constitutive Nrxn1-SS4- variants. (G & H) Same as (E & F) but for Nrxn3SS4+ knockin mice in which ΔCre retains a constitutive expression of Nrxn1-SS4+splice variants, whereas Cre converts the Nrxn1-SS4+variants into constitutive Nrxn1-SS4- variants. Data are means ± SEM. Number of neurons/mice are indicated in bars. Statistical significance was assessed by two-way ANOVA or unpaired two-tailed t-test (*p≤0.05, **p≤0.01, and ***p≤0.001).