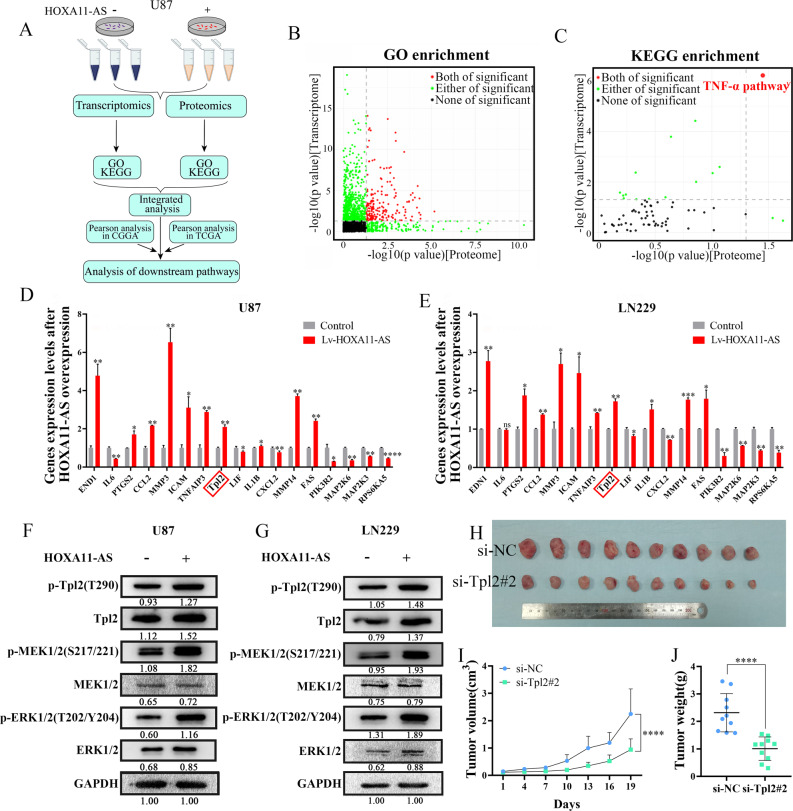
Fig. 4. TNF-α pathway was an important downstream pathway for HOXA11-AS via integrated analysis of transcriptomics and proteomics.
A The overall workflow of integrated analysis of transcriptomics and proteomics. B Scatterplot of GO items in both transcriptomic and proteomic data sets. C Scatterplot of KEGG items in both transcriptomic and proteomic data sets. The red plots indicated these items significant enrichment of both omics; the green plots indicated these items enrichment of single omics; the black plots indicated these items were not enriched in both omics. The expression levels of 17 DEGs enriched in the TNF-α pathway from transcriptomics were verified by RT-qPCR in U87 (D) and LN229 (E) cells. Each data point was calculated from averages of biological triplicates. Key proteins in TNF-α pathway were validated by western blot in U87 (F) and LN229 (G) cells infected with Lv-HOXA11-AS. GAPDH was used as an internal control. Xenograft mouse models were generated by subcutaneous injection of U87 cells into nude mice. Seven days after implanted, mice were treated with si-NC (n = 10) or si-Tpl2 (n = 10). Representative image of tumor dissected tumors (H), growth curve (I) and tumor weight (J) in si-NC or si-Tpl2-treated xenograft mouse models. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.