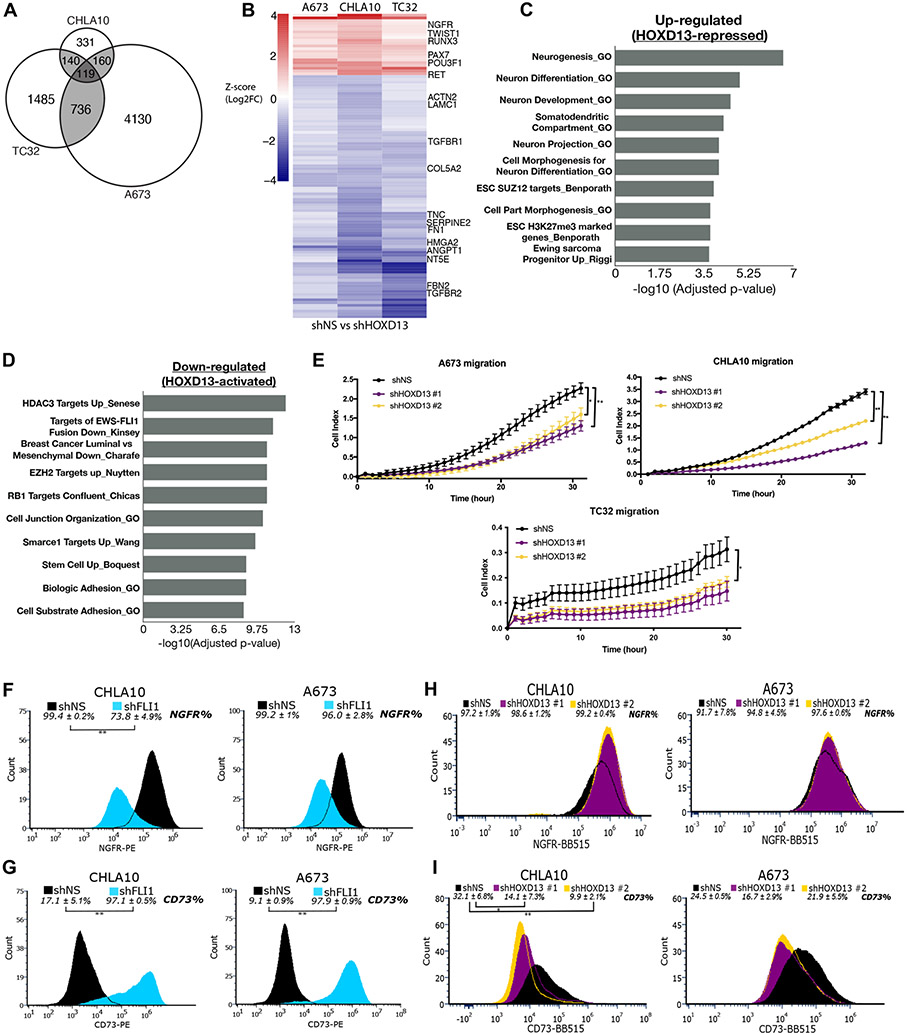
Figure 4. HOXD13 regulates neuro-mesenchymal gene programs and influences cell states.
A) Venn diagram showing overlap of significantly differentially expressed genes in each cell line. B) Heatmap depicting the differentially expressed genes (padj <0.05) between shHOXD13 and shNS for all cell lines. Scale: Z-score (Log2FC). Overrepresentation analysis of top 10 gene sets C) up- and D) down-regulated following HOXD13 knockdown in all cell lines. E) xCELLigence Real Time Cell Migration assay of control and HOXD13 knockdown cells. Error bars are representative of SEM from at least two independent experiments with 5 technical replicates per condition. * p<0.05; ** p<0.01. Flow cytometry histograms showing the shift in F) NGFR + cells and G) CD73+ cells upon EWS::FLI1 knockdown. Flow cytometry histograms showing the shift in H) NGFR + cells and I) CD73+ cells upon HOXD13 knockdown. Flow Error represents SD from at least three independent experiments. * p<0.05; ** p<0.01; Two-tailed t-test.