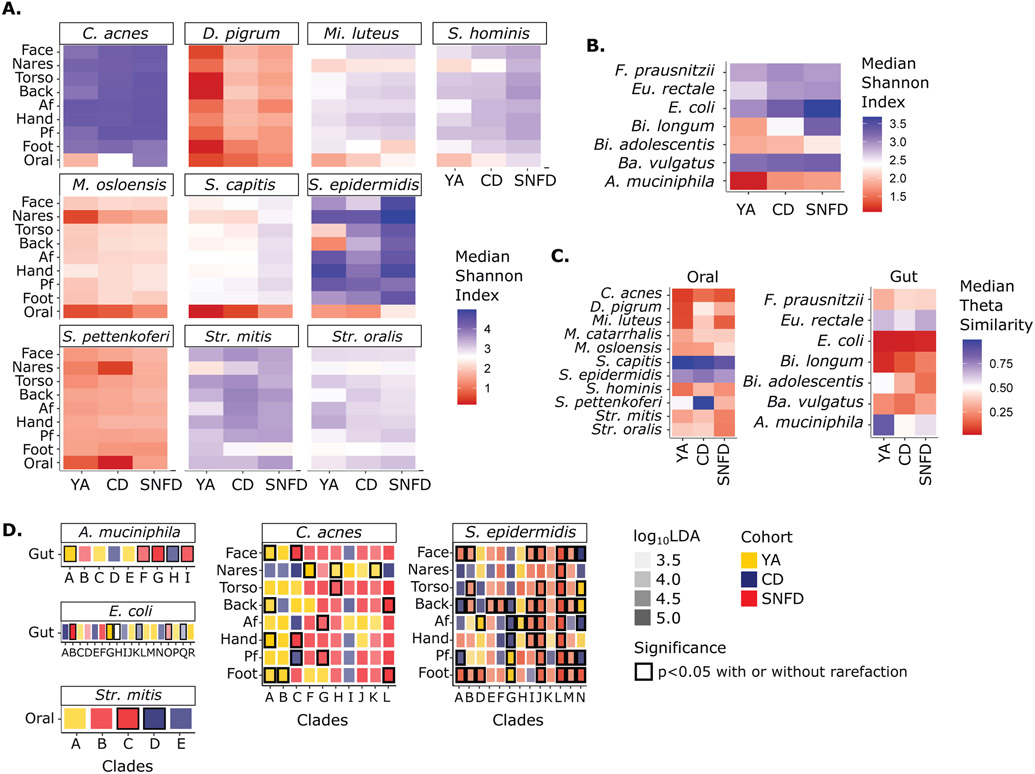
Fig. 5. Strain-Level Diversity, Heterogeneity, and Differential Abundance of Clades.
Only the in-house YA cohort was used. Strain diversity of select oral & skin (A), and gut (B) species. Previously unabbreviated species: Dolosigranulum (D.), Micrococcus (Mi.), Faecalibacterium (F.), Eubacterium (Eu.) Bifidobacterium (Bi.), Bacteroides (Ba.). Median Shannon Index of strains within species grouped by body site and cohort. (C) Heterogeneity as represented by median θ similarity of strain composition within a cohort. θ=1 represents 100% similarity and thus minimal heterogeneity. θ=0 represents no (0%) similarity, maximum heterogeneity. For A-C, Analogous Bray-Curtis dissimiliarities and analyses using data rarefied to 500,000 reads are reported in fig. S14. (D) Strain clades significantly enriched in YA, CD, or SNFD cohort. Outline color indicates significance (Kruskal-Wallis test p < 0.05). Opacity of the fill color indicates the corresponding log10LDA value. Clade assignments in D are arbitrary letters assigned to primary branches of unrooted phylogenic trees of all genomes for that species (fig S15, table S5-7). Analogous analyses using the expanded YA cohort are reported in fig. S16.